Abstract
Clonality assays based on antigen receptors are used as adjunct examinations in the diagnosis of lymphoproliferative diseases. We investigated the usefulness of the T-cell receptor beta (TRB) and T-cell receptor delta (TRD) loci in clonality assays for high-grade gastrointestinal (GI) lymphoma in dogs. For TRB, we used primers reported previously; for TRD, we designed primers for each of the V and J genes based on genomic sequences. Genomic DNA was extracted from 39 formalin-fixed, paraffin-embedded sections of high-grade GI lymphoma diagnosed histologically. The sensitivity of TRB and TRD primers for GI lymphoma was 41.0% and 38.5%, respectively, which was lower than the 82.1% sensitivity of T-cell receptor gamma (TRG) primers However, some cases that could not be detected using TRG primers had clonality with either TRB or TRD primers. We found the TRG locus to be more suitable as a first choice for the assay of canine lymphoma clonality than the TRB and TRD loci. However, the detection rate of T-cell clonality may be enhanced using TRB and TRD primers for lymphoma cases not detected using TRG primers.
Clonality assays based on PCR analysis of antigen receptor genes have been used as adjunct examinations in the diagnosis of lymphoproliferative diseases, including lymphoma.1,7 The PCR targets are the immunoglobulin genes for B-cell and T-cell receptor genes of T-cell origin. Although the diagnosis of lymphoma is made based on histologic examination and particular immunologic features, wherein CD3 is a marker for T cells and CD20 is a marker for B cells, some cases may remain ambiguous based on these examinations.
Clonality assays could assist in distinguishing lymphomas from reactive lesions, wherein obvious PCR products indicate clonal expansion of lymphocytes. 9 Particularly in the case of gastrointestinal (GI) lymphoma, clonality assays are useful not only as adjunct examinations but also for prognostication. The prognosis of cases diagnosed as chronic enteropathy could be guarded if clonality is detected.5,17 In such cases, GI lymphoma may be suggested.5,17 Furthermore, studies have reported the beneficial use of clonality assays in differentiating chronic enteropathy from lymphoma.10,14 Clonality assays are essential for the diagnosis and prognosis of GI lymphoma in dogs.
In a human clonality assay, 3 T-cell receptor (TR) genes, including beta (TRB), gamma (TRG), and delta (TRD), have been used. 11 TRG is a first-line target given that it has been observed to rearrange in most types of T cells. TRB is transcribed in αβ-T cells, which are the predominant lymphocyte population. 3 Therefore, TRB, along with TRG, is used preferably as a target in human clonality assays. 21 Although TRD is transcribed in γδ-T cells, TRD genes are deleted in αβ-T cells because these genes are present at the locus that is the same as that for T-cell receptor alpha.3,4 Therefore, TRD may still be a target for a T-cell clonality assay, especially in γδ-T-cell proliferation and immature T-cell diseases.6,11 The human clonality assay that utilizes 3 TR gene loci can provide a high detection rate. 2
In contrast, TRG has been used as the only target in a canine clonality assay.7,9 A case report addressed concurrent T-zone lymphoma and B-cell lymphoma in a dog in which primers for the TRB locus were described, 12 but detailed descriptions of the design and specificity of primers were not mentioned. Furthermore, the TRD locus has not been reported for canine clonality analysis. Another study showed that the detection rate of a clonality assay using the TRG locus in high-grade intestinal lymphoma in dogs was 82%, 18 which was lower than that in a human T-cell clonality assay. 2 A combination of TRG, TRB, and TRD in a feline clonality assay improved the detection rate of lymphoma. 16 The use of the TRB and TRD loci in a canine clonality assay may be valuable.
We examined the TRB and TRD loci as targets in a T-cell clonality assay of high-grade GI lymphoma in dogs. We investigated 39 cases of canine GI lymphoma and 23 cases of lymph nodes without lymphoma, as described previously, 18 wherein lymph nodes without lymphoma were obtained for evaluation of metastasis of non-lymphoid malignant tumors, including mammary tumor, squamous cell carcinoma, thyroid carcinoma, and undetermined epithelial tumor. 19 In brief, genomic DNA (gDNA) was extracted from formalin-fixed, paraffin-embedded (FFPE) GI lymphoma and lymph node sections (High pure PCR template preparation kit; Roche) per the manufacturer’s instructions. Extracted gDNA was adjusted to 10 ng/µL and stored at −20°C until used further. The integrity of the gDNA was verified using 3 primer sets, including immunoglobulin heavy-chain constant region, ATP7B, and SOD1, as described previously. 19
The primers of the TRB locus that we used have been reported previously (Table 1). 12 TRB primers were compared with the TRBV and TRBJ1 genes mapped on the International ImMunoGeneTics (IMGT) database (accession IMGT000005, https://www.imgt.org/) as well as with the TRBJ2 gene, which was reported in a previous study, 13 because of the absence of annotation of TRBJ2 genes in IMGT000005. To design the primers for the TRD locus, a genomic sequence containing TRDV and TRDJ genes (accession IMGT000004) was chosen from the IMGT database. Five V genes and 4 J genes were mapped on the TRD locus, and primers for each of the V and J genes were designed using Primer3Plus (https://www.bioinformatics.nl/cgi-bin/primer3plus/primer3plus.cgi; Table 2). Genomic DNA derived from FFPE sections may have low integrity of DNA degraded by formalin given that a study of a feline clonality assay showed less amplification efficiency of ~300-bp fragments. Therefore, we designed the primers to amplify <200 bp.
TRB primer sequences and matched TRB genes used in our study of gastrointestinal lymphoma in dogs.
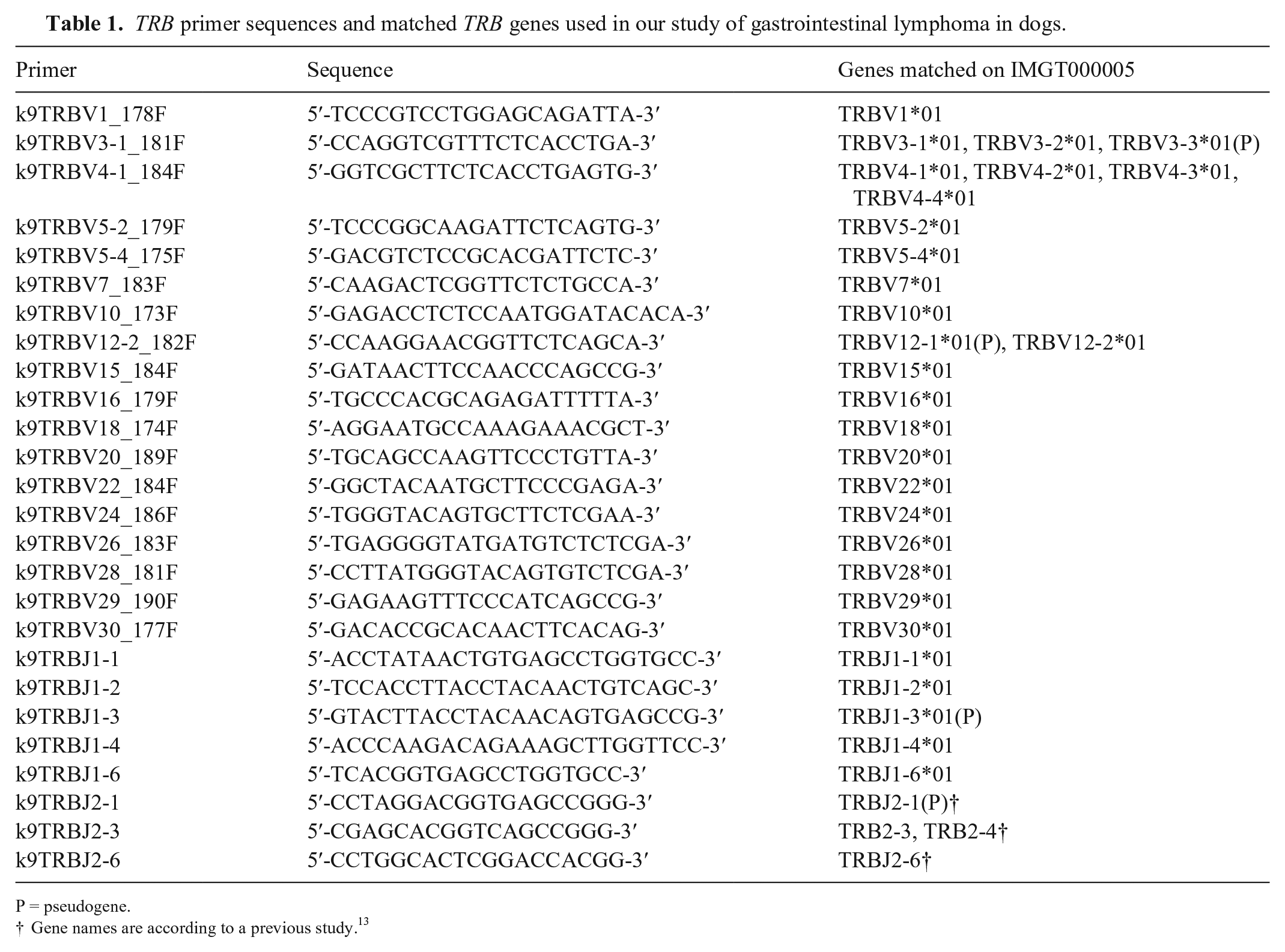
P = pseudogene.
Gene names are according to a previous study. 13
Primers for each TRD gene designed in our study of gastrointestinal lymphoma in dogs.
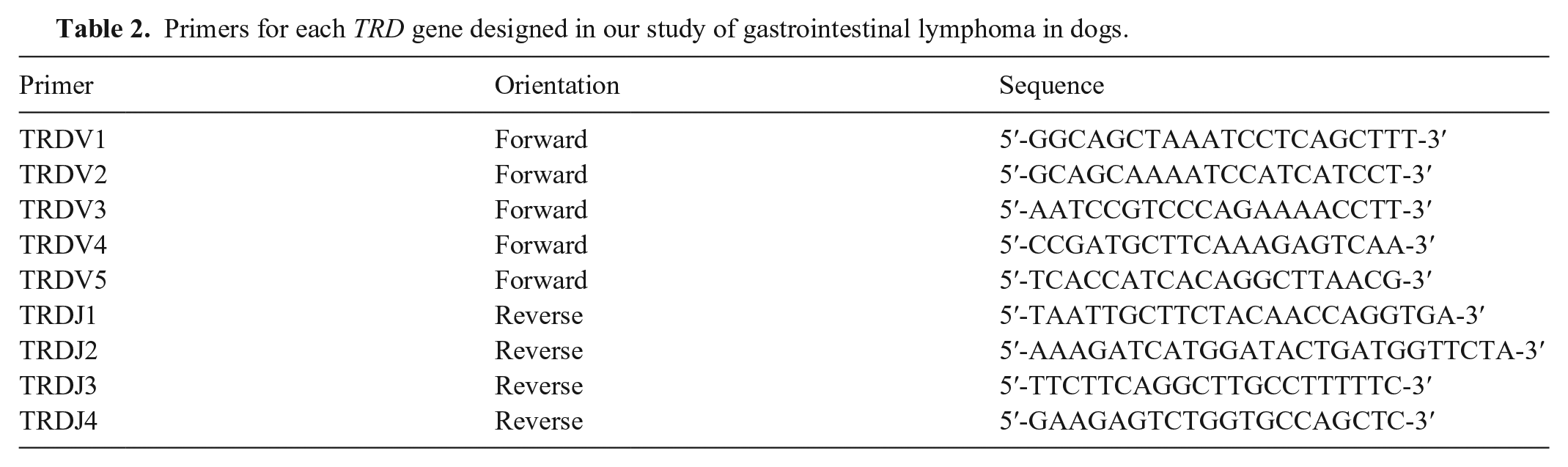
All primers were used in the same tube for TRB or TRD. PCR was performed using 20 ng of gDNA in a total volume of 10 µL containing 1 µL of 10× PCR buffer, 1 µL of 2-mM dNTPs, 1 µL of each forward and reverse primer (2 µM), and 0.5 units of DNA polymerase (Blend Taq plus; Toyobo Life Science). The PCR conditions were as follows: 95°C for 1 min and 40 cycles of denaturation at 94°C for 20 s, annealing at 60°C for 20 s, and extension at 72°C for 20 s. The PCR products were analyzed using capillary electrophoresis with a high-resolution cartridge (Qsep100; BiOptic). A clonality assay was prepared in duplicate tubes for each locus to verify reproducibility. Clones with 1, 2, or several distinct and reproducible peaks that were 2-fold higher than the polyclonal base were defined as monoclonal, biclonal, and oligoclonal, respectively. Non-reproducible peaks in duplicate samples were defined as pseudoclonal. Clones with a Gaussian distribution were defined as polyclonal. The absence of a PCR product was defined as “no amplification.” The sensitivity, specificity, and 95% CI of each locus were calculated from the obtained data.
To verify the alignment of TRB primers to a sequence consisting of 18 V genes and 8 J genes, each primer sequence was aligned with the gDNA obtained from the IMGT database and a previous study 13 (Table 1). Of the 36 TRBV genes mapped on IMGT000005, 23 V genes completely matched with one of the V gene primers. The remaining 13 V genes remained unmatched with any of the TRBV genes. Six TRBJ genes, named TRBJ1-6, were annotated on IMGT000005, with 5 of these genes matching 5 TRBJ primers. One TRBJ1 gene (TRBJ1-5*01), which is a functional gene, was not covered by the primer set. Three TRBJ2 primers were matched to TRBJ2-1, 2-3, 2-4, and 2-6 genes. TRBJ2-2 and 2-5, which are functional genes, were not covered by the primer set. In our clonality PCR, the TRB primers amplified 150–220-bp products, and the TRD primers amplified 110–150-bp products (Fig. 1; Suppl. Table 1).
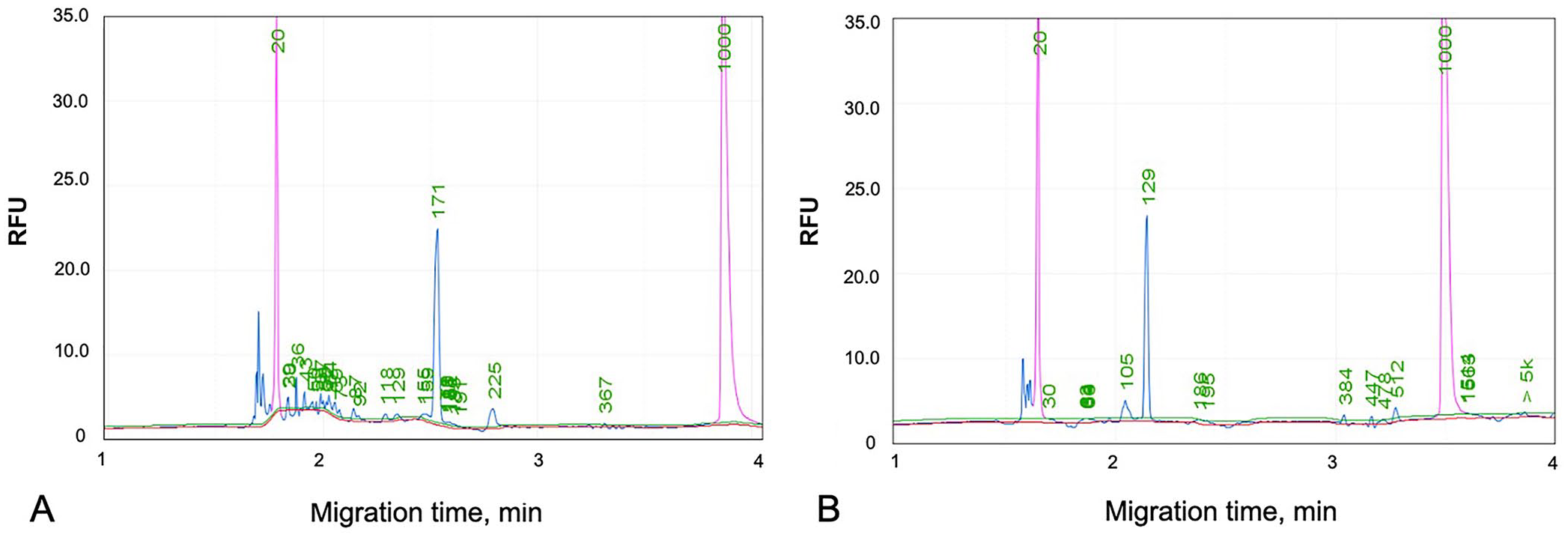
Elution profiles of capillary electrophoresis in clonality assays using TRB and TRD primers. The 2 pink peaks are markers of 20 and 1,000 bp.
Of the 39 lymphoma cases, we obtained clonal results from 16 samples for the TRB locus (41.0%, 95% CI: 27.1–56.6%; Table 3) and 15 samples for the TRD locus (38.5%, 95% CI: 24.9–54.1%; Table 3). Of the 16 cases with clonality for the TRB locus, 5 had clonality in both TRB and TRD primers. Of the 39 cases, 27 had clonality in either TRB or TRD primers (69.2%); clonality for TRG primers was 82.1%. The detection rates of TRB and TRD primers were also lower than those of TRG primers reported previously. 18 Evaluation of primers using gDNA derived from lymph nodes without lymphoma showed that 2 cases had clonality in either TRB or TRD primers. The specificity of both loci was 95.7% (95% CI: 77.3–100%).
Sensitivity and specificity of each primer set for TRB and TRD loci in cases of gastrointestinal lymphoma in dogs.
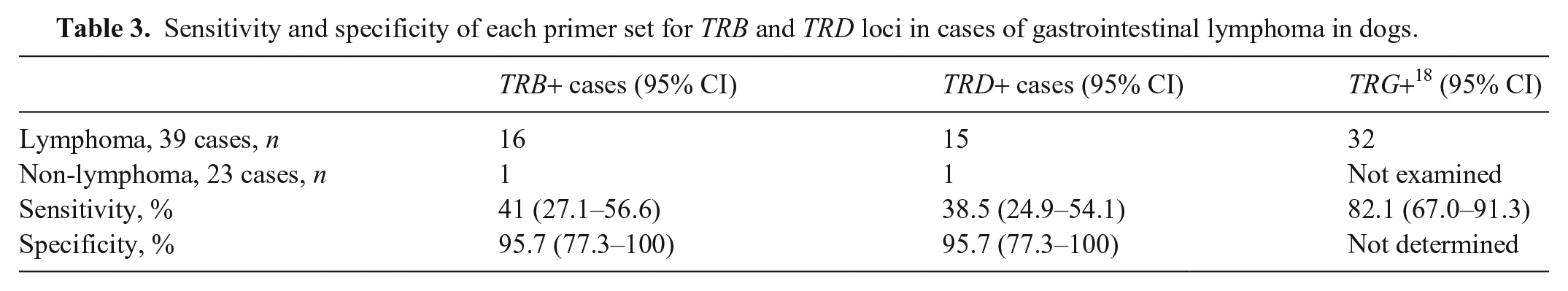

We found that the detection rate using TRB primers in high-grade canine GI lymphoma was lower than that using TRG primers. This may be a result of the restriction of coverage of TRB primers to the target sequences. 12 The reason for this is given below; a previous study has shown 36 V genes of TRB on the IMGT contig. 15 Only one functional TRBV gene (TRBV25*01) showed no match to any primer. Of the 10 functional TRBJ1 and TRBJ2 genes, 3 TRBJ genes (TRBJ1-5, TRBJ2-2, TRBJ2-5) were not covered by the primer set used in our study. A study reported that 55% of cDNA clones of TRB prepared from peripheral blood retained the TRBJ2 gene. 13 Furthermore, the use of pseudogenes in TRBV and TRBJ genes instead of functional genes during rearrangement has been speculated. 13 Therefore, using primers to identify pseudogenes or adding primers to match the uncovered TRBJ genes may improve the detection rate of the clonality assays.
We found that the detection rate of TRD primers was lower than that of TRG primers, although the primers were designed to fully match all TRDV and TRDJ genes. No PCR products, even polyclonal products, were observed in all cases without clonal reaction, suggesting the removal of TRD genes. This phenomenon is similar to that observed in a study of a feline TRD clonality assay. 16 A few reports have described the incidence of γδ-T-cell lymphoma in dogs 8 ; however, no studies have focused on the incidence of GI lymphoma. Therefore, compared with TRG, the use of TRD in clonality assays is restricted.
Our study has some limitations. In our clonality assays using gDNA prepared from lymph nodes without lymphoma, 2 samples had clonality for either TRB or TRD primers. None of the other non-lymphoma samples had clonality, indicating that the specificity of the clonality assay for TRB and TRD was 95.7% (95% CI: 77.3–100%). There are 2 hypotheses regarding clonal reaction in normal lymph node samples: true clonal or clonal expansion in response to antigenic stimulation. 9 For the former hypothesis, given the lack of follow-up clinical information on the cases, it is unclear whether these cases with clonality in TRB or TRD primers subsequently developed lymphoma. It was impossible to determine the true clonal nature of these cases. For the latter hypothesis, clonal expansion in response to antigenic stimulation cannot be ruled out because all lymph nodes without lymphoma were obtained from dogs that were being examined for metastasis of a malignant tumor. 20
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221116285 – Supplemental material for A clonality analysis based on T-cell receptor beta and delta loci for high-grade gastrointestinal lymphoma in dogs
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221116285 for A clonality analysis based on T-cell receptor beta and delta loci for high-grade gastrointestinal lymphoma in dogs by Masamine Takanosu and Yumiko Kagawa in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
