Abstract
Technologic advances in information management have rapidly changed laboratory testing and the practice of veterinary medicine. Timely and strategic sampling, same-day assays, and 24-h access to laboratory results allow for rapid implementation of intervention and treatment protocols. Although agent detection and monitoring systems have progressed, and wider tracking of diseases across veterinary diagnostic laboratories exists, such as by the National Animal Health Laboratory Network (NAHLN), the distinction between detection of agent and manifestation of disease is critical to improved disease management. The implementation of a consistent, intuitive, and useful disease diagnosis coding system, specific for veterinary medicine and applicable to multiple animal species within and between veterinary diagnostic laboratories, is the first phase of disease data aggregation. Feedback loops for continuous improvement that could aggregate existing clinical and laboratory databases to improve the value and applications of diagnostic processes and clinical interventions, with interactive capabilities between clinicians and diagnosticians, and that differentiate disease causation from mere agent detection, remain incomplete. Creating an interface that allows aggregation of existing data from clinicians, including final diagnosis, interventions, or treatments applied, and measures of outcomes, is the second phase. Prototypes for stakeholder cooperation, collaboration, and beta testing of this vision are in development and becoming a reality. We focus here on how such a system is being developed and utilized at the Iowa State University Veterinary Diagnostic Laboratory to facilitate evidence-based medicine and utilize diagnostic coding for continuous improvement of animal health and welfare.
Introduction
Diagnostic coding is widely used in human medicine, and to various degrees across veterinary medicine, and can standardize reporting of clinical signs and symptoms, test ordering, or disease diagnosis. However, unlike human medicine, there is no external or widely used uniform veterinary coding system. Individual medical centers and laboratories often use their own coding systems or occasionally the system is shared between institutions. We will use diagnostic (Dx) coding here to reflect the diagnosis of disease based on the available evidence provided by pathology and ancillary testing, including but not limited to bacterial culture, molecular testing, serology, toxicology, and virology, with a system designed specifically for a veterinary diagnostic laboratory (VDL).
Dx codes and associated case data are useful for monitoring health changes over time in a patient or population, for disease surveillance, or for regulatory, administrative, or insurance purposes. Additional value can be derived from certain categories of Dx codes in production animal medicine, where aggregation of case codes across designated populations can be analyzed to inform health intervention strategies and management decisions and monitor their outcomes. Ideally, a coding system design fits the intended purpose of the data output with considerations of utility during coding. Here, we describe the disease Dx coding system employed at the Iowa State University VDL (ISU-VDL; Ames, IA), extensively modified for greater accessibility and applicability of data for both internal (diagnosticians, researchers, veterinary medicine educators, administrators) and external (veterinary clinicians, government officials, and pharmaceutical companies) users. We describe what we did and why, present a preliminary assessment of interobserver agreement, how we address continuous improvement, and discuss future applications of Dx coding within our laboratory, by our internal and external users, and across veterinary medicine.
Diagnostic coding background and vision
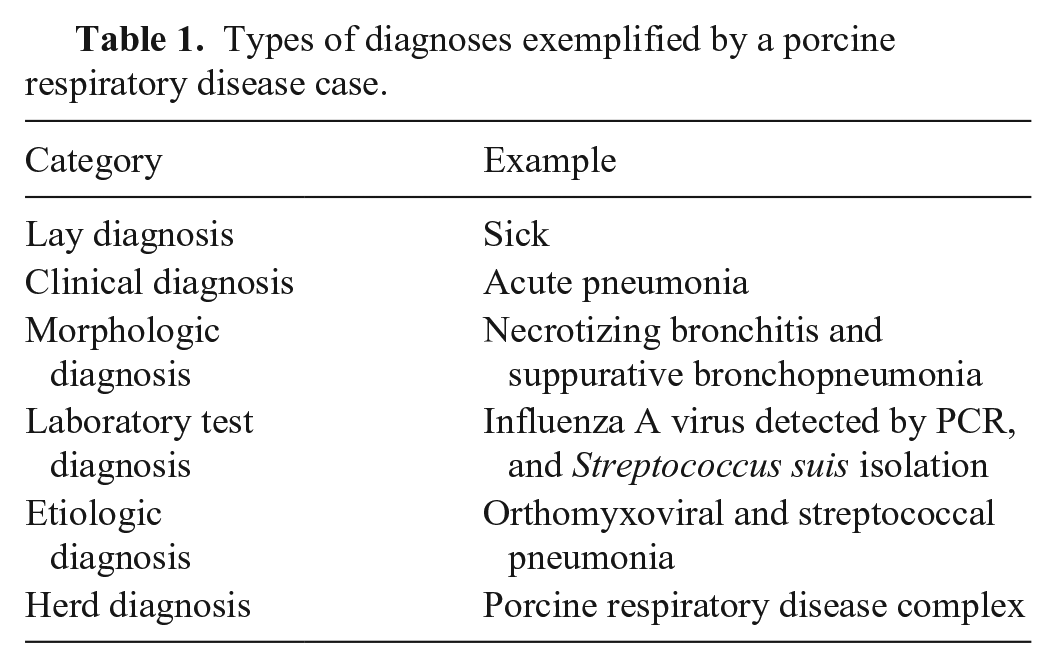
Submission history and laboratory data of various types (patient identification, signalment, demographics, medical history, tests, and results) are routinely captured and stored in some type of data warehouse in most diagnostic laboratories. Codes are commonly used in VDL laboratory information management systems (LIMSs) to indicate patient type, signalment, demographics, animal location, sample type, test type, test result, lesion, or system(s) affected clinically. None of this information confirms a particular disease or etiology without some interpretation of context. Context is provided in the submission history, clinical signs, and/or pathologic changes that are observed in tissues. Without the context of clinical signs or pathologic changes, results of tests are useful but may not confirm disease status or causation, particularly with endemic agents in populations. However, this information coupled with diagnostician input is the basis of individual case reports, which are often accompanied by interpretive comments that may state the disease process(es) present and confirmed or likely cause(s) as the “diagnosis.” The diagnostic pathologist may state their findings in the form of a morphologic diagnosis, an etiologic diagnosis, or a summary of their interpretation of available information, without clearly stating a laboratory diagnosis in the case report. The lab-rendered diagnosis is not necessarily a definitive diagnosis but one of several interconnected diagnoses (Table 1). Although the term diagnosis may imply a different meaning to different users, a Dx code breaks a diagnosis into its critical components in a repeatable, understandable, and searchable format.
Types of diagnoses exemplified by a porcine respiratory disease case.
In population medicine, a diagnostic pathologist must consider whether the animals sampled, sample types, and timing of sampling accurately represent the clinical syndrome reported or the diagnostic question posed by the submitting veterinarian, or both. Thus, Dx coding is an attempt to summarize the pertinent medical issues and the likely causes for them. However, the ultimate benefit of Dx codes is not for the individual case; it is the ability to generate aggregate data that can help define epidemiologic trends, influence management decisions, or identify underlying comorbidities or conditions that would not be evident at the individual case report level. The purpose of a disease Dx code is to provide a method to identify and retrieve cases of historical interest, either for academic purposes (particular lesions, case types, etiology) or for stakeholders to monitor or benchmark key disease parameters in their herds or practices. Dx coding is not intended to replace a case report, but to augment its utility.
Diagnostic coding at the ISU-VDL
Disease Dx codes at the ISU-VDL were originally developed for laboratory personnel to capture and archive results of diagnostic investigations. Accuracy in coding was important to facilitate retrieval of specific cases along with basic demographic data to capture species, geography, time, and disease process or etiology. However, this coding system had become cumbersome and obsolete because of drift in the structure of codes, gaps in coding, and some redundancies. The system was designed originally to not just capture Dx codes, but test-report codes as well, such as bacterial isolation of an organism, and the system was not hierarchical. All of these factors resulted in difficulty when searching for appropriate codes while coding or querying the database, as well as considerable variation in the number and specificity of Dx code(s) assigned as relevant on a specific case. Search and analytical capability in the previous system was limited and challenging because it required the user to have historical knowledge of the system and an understanding of how each pathologist would have coded their cases. Additionally, there was not an institutional inclination, at the diagnostician level, to code with purpose, nor was there feedback and discussion of how cases were coded. For the new system, we envisioned an information system to capture disease and etiology data that is intuitive, accurate, and consistent for disease diagnoses across animal species with analytical capability for real-time summaries, historical analysis, and benchmarking at individual animal, farm site, production system, and regional and national levels.
Germane to the development of our coding system is an understanding of case submission at the ISU-VDL. The full-service ISU-VDL receives nearly 100,000 accessions (cases) per year and performs over one million individual tests. Approximately 14,500 cases per year have submission history, tissues, and gross and/or microscopic pathology performed. Species breakdown heavily favors food animal, predominantly swine (70–75% of cases) and bovine (10–15% of cases), as well as over 2,000 cases annually representing other animal species that are required to have one or more Dx codes assigned. Most of these cases include histopathology. Most cases submitted from herds or flocks have additional testing performed sufficient for confidence in diagnosis, and these tests are considered in the assignment of appropriate Dx codes.
A critical driver for a new coding system was the desire to better link information within and between cases and across populations. For temporal analysis of disease status within populations, it is useful to have search and analytical functions that allow summaries to examine combinations of diagnosis and test outcomes. This implies the ability to acquire both positive and negative results from all cases meeting a case definition and/or clinical syndrome. Given that potential pathogens are common in animal populations, it is often useful to determine if disease is occurring from single agents or as disease complexes. The permutations of multiple Dx codes, with consideration of the sequence in which those agents occur over time, might be clinically significant and affect intervention strategies. Thus, a desirable feature for data analysis for diseases A, B, and C is tracking the presence and the age of onset of the various permutations in which these diseases occur in different populations (e.g., disease A and B and C, A and C not B, C and B not A, etc.). Because of the idiosyncrasies of the previous system, which included some colloquial or outdated nomenclature, and inconsistencies in the number of Dx codes applied, these types of inquiries were nearly impossible, even by seasoned users. An example of drift in the system is rumen metabolic acidosis. This was coded simply as “GRAI OVER” with a description of acidosis/grain overload-lactic acidosis. Codes in the previous system typically began with the process: neoplastic (N designation), toxic (TOXI or DEFI), or inflammatory (PNEU for pneumonia, ENTE for enteritis), for example, but there was not a metabolic category, therefore the lay term became the code. Another goal for a new coding system was to make search functions accessible to external users, with no coding experience, and the ability to easily feed the data into analytical tools (algorithms).
We developed a Dx coding system that allowed integration of historical Dx code data with a new, expanded, and more versatile coding system with the specific objectives to provide basic summary and trend information to stakeholders of the ISU-VDL. Concurrently, the development of algorithms and data visualization tools to make data analysis more user-friendly and convenient was implemented. (See Trevisan et al. in this issue for in-depth information on those tools.) This coding system was fully implemented and has been utilized for all pathology case coding as of 1 July 2019. All cases dating back to 1 January 2003 were recoded using the new system. Mapping old codes to new codes necessarily resulted in some compromise in code specificity and some aberrations in trends for a few diseases, but most of the major and common diseases diagnosed at the ISU-VDL mapped directly. We do have a parallel search system for the sample-analyte-test-result that can assist us in the interpretation of step changes in Dx codes as well as trends in detection frequency or rates.
ISU-VDL diagnostic coding process
Briefly, the diagnostic process at the ISU-VDL occurs as follows (Fig. 1): a submitter provides samples along with clinical and gross lesion information, with their diagnostic question(s) to the laboratory (dataset 1). Using the clinical data provided by the submitting veterinarian, the diagnostic pathologist reviews tissue changes and selects testing commonly consisting of histopathology, bacteriology, and molecular testing (PCR) to address differential diagnoses for the reported clinical syndrome, clinical history, and tissue changes (dataset 2). Through the aggregation of dataset 1 (clinical data) and dataset 2 (laboratory data), the diagnostic pathologist reaches a laboratory diagnosis or diagnoses that are then captured in the case report and coding system as a Dx code. This system moves beyond the test and report format in which output data are not necessarily reflective of disease status or impact. In animal groups in which agents are endemic and constantly circulating, this system can ultimately have a greater impact on the understanding of disease processes, epidemiology, efficacy of clinical intervention, or deviations in disease manifestation.
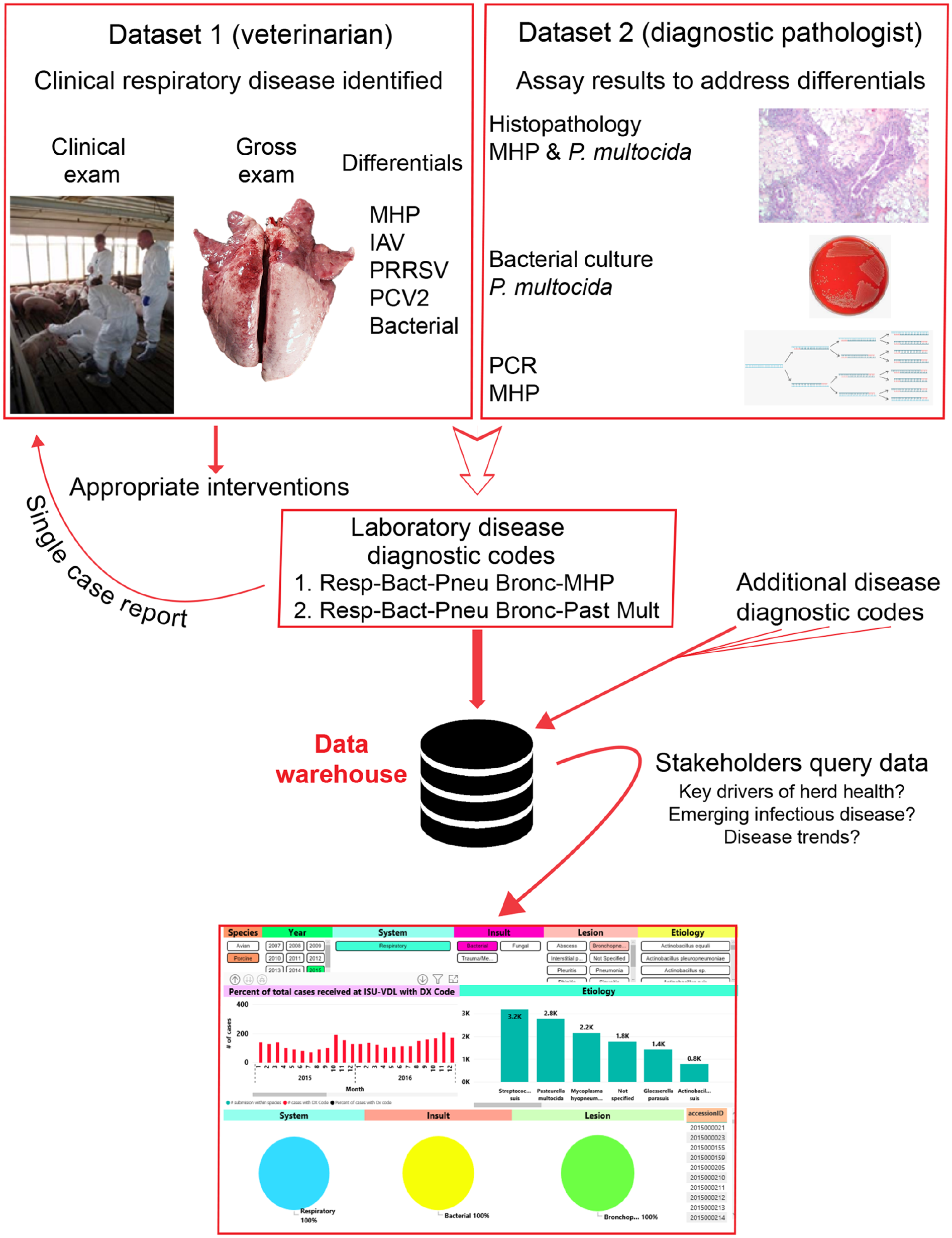
The ISU-VDL diagnostic process and relevant feedback loops. From the top, examples of datasets submitted by a veterinarian and generated by the laboratory are combined to create a case report with associated diagnostic codes. This single case report is utilized to inform appropriate interventions, but also is stored in the data warehouse, which can be queried to retrieve output data. IAV = influenza A virus; MHP = Mycoplasma hyopneumoniae; P. multocida = Pasteurella multocida; PCV2 = porcine circovirus 2; PRRSV = porcine reproductive and respiratory syndrome virus.
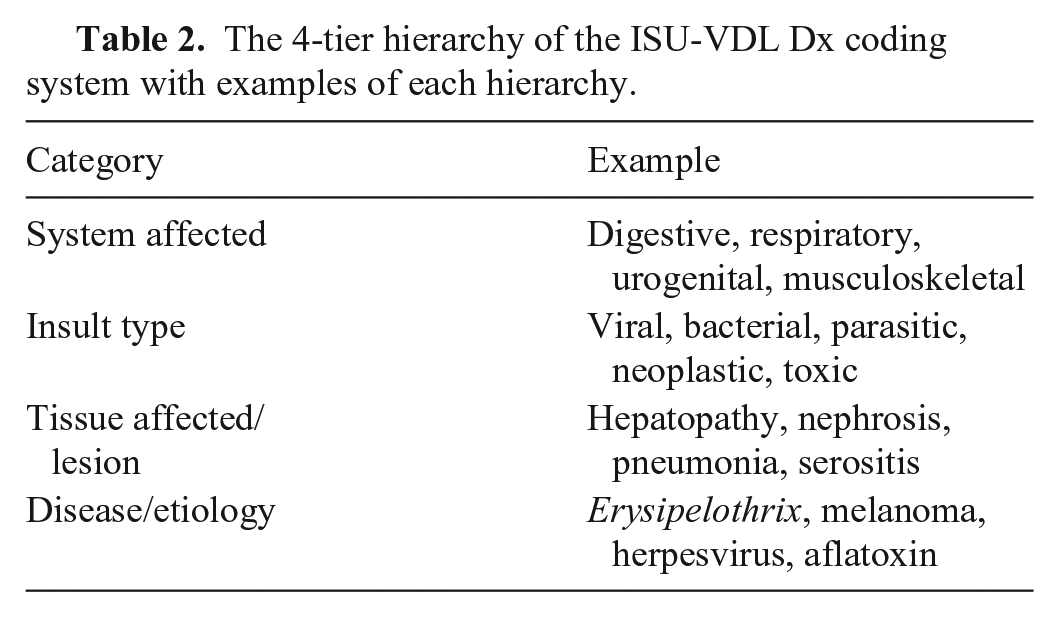
Within the system, all cases assigned a test within autopsy or histology sections require a Dx code to close the case. Etiology or causation is only included in the code when sufficient evidence is available to support that conclusion. Any additional case in which there is sufficient data to establish a disease status or observed clinical syndrome can be coded. Specifically excluded from Dx code assignment are ISU-VDL cases with only test-analyte-report systems in place. Available Dx codes are automatically filtered by the animal species represented in the case to help reduce potential coding errors. Then, one or more relevant Dx codes are selected by the diagnostic pathologist coordinating the case. Each Dx code has 4 components (system affected, insult type, tissue affected/lesion, and disease/etiology; Table 2) that can be entered in any order, with “type-ahead” filtering and automatic selection tools augmenting the process.
The 4-tier hierarchy of the ISU-VDL Dx coding system with examples of each hierarchy.

Assignment of a disease diagnosis (Dx code) usually requires 2 or more lines of evidence. This can be straightforward, such as with typical clinical signs and confirmatory test (e.g., toxicities, metabolic diseases, parasitism), or by combinations of evidence types that include lesions (i.e., gross and/or microscopic pathology) with ancillary information (e.g., clinical signs, test results; Fig. 1). To arrive at a laboratory diagnosis, a typical or classic lesion might be sufficient, or a compatible lesion coupled with an ancillary or confirmatory test.
The consistency of Dx codes across diagnostic pathologists is improved when Dx code entry is intuitive, flexible, rapid, and easily understood without requiring extensive training or institutional knowledge. In the ISU-VDL system, the “type-ahead” filter for Dx codes allows rapid visualization and entry of multiple codes simultaneously. Additionally, by narrowing available Dx code options as users type, the system helps guide users to similar coding endpoints regardless of the starting Dx code component that is entered into the search tool. Users may start with the etiology and work backward to find the proper code or may begin with insult and lesion terms to find a list of potential etiologies available in the system (Fig. 2).
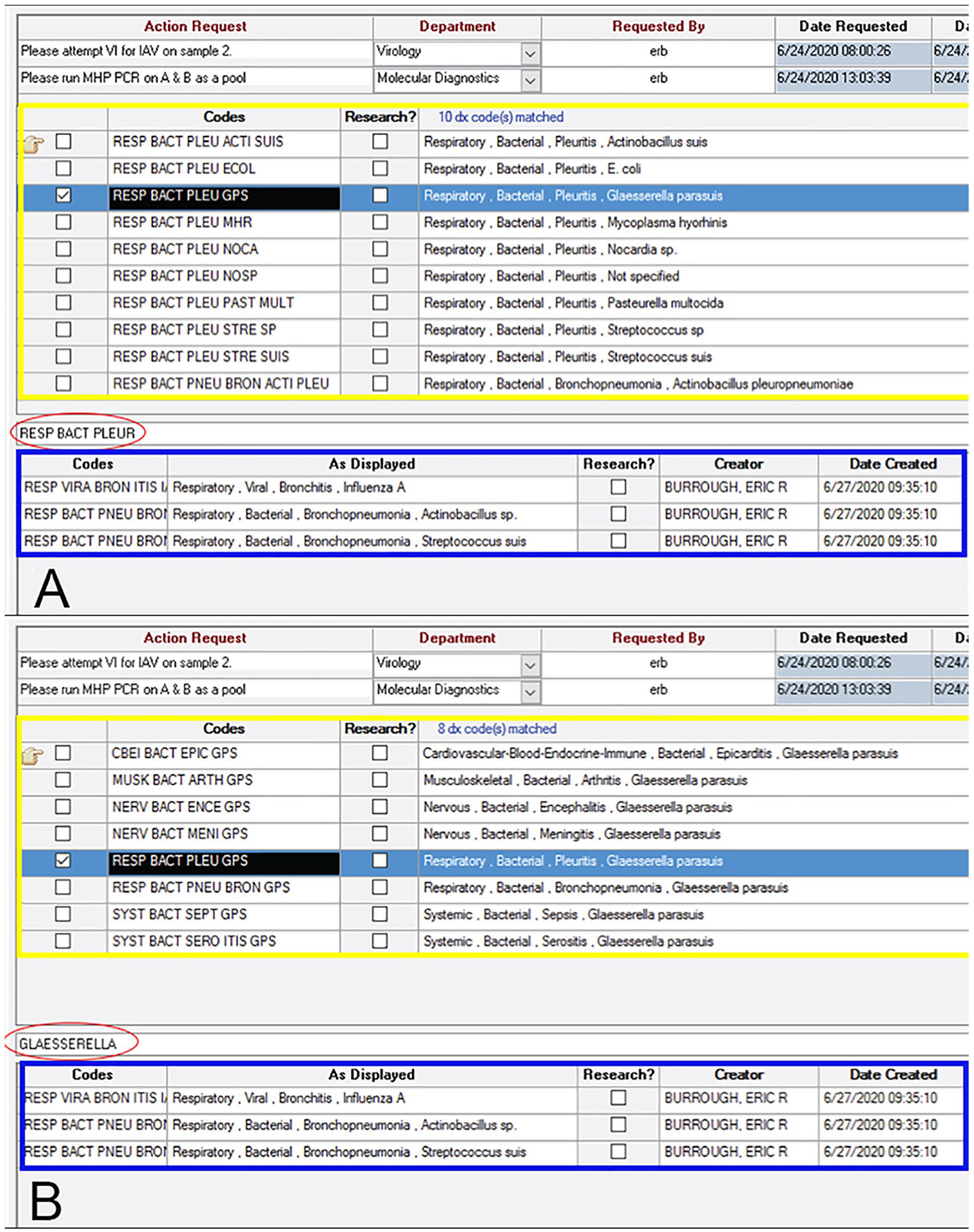
Examples of diagnostic (Dx) code entry within the LIMS at the ISU-VDL. In this case, Dx codes for influenza A virus infection and bronchopneumonia associated with Streptococcus suis and Actinobacillus sp. have already been applied (outlined in blue) and the diagnostician is applying a code for pleuritis associated with Glaesserella parasuis infection.
At each level, except system affected, the option “not specified” can be selected. For example, if a lesion appears to be bacterial but no specific bacteria are identified, the etiology code is “NOSP.” If criteria for specific etiologic or disease diagnosis are not met, cases are then coded by the body system reportedly affected in the submission history. For example, if coughing is indicated on the submission form, but no lesions are observed in the lung and no definitive pathogens are detected, the case is given a clinical diagnosis designation, and the case remains part of the disease database. In our system, the Dx code would be RESP NOSP CLIN DX NOSP (respiratory, not specified, clinical diagnosis, not specified) and would still be retrieved with a search by system-respiratory, given that this was the reason for case submission. Dx codes were migrated to the new system from the former system, with multiple reviews by pathologists to amend, update, add, or omit codes, as necessary. There is also a designation of research, providing an easy method to separate research cases from diagnostic cases when compiling disease data related to clinical submissions.
Disease states in both individual animals and groups may not have an etiology identified, but if coded by system, insult type, and lesion or tissue involved, can still be sought and found in our system. This feature is useful for many reasons, including identifying or tracking new or emerging lesions or disease and identifying trends to better define them. In the previous system, cases with lesions but without an identified etiology would be difficult to find. An example of a diagnosis without an etiology would be a lesion code of bronchointerstitial pneumonia. In the new system, this is easier to find, given that the specific term bronchointerstitial pneumonia does not need to be known, but can be found searching in “respiratory” and “not specified.” Our system allows for common names to be useful to clients, syndrome names for diseases of unconfirmed etiology, disease names as well as etiology/causation components. The database and entry/search platform make information available and understandable to a variety of potential users.
There are multiple stakeholder perspectives to consider in the development of a system, as well as questions of confidentiality, data accessibility, and ownership. The ideal system allows flexibility to find granular information yet allow organization and summarization over time. The vision for this system is to integrate Dx codes with all tests and results from the LIMS by using Dx codes to find relevant cases and using those accession numbers to return all test-result and anonymized demographic data from the LIMS. Importantly, the development of this coding system was designed specifically to include value to external clients. In particular, the desire of swine clients to have better analytical tools to support intervention decisions and monitor relevant diagnostic information for the health of large populations was considered; user-friendly tools to access, search, and analyze this large database are a priority. 7 Currently, 2 swine production systems are beta-testing this system and finding it valuable to identify their most frequently diagnosed diseases and benchmark frequency to the aggregate of the database, thus identifying potential areas of improvement. The system also provides insight to these systems of diseases not present, or perhaps overlooked, compared to the aggregate of their cohorts. The frequency and trends in identification and characterization of endemic potential pathogens become easier to identify and act upon. Production systems are linking disease diagnostic data to productivity and economic data to better understand disease agent impact and investigating the utility of diagnostic monitoring as part of the key indicators for productivity or lack thereof. Swine veterinarians can also use these data to support improved interventions for disease control to skeptical owners, nutritionists, and managers. Importantly, VDL external users are keenly interested in tools to identify step changes in disease occurrences. Step changes are abrupt or significant changes of frequency, location, or season such as can occur when a new agent enters a susceptible animal flow or a reporting system changes. These are more abrupt than fluctuation in trends.
Established systems such as VeNom or SNOMED-CT are specifically designed for academic or regulatory use and are cumbersome for clinicians involved in population medicine in the field. SNOMED-CT was originally designed by the American College of Pathology and has extensive terminology, and although it establishes relationships between codes, 1 it is not fully hierarchical, nor is it intuitive. As an example, we performed a search for disease in swine caused by Actinobacillus pleuropneumoniae (APP) in the SNOMED-CT vet extension (https://vtsl.vetmed.vt.edu/). It was unclear which codes or coding would be sufficient to code a case of APP; a search of “pleuropneumoniae” brings up 17 hits, all related to APP, but none of them a disease diagnosis. As a disease (disorder, in the SNOMED-CT terminology), it is listed under “Porcine contagious pleuropneumonia”, which is not standard terminology in the industry. Additionally, similar to our previous system, each SNOMED-CT code came with a description or explanation, which is unnecessary in the new paradigm. In the new system, a search of “pleuropneumoniae” on a swine case brings up 2 options: RESP BACT PNEU BRON ACTI PLEU (respiratory, bacterial, bronchopneumonia, Actinobacillus pleuropneumoniae) or SYST BACT SEPT ACTI PLEU (systemic, bacterial, septicemia, Actinobacillus pleuropneumoniae). Going back to the previous example of bloat, in the new system, this is DIGE TRAU RUME BLOA (digestive, trauma/mechanical, rumen, bloat). Our system adheres to basic findability, accessibility, interoperability, and reusability principles of database development,2,9 while maintaining simplicity, to the benefit of coders as well as database users.
The new Dx coding system at the ISU-VDL is sufficiently versatile to provide basic summary and trend information to submitters, to internal VDL users, and to allied stakeholders to ask better questions as well as allow for the aggregation of all relevant cases and their test results to answer high-level questions. Submitters benefit at the reporting level by receiving the diagnosis components in words that are consistent across multiple diagnosticians. As well, search and analysis are not only possible but simple, with visual tools to enhance analysis. Tools in beta testing are similar to those described and utilized in the companion article in this issue of JVDI by Trevisan et al. Our system allows for searching historical codes and cases to determine frequency of disease diagnosis, changes in frequency of diagnosis over time, and cases with multiple or concurrent diagnoses. There is also the capability to integrate Dx code with accession number and all other recorded laboratory data, such as demographic data on written or electronic submission forms, test-analyte-result testing, analytical capability for real-time summaries and historical analysis, and benchmarking at farm, system, regional, and national levels. For one of the beta testers, this allowed a better picture of the timing of onset of porcine reproductive and respiratory syndrome (PRRS) from a detection of PRRSV, as well as the manifestation of disease aspect. It also allowed the linking of sow farm data with disease diagnostic data to understand the impact of upstream factors in disease manifestation downstream. One of the 2 systems has moved to an automated model-building system, wherein close-out data and other on-farm data can be combined with the diagnostic data from ISU-VDL. This real-time data aggregation could become the gold standard in the industry. Although the algorithms and data aggregation tools are critical to link the Dx coding data, the simplicity and findability of the Dx coding system are foundational to making the data accessible. Currently, accessibility is limited to internal users and our beta testers. Our external users access the data through a password-protected link. We are continuously improving our system to assure relevance to stakeholders for trend analysis, anonymity for confidentiality concerns, and a concurrent institutional emphasis on accurate and comprehensive coding. A web-based feature is under development for submitters to offer feedback on utility and accuracy of diagnostic data provided by the laboratory.
Dx coding agreement
As discussed, one of the objectives of the Dx code system was to allow for accurate and consistent generation of Dx codes by diagnosticians, thus resulting in reliable data for stakeholders. Assessment of diagnostician Dx code agreement was performed by evaluation of 40 previously coded cases by 4 diagnosticians from a 10-mo period (1 July 2019–30 April 2020). Cases were limited to porcine or bovine species but were required to have histopathology, bacteriology, and molecular testing, and could have toxicologic testing. Eligible cases were randomly selected from the participating diagnosticians’ case logs with redaction of laboratory diagnoses and names of diagnosticians from the final report by an independent diagnostician.
Most cases (26 of 40) were designated as tier 1 cases given that they had 1 or 2 Dx codes for each case. There was general Dx code concordance on 26 of 27 (96%) cases; all 4 diagnosticians selected 1 or 2 Dx codes with the same system and etiologic agent. Tier 2 cases were defined as cases with ≥ 3 Dx codes assigned. Dx code agreement among all 4 diagnosticians on tier 2 cases was more varied (9 of 14). Discordance was driven by a mix of differences in diagnosis as well as in coding. For example, in one case, all diagnosticians agreed on the lesion but there was discord over whether there was enough evidence to add an etiologic agent to the lesion. Consensus was challenged in other cases because several systems were affected by different pathogens along with several secondary bacterial agents, resulting in numerous Dx codes by each diagnostician. Before our study, we were aware that differences in training and experience between diagnosticians may influence their selection of Dx codes; however, these few examples of discordance highlighted the necessity of collective training in proper Dx selection, given that there had been limited formal training of diagnosticians in appropriate Dx coding. In addition, discussion and ongoing training within our diagnostic pathology group are expected to provide clarity on tier 2 cases and to increase overall concordance in these cases.
From the presented investigation, as well as increased discussion of coding when reviewing cases as part of our laboratory’s monthly proficiency training for diagnosticians, we are improving our coding system by increasing consistency of data. Although each case at the ISU-VDL is coded by a single diagnostician, previous studies have shown that increasing the number of coders per case does not have significant advantages over a single-coder system. 4 Accordingly, as we continue to refine our coding system, we are confident in the accuracy of aggregated disease diagnosis information, particularly at the body system, insult, and etiology levels. For many studies examining agreement of coding or pathology grading systems, cases are narrowed to a disease entity (pathogen or neoplasia type),3,6,8 a system or body area (e.g., amputated digits or liver),5,10 and typically species (all-human reviews). Additional evaluation of our coding system, utilizing some of these limits (species, pathogen, system), will help to refine the system further or identify areas for additional training and discussion to reach better agreement among pathologists.
Implementation of the new Dx coding system and execution of this small study has generated valuable discussion, in hallways, offices, and particularly in our monthly reviews of diagnostician competency. Planned additional and more extensive analysis will identify discordances and potential pitfalls of the coding system, which can foster improvement toward a more consistent and structured framework for the coding process for our diagnosticians.
Future applications of Dx coding
We have developed a system that allows aggregation and visualization of Dx codes across populations and within systems. Dx coding moves beyond agent detection and associates an agent or abnormality with disease and represents an important feedback loop in the diagnostic process. To further leverage Dx codes, feedback loops with interactive capabilities between clinicians and diagnosticians are needed. A system in which near-real-time feedback could be provided to the clinician by the diagnostician in terms of case quality (diagnostic question posed, samples submitted, and confidence of the diagnostician in the diagnosis) as well as to the diagnostician by the clinician in terms of probable accuracy and applicability of laboratory diagnosis to the herd would facilitate continuous improvement and accuracy of the diagnostic process (Fig. 1).
Additionally, the creation of an interface that allows the collection of existing data from clinicians, including herd metadata (animal source, parity distribution, vaccination protocol), final diagnosis, interventions, or treatments applied, and measures of outcomes would provide data for evidence-based medicine in the form of effective disease intervention and prevention strategies. We have linked our coding system with a web-based, password-protected interface and are currently beta testing it. In production animal systems, this information can be organized and applied via statistical process control tools at the site, flow, or system level. Prototypes for stakeholder cooperation, collaboration, and beta testing of this vision are in development and will further inform Dx coding at the ISU-VDL. Specifically, stakeholders are currently evaluating by site (farm) the relationships between Dx codes, days on feed at the time of diagnosis, and close-out data including mortality, treatment cost, and average daily gain. Automation of these analyses on a predefined schedule would provide highly customizable applications to identify and prioritize preeminent, ultimate, and proximate causes of disease to improve animal health.
The ability for stakeholders at all levels to utilize accurate denominators to inform decisions is an important component of future use and process improvement. Panel PCR testing to ensure that differential diagnoses are assayed concurrently is one method by which to establish denominators. However, the ability to overlay agent detection results outside (surveillance and/or monitoring) and within cases with a Dx code that includes test, sample type, test result, and quantitative value (e.g., Ct, S/P ratio) may also facilitate improved understanding of epidemiology, infection dynamics, and microbial ecology within a population. Currently, our system does not allow for such analyses, but the infrastructure of the coding system should allow for these applications.
Conclusion
Dx coding is not a new concept, having been used in both human and veterinary medicine prior to moving to computer databases. To maximize utility, Dx coding schemes must be designed with a purpose—an understanding of how they will be used and by whom. Ideally, the system would be adaptable, searchable, and easy to understand. We propose that our system is all of these things and designed for use not only by institutional and regulatory users but also by clinical veterinarians, allowing the massive amounts of data produced to be aggregated to better drive evidence-based medicine and provide the portal to aggregate test results of individual cases for deeper analysis. Although our system was designed with the idea of querying production animal data, the flexibility of the system allows applicability to disease in other species as well. Given the tiered system, it should be more straightforward to query as well as provide more data, even in cases in which a specific etiology or process is not identified.
Looking forward, should a similar 4-tier Dx coding system be adopted at other VDLs, visualization of aggregated anonymized Dx code trends across VDLs becomes possible, similar to what has been done for PRRSV detection. 7 Other coding systems may be considered as well for wider use across VDLs. Discussion of what is in use, why, and the strengths and weaknesses of these systems as well as the goals when developing them should guide the veterinary diagnostic industry. Ultimately, such inter-VDL data aggregation of disease diagnoses would provide a platform to monitor disease trends for all species in a geographic region that is less susceptible to the inherent biases of a single laboratory reporting their own data. The tools are already available to make such analyses possible, which would further support evidence-based veterinary medicine. The next step is for VDLs to decide if it is worth the investment of time and resources to make a standardized veterinary disease coding system a reality to facilitate continuous improvement of animal health and welfare.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
