Abstract
The factor VII (FVII) protein is an integral component of the extrinsic coagulation pathway. Deleterious variants in the gene encoding this protein can result in factor VII deficiency (FVIID), a bleeding disorder characterized by abnormal (slowed) clotting with a wide range of severity, from asymptomatic to life-threatening. In canids, a single FVIID-associated variant, first described in Beagles, has been observed in 24 breeds and mixed-breed dogs. Because this variant is present in breeds of diverse backgrounds, we hypothesized that it could be a contributing factor to unexplained bleeding observed in some canine autopsy cases. DNA was extracted from paraffin-embedded tissue samples from 67 anticoagulant-negative autopsy cases with unexplained etiology for gross lesions of hemorrhage. Each dog was genotyped for the c.407G>A (F71) variant. Experimental controls included 3 known heterozygotes and 2 known homozygotes for the F71 variant, 2 normal dogs with known homozygous wild-type genotypes (F7WF7W), and 5 dogs with bleeding at autopsy that tested positive for anticoagulant rodenticide and were genotyped as F7WF7W. All 67 cases tested homozygous for the wild-type allele, indicating that the common FVIID variant was not responsible for the observed unexplained bleeding. Our work demonstrates the usefulness of retrospective studies utilizing veterinary diagnostic laboratory databases and tissue archives for genetic studies. In the case of FVIID, our results suggest that a singular molecular test for the F71 variant is not a high-yield addition to postmortem screening in these scenarios.
Factor VII deficiency (FVIID) is an autosomal recessively inherited bleeding disorder that affects ~1 in 500,000 humans.32,41 People with this disorder can have symptoms that vary widely in severity. 33 In humans, there are >250 identified variants in the F7 gene, which encodes the clotting factor VII (FVII) protein; these include missense, nonsense, and splice site changes, 38 resulting in a varied phenotypic spectrum. Indeed, the clotting phenotype can vary even in patients with identical genotypes. 19 As such, efficiently diagnosing patients using a genetic test is difficult; rather, laboratory coagulation panels are the gold standard. In contrast to the >250 human F7 variants, in canine models to date, there is only 1 published F7 variant associated with a bleeding phenotype.7,22 Although this variant was published more than a decade ago (~2006), it has not yet been investigated as the cause of unexplained bleeding in canine autopsy cases, to our knowledge.
The mammalian clotting cascade consists of proenzymes requiring activation to function, and is divided into 3 parts: 1) the intrinsic pathway, 2) the extrinsic pathway, and 3) the common pathway; the extrinsic pathway is considered more important clinically and is initiated by the interaction of tissue factor (TF) and activated FVII.21,44 The FVII protein is normally found at low plasma concentration in the bloodstream, only ~0.5 mg/L in humans. 21 FVII is a vitamin K–dependent proenzyme that is activated when a signal peptidase cleaves one Ala-Ile bond in response to TF, which is itself exposed in response to tissue damage.6,28 To continue the coagulation pathway, the FVII protein active site cleaves and activates factor IX (intrinsic pathway) and factor X (common pathway).18,31 The end point of the coagulation cascade is the fibrin clot. 44 Absence or deficiency of FVII results in a bleeding disorder. 21
FVIID can be acquired (typically as a result of an underlying lesion, e.g., liver failure, consumptive coagulopathies), but as a single factor deficiency, FVIID is more commonly recognized as a congenital inherited disorder. 38 FVIID is characterized by an inadequate amount of effective FVII protein, which results in abnormal (slowed or absent) clotting that can range in severity from asymptomatic to life-threatening.33,38 Genetic FVIID results from any variants in the F7 gene that decrease FVII protein function. 39 The mode of inheritance is typically autosomal recessive, although compound heterozygotes are also observed.19,38
FVIID can be confounding, in that severity is not always directly related to the amount of circulating FVII protein.38,39 For example, individuals with circulating FVII coagulant activity of only 1% can still be asymptomatic, or may experience only mild symptoms such as easy bruising, or can experience life-threating bleeding (such as into the central nervous system or gastrointestinal tract).21,27 Given that there are >250 documented F7 gene variants in humans, 38 and because compound heterozygotes can also experience clotting dysfunction, clinical coagulation tests are required to definitively diagnose FVIID in an individual.7,39 A FVII-deficient patient characteristically has an increased prothrombin time (PT)7,21; however, no single test is able to accurately predict bleeding risk, 38 and even specifically measuring FVII coagulant activity (FVII:C) can have its own discrepancies depending on the origin of the thromboplastin reagent used for the test itself.27,38
Direct measurement of FVII:C activity level is not conducted routinely in dogs, despite hereditary FVIID being described in dogs since 1962. 29 However, unlike humans, to date, only a single genetic variant associated with this condition in dogs is described (OMIA 000361-9615). This variant, hereafter referred to as F71, was originally described in a colony of research Beagles, in which affected dogs typically had only mild bleeding or were subclinical and were identified incidentally via pre-surgical coagulation screening. 7 The variant was also identified in pet Beagles. 7 This variant consists of a G to A missense mutation (c.407G>A; all positions used are from CanFam3.1) in exon 5 of F7, located on canine chromosome 22. The resulting amino acid substitutes glutamic acid for the original glycine (p.Gly136Glu) and results in a decrease in plasma FVII coagulant activity, inherited in a recessive manner. 7 The identical F71 variant was later observed in Alaskan Klee Kai dogs; in this breed, some homozygous dogs were subclinical or had only mild bleeding similar to the Beagles, whereas another Klee Kai experienced severe bleeding. 22 As in humans, these FVIID dogs had increased PT and were within RIs for activated partial thromboplastin time. 22 Subsequently, the F71 variant has been identified in 22 additional dog breeds (24 total) and in mixed-breed dogs (Suppl. Table 1),7,12,13,22,40,45 with a reported allele frequency of 0.487–0.614% across all mixed-breed dogs, and 0.707% in combined purebred samples.13,45 This demonstrates that F71 is an old allele, widely dispersed across and pre-dating modern dog breeds.
Infrequently, a canine cadaver submitted for diagnostic postmortem examination is observed to exhibit bleeding (e.g., into a body cavity, into the gastrointestinal tract), but this hemorrhage is not explained by rodenticide anticoagulant toxicity test results, trauma, or obvious bleeding tumors (e.g., hemangiosarcoma). Given that the F71 variant is dispersed across a wide range of breed backgrounds in the modern dog population, we hypothesized that this variant might be present in some of the dogs exhibiting unexplained bleeding at autopsy. Because the F71 variant is associated with a spectrum of bleeding phenotypes, our study is an important step in determining the variant’s broader impact. If F71 is a significant contributor to unexplained hemorrhage discovered during a postmortem examination, then including a genetic test for this variant in some autopsy screening scenarios would be indicated. Our work also exemplifies the valuable resources provided by veterinary diagnostic laboratories for genetic studies.
Materials and methods
Canine samples
DNA samples from 2 validated F71F71 dogs (breed not reported) were provided by Dr. Urs Giger at the University of Pennsylvania School of Veterinary Medicine (Philadelphia, PA, USA) to serve as positive controls. Two Shetland Sheepdogs with no known history of any clotting disorder and with previously banked DNA were used as homozygous reference wild-type (reference allele hereafter referred to as F7W) controls (approved by Purdue University’s Institutional Animal Care and Use Committee [IACUC], protocol 1812001840). Sequences from these 2 dogs matched the reference canine genome (F7WF7W). DNA from 3 known F7WF71 dogs (1 Beagle, 2 Scottish Deerhounds) was obtained to serve as heterozygous controls (carriers); these were provided via the Canine Health Information Center’s DNA bank (CHIC Program, managed by the Orthopedic Foundation for Animals, https://www.ofa.org/about/chic-program).
The database at the Animal Disease Diagnostic Laboratory (ADDL) at Purdue University’s College of Veterinary Medicine (West Lafayette, IN, USA) was searched retrospectively, and 67 cases, spanning from 2005 to 2020, were identified that fit the following inclusion criteria (Table 1, expanded in Suppl. Table 2). Cases were defined as dogs identified at postmortem with bleeding (e.g., into a body cavity or cavities or hollow organ, without obvious bleeding tumors) sufficient to trigger the request for an anticoagulant rodenticide toxicity test. All cases were required to have a negative test result for anticoagulant rodenticides. Specifically, fresh or frozen liver from each case was analyzed for the presence or absence of anticoagulant rodenticides by high-performance liquid chromatography with UV and fluorescence detection, a procedure appropriate for the detection of warfarin, coumachlor, diphacinone, dicoumarol, chlorophacinone, bromadiolone, brodifacoum, and difethialone. Excessive bleeding did not necessarily need to be identified as the primary cause of death.
Cases identified with bleeding at postmortem, grouped by assigned designation code.
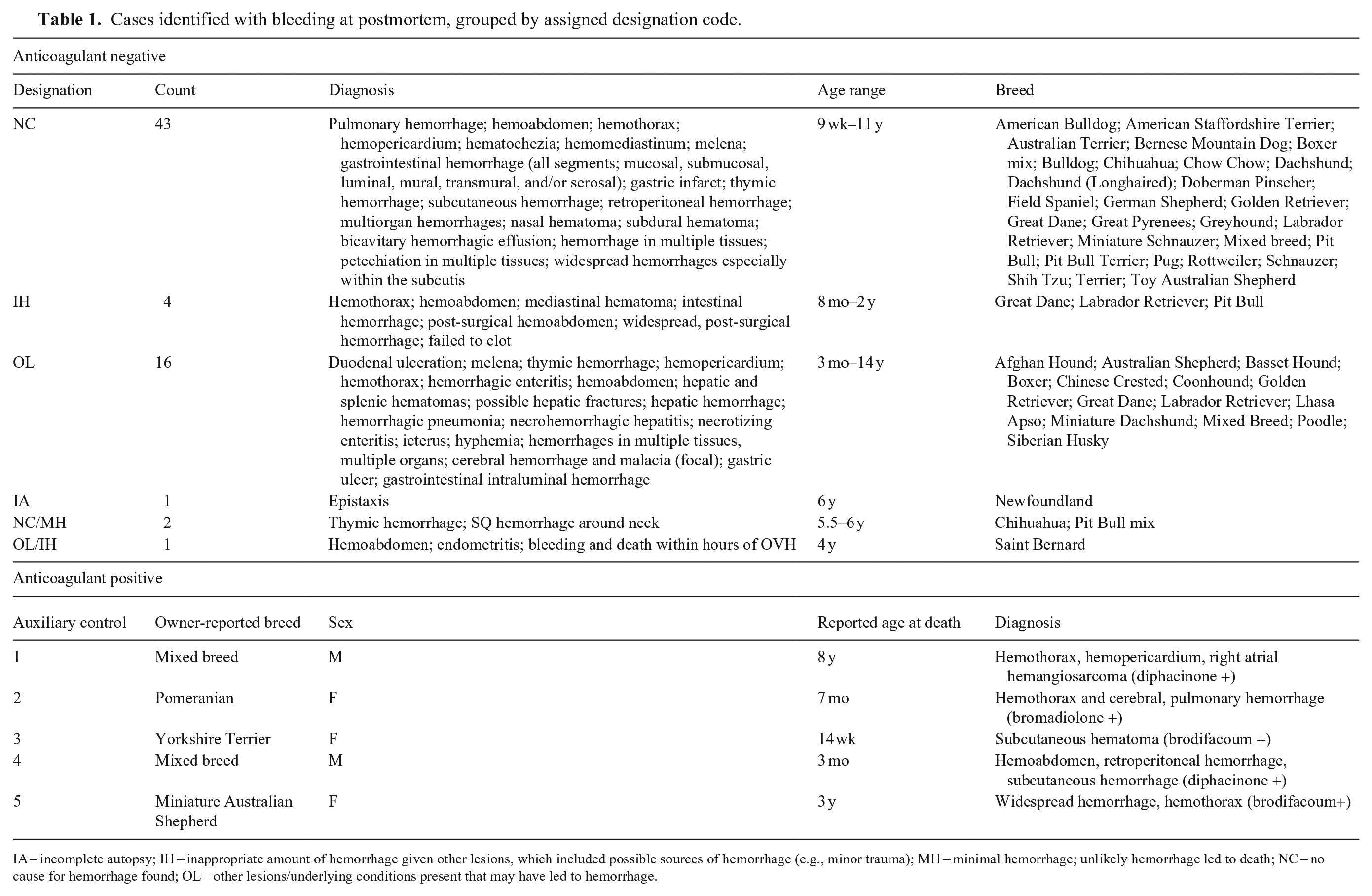
IA = incomplete autopsy; IH = inappropriate amount of hemorrhage given other lesions, which included possible sources of hemorrhage (e.g., minor trauma); MH = minimal hemorrhage; unlikely hemorrhage led to death; NC = no cause for hemorrhage found; OL = other lesions/underlying conditions present that may have led to hemorrhage.
A board-certified veterinary pathologist (G.N. Burcham) reviewed the gross findings or clinical signs and histologic findings in all cases and assigned designation codes, as appropriate (Table 1; Suppl. Table 2). In some instances, reported clinical signs (e.g., epistaxis) were considered when making designations, but no premortem testing (e.g., platelet counts, clotting time) data were available for consideration. Five codes were used: 1) NC = no cause for the hemorrhage was found; 2) IH = inappropriate amount of hemorrhage given other lesions, which included possible sources of hemorrhage (e.g., minor trauma); 3) OL = other lesions or underlying conditions that were likely or possible sources of hemorrhage; 4) IA = incomplete autopsy; and 5) MH = minimal hemorrhage that was unlikely to have led to death.
Cases coded as NC, IH, and OL had a degree of hemorrhage that was considered the likely cause of death (or, in the case of OL, an amount of hemorrhage that contributed to death alongside another serious lesion or condition). Cases designated as MH likely died of causes unrelated to hemorrhage. One case was designated as IA; this was because limited tissues had been submitted to the ADDL for examination, and the entire dog was not available for gross examination by ADDL personnel. Thus, the degree of hemorrhage, other lesions, and cause of death were not determined in this dog.
The designation codes IH and OL included lesions that may have been related to the observed hemorrhage. In cases designated IH, other lesions that would have led to small amounts of hemorrhage in a normal dog were present (e.g., minor trauma). However, in the study cases, a greater degree of hemorrhage than would have been expected given the lesions was recorded. In cases designated OL, other lesions that may have led to large amounts of hemorrhage in a normal dog were present. Thus, these cases had a large amount of hemorrhage present, as well as lesions that would explain the degree of hemorrhage observed. For example, in one case designated OL, hemoabdomen was considered a major contributing factor in death; bacterial valvular endocarditis and septic emboli were also identified in this dog, both of which likely led to disseminated intravascular coagulation, a known cause of excessive hemorrhage. Together, these categories demonstrate that our study cohort included many types of cases.
Breed, sex, and age at death were recorded. Reported breed was not verified via genetic testing. Samples from 5 dogs exhibiting similar bleeding, but with positive identification of an anticoagulant rodenticide, were also genotyped as auxiliary controls. All ADDL samples collected (n = 67 + 5 = 72) consisted of banked postmortem tissues (no IACUC approval required), either as sections cut from 64 formalin-fixed, paraffin-embedded (FFPE) blocks or 8 entire FFPE blocks.
DNA extraction
Genomic DNA was extracted from FFPE cut sections (GeneRead DNA FFPE kit; Qiagen). For entire FFPE blocks, tissue was removed in bulk (gouged out), and DNA was then extracted (Gentra Puregene tissue kit; Qiagen), following the Hemo-De protocol according to the manufacturer’s instructions.
Primer design and PCR
Given that the originally published location for F71 was from the CanFam2.0 reference genome (g.63537727G>A), this position was converted to CanFam3.1 (g.60578895G>A) via the University of California–Santa Cruz BLAT program (https://genome.ucsc.edu/cgi-bin/hgBlat). Primers were designed to amplify the region surrounding the F71 variant, located on CFA 22, in exon 5 of F7, using Primer3 (https://primer3.ut.ee/). The resulting primers were: forward primer 5′-TGGGCTAAGCTCTTCAGACG-3′ and reverse primer 5′-ACATGCTCACCCTTCCTGTC-3′; these generated a 443-bp product. Standard PCR conditions, using KOD Xtreme hot start DNA polymerase (MilliporeSigma), were used to generate PCR products.
Sanger sequencing and sequence analysis
PCR products were cleaned and prepared for sequencing using the ExoSAP-IT PCR Product Cleanup (Thermo Fisher) protocol. Samples were then submitted to the Purdue Genomics Core Facility or Eurofins Genomics for Sanger sequencing.
Sequencher v.5.4.6 (Gene Codes) genetic sequence analyzer was utilized to visualize the sample sequence chromatograms. All sequences were manually examined for the presence or absence of the F71 variant. All dogs were classified as homozygous reference or wild-type (F7WF7W), heterozygous (F7WF71), or homozygous for the known variant (F71F71).
Results
The 15-y period that we searched resulted in a case population of 67 dogs. Sexes were essentially equally represented (male = 34, female = 33). The reported age at death ranged from 9 wk to 14 y. The bleeding phenotypes represented a wide array of clinical pictures, from cases with extensive, whole-body hemorrhage, to cases with more localized, mild hemorrhage. We designated 43 cases NC (Table 1; Suppl. Table 2), indicating that no cause for hemorrhage was found. The other 24 cases represented scenarios such as minimal hemorrhage (in which the hemorrhage was unlikely to have led to death, but may still have represented a bleeding disorder), or a potential source of bleeding (e.g., minor trauma) in which the amounts of hemorrhage were inappropriate for that potential source. Breeds within our case population in which the F71 variant has been reported included the Basset Hound, Miniature Schnauzer, Miniature Dachshund, and mixed-breed dogs.
Sequences from all experimental control samples (known homozygous reference, F7WF7W; heterozygous, F7WF71; and homozygous for the known variant, F71F71) generated the expected genotypes (Fig. 1A–C), wherein the reference or wild-type nucleotide at this locus is a guanine, and the F71 nucleotide at this locus is an adenine. FVIID-affected Beagles and Alaskan Klee Kais are reported to be homozygous for adenine at this locus (as is the dog in Fig. 1C).7,22 All of the 67 case dogs as well as all 5 of the anticoagulant rodenticide–positive dogs genotyped as F7WF7W, identical to the homozygous reference or wild-type control (Fig. 1A).
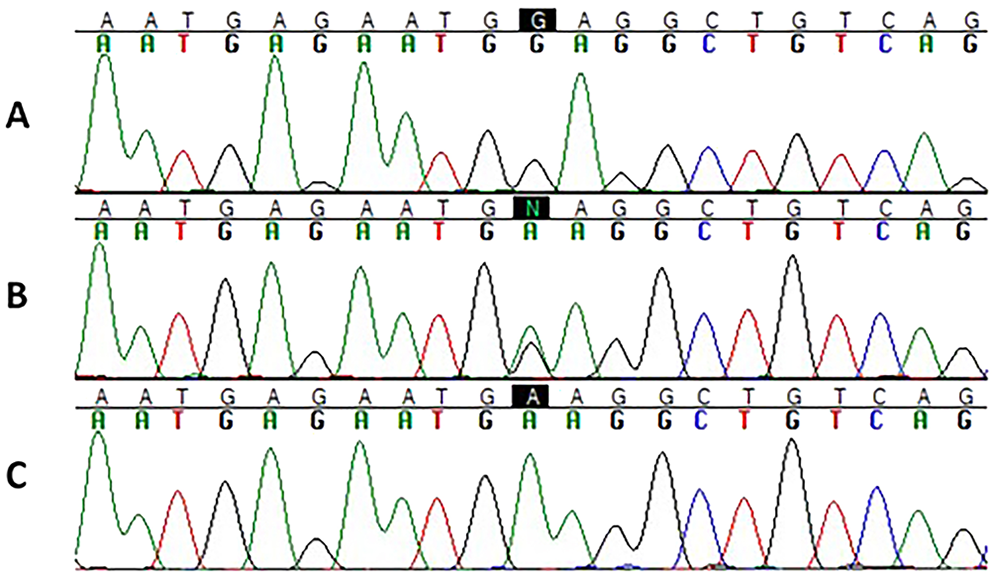
Sequence electropherograms of the F71 locus. The black boxes highlight the nucleotide of interest.
Discussion
Unlike the situation in humans, only one disease-associated variant has been described to date in the canine FVII gene. Here we report that this variant (F71) is unlikely to be the cause of unexplained bleeding in most autopsy cases, given that the variant was not detected in our sample cohort. We can further conclude that a singular genetic test for this variant in autopsy cases with unexplained bleeding is not a high-yield test. The lack of representation of the F71 allele was supported by the fact that all 5 of the anticoagulant rodenticide–positive cases also did not have the F71 variant.
Our assembled sample cohort was strikingly balanced by sex, with a nearly equal proportion of males and females. This finding suggests that, overall, an X-linked disorder (either hemophilia A, caused by a deficiency of factor VIII, OMIA 000437-9615; or hemophilia B, caused by a deficiency of factor IX, OMIA 000437-9615; both of which occur in humans and dogs5,24,30) is unlikely, if all cases experiencing the same condition. It is still possible that males in the study experienced an X-linked bleeding disorder; none of the case cohort dogs were genotyped for any hemophilia A or B variants. Interestingly, although 2 different hemophilia A variants have been described in German Shepherds,9,26 and our sample cohort contained 6 German Shepherd cases, 5 of the 6 were female. Although this finding does not rule out hemophilia A in these cases, the sex pattern is not consistent with an X-linked disease.
The wide range of ages at death suggests a varied clinical picture, as do the designation codes (i.e., several different types of cases were investigated), and the breed variety. Of breeds previously reported to carry the F71 variant, very few were represented among our cases. These included the Basset Hound, Miniature Schnauzer, and Miniature Dachshund. None of the breeds in our case cohort were confirmed genetically, and some lacked important details; for example, the F71 variant has been identified in the Redbone Coonhound, whereas in our cohort a dog was reported simply as Coonhound, which could represent one of many breeds. Similarly, one of our cases was identified as Schnauzer, without further classification; the F71 variant has been found in both the Giant and Miniature Schnauzer 12 ; therefore, this case could be from an F71 carrier breed. We also had many mixed-breed dogs in our cohort, and mixed-breed dogs are reported to carry the F71 allele at a very low frequency13,45; however, this situation is highly dependent on the contributing breed background, which is unknown for samples in our cases.
The F71 variant is located within the second epidermal growth factor–like domain (EGF-2) of canine F7. 7 This domain is an integral component of protein–protein or protein–ligand interactions 1 ; specifically, the EGF-2 domain is necessary for FVII and TF to interact, 8 and TF binding is the step required to activate FVII. 34 In humans, EGF-2 mutations have been shown to dramatically impair FVII:C activity by specifically affecting protein–protein interactions. 39 Therefore, it is likely that the amino acid change seen in this domain in dogs also impacts this interaction, decreasing the coagulation activity of the FVII protein. Other work with the human FVII protein has shown that 2 missense mutations in the EGF-2 domain markedly reduced activity levels as a result of reduced secretion, 20 and indeed, the canine F71 allele’s protein was retained intracellularly significantly more often than the wild type (p < 0.0001). 7 When the canine F71 allele was first characterized, dogs homozygous for the variant were shown to have a marked decrease in FVII activity. 7 Further supporting this hypothesis is the fact that a p.G156D variant was observed in a human patient who had measurably reduced FVII activity, 19 which is significant because residue 156 in humans is the homologous residue of interest (residue 136) in dogs.
Given that all 67 cases genotyped as homozygous for the reference or wild-type allele (F7W), the F71 allele cannot be contributing to the unexplained bleeding seen in these autopsy cases. Hence, continued testing for this variant is likely unnecessary, even in cases with unexplained bleeding and no detection of anticoagulant rodenticide. It is interesting to note that our cohort represents 4–5 cases per year at the Purdue ADDL that receive no definitive explanation for their bleeding. Despite the negative findings, our study clearly demonstrates the valuable resources found in veterinary diagnostic laboratories, such as the Purdue ADDL, for these types of genetic studies. Veterinary diagnostic laboratory databases allow for retrospective case identification, and archived FFPE tissues can be a key source of DNA samples.
Considering that the F71 allele is present in many dog breeds, and that the phenotypic expression varies between individuals and breeds, it is possible that these unexplained bleeding cases may involve other genetic variants, perhaps within the F7 gene, or elsewhere, such as in other clotting factor genes or unknown regulators. For example, in humans, mutations in the F7 promotor have led to severe, or even lethal, FVIID by impairing binding of transcription factors. 17 Similarly, also in humans, an HSA 8 gene locus (a remote locus; F7 is located on HSA 13) may play a role in regulating FVII levels 14 ; no similar regulatory regions have yet been published in dogs. The >250 known human F7 variants include missense, nonsense, small indel, and splice site changes that can be found in every region of the gene and affecting any of the protein domains25,39; therefore, other deleterious F7 variants could exist in dogs.
Another well-known inherited bleeding disorder in dogs, von Willebrand disease (VWD; OMIA 001057-9615, 001339-9615, 001058-9615), could possibly be undiagnosed in some of our cases. VWD is separated into types associated with several variants (at least one of which may not be causal) and is found in dozens of dog breeds.2,5,10,12,13,23,24,36,37,42,43 None of the case cohort samples were tested for any of the VWD variants; however, several of the breeds represented in our cases are documented to carry VWD variants, including the Doberman Pinscher, Chinese Crested, Great Dane, and others. Given that VWD variants are also common in mixed-breed and purebred dogs, 13 testing our case population for appropriate VWD variants is a logical next step for future work.
Although FVII deficiency, hemophilia A and B, and VWD are all found in many breeds, most other canine genetic hemostatic disorders with known contributing variants are highly breed-specific (e.g., thrombasthenic thrombopathia in Otterhounds, 3 macrothrombocytopenia in Cavalier King Charles Spaniels 11 and Cairn and Norfolk Terriers, 16 and P2Y12-related bleeding disorder in Greater Swiss Mountain dogs4,15). As in antemortem testing, such tests are applicable only in breed-specific scenarios, and would not be broadly applicable in a study such as ours.
The lack of complete clinical history for our case dogs is a significant limitation of our study; without this information, the dog’s previous coagulation history (clinical and laboratory) was unknown. For example, for most cases the dog’s antemortem liver function was unknown; given that FVII is synthesized exclusively by the liver, 38 hepatic disease (e.g., malignancy, autoimmunity) could impact FVII production despite a normal F7 genetic sequence. Similarly, antemortem coagulation testing results would have been helpful, particularly for identifying acquired defects in hemostasis. This dearth of health history information is not uncommon with whole-body cadaver autopsy submissions. Because we used only one pair of primers, an additional polymorphism(s) within the primer location could possibly, in a breed-specific manner, create allele dropout. This situation was reported by one testing laboratory for Beagles. 35 In our study, allele dropout could have created false negatives that would only have potentially been detected with a second genotyping assay. Another limitation is the scope of our study: we focused on only one specific nucleotide (and therefore one amino acid) within canine F7. It is possible that some of the homozygous F7WF7Wcases have a FVIID genetic cause for their bleeding, but any variants outside the targeted region in F7 would not have been detected.
We have demonstrated that the widespread F71 variant is not a common cause of unexplained bleeding, with the implication that testing dogs for this solitary variant is not a logical step. Taken together, the most expeditious future approach on a research basis would be whole-genome sequencing (WGS) of case samples, which allows simultaneous enquiry of all genes, including the entire coagulation pathway and all known variants (such as VWD variants), and could unearth new genetic causes of congenital hemorrhagic disease. This approach would genotype all known variants previously associated with canine bleeding disorders, and also reveal any new or different variants within F7 or other genes. However, WGS is not yet practical as a routine screening test given the volume of data generated, time required for analysis, and cost. A genetic panel test combining all of the known canine coagulopathy variants could still be a useful screening assay for the types of cases described here.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221118581 – Supplemental material for Investigation of a common canine factor VII deficiency variant in dogs with unexplained bleeding on autopsy
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221118581 for Investigation of a common canine factor VII deficiency variant in dogs with unexplained bleeding on autopsy by Jessica A. Clark, Stephen B. Hooser, Dayna L. Dreger, Grant N. Burcham and Kari J. Ekenstedt in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
Shawna Cook is sincerely thanked for her assistance with preparation of this manuscript. We also gratefully acknowledge the many pathologists and pathologists-in-training at the Purdue ADDL who contributed gross and histopathology reports and diagnostic expertise to the ADDL database used to identify the study population, and to Dr. Christina Wilson-Frank for analytical testing.
Declaration of conflicting interests
The authors declared no conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Kari J. Ekenstedt was supported by the Office of the Director, National Institutes of Health (NIH), under award K01-OD02751.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
