Abstract
Poor reproductive performance in beef cattle caused by infectious agents results in major financial losses as a result of reduced pregnancy rates and extended calving intervals. Bulls can be subclinical chronic carriers of bacterial and protozoal agents involved in cow infertility, such as Campylobacter fetus subsp. venerealis, Ureaplasma diversum, Mycoplasma bovigenitalium, Mycoplasma bovis, and Tritrichomonas foetus. Bulls harbor these microorganisms in their preputial crypts and transmit the agents to cows during natural mating. To obtain an overview of the etiologic agents in the preputial mucus of bulls, we aimed to identify, by PCR assay, C. fetus subsp. venerealis, M. bovis, U. diversum, M. bovigenitalium, and T. foetus in Brazilian bulls from farms with high infertility rates. We collected preputial mucus from 210 bulls on 18 beef cattle farms in Brazil between 2019 and 2020. We found at least one of the infectious agents that we were studying in bulls on 16 of the 18 beef cattle farms tested. We detected at least one infectious agent from 159 of 210 (76%) bulls tested, namely C. fetus subsp. venerealis, M. bovis, U. diversum, M. bovigenitalium, and T. foetus in 87 (55%), 84 (53%), 45 (28%), 28 (18%), and 1 (0.6%) animal, respectively. We found 95 bulls (60%) positive for only 1 etiologic agent (single infection) and 64 bulls (40%) carried multiple agents. Our results demonstrate the occurrence of bacterial and protozoal infectious agents that may be related to infertility in Brazilian beef cattle herds.
Brazil is the second-largest beef producer in the world and the largest exporter, exporting 2 million tons in bovine carcass weight each year; a trade worth >$5.4 billion per year. 41 For the beef industry worldwide, reproductive infections are a serious economic and health problem. The poor reproductive performance caused by infectious agents results in major financial losses as a result of reduced pregnancy rates and extended calving intervals. 25 In many countries, including Brazil, there are no official surveys of etiologic agents involved in poor reproductive performance; consequently, the economic losses are not quantified. 3 Taken together, these factors, and the lack of testing, result in lack of understanding of the effects of reproductive losses by infectious agents in the beef cattle industry. 22
Bovine reproductive diseases, especially infertility, are related mainly to bacterial, viral, and protozoal infections.5,38 Agents can be present in and infect the reproductive tract, leading to consequences either for the cow or the embryo.5,27 The main described agents related to infertility in cows are the bacteria Campylobacter fetus subsp. venerealis, Ureaplasma diversum, Mycoplasma bovigenitalium, and Mycoplasma bovis,2,5,22,28 and the protozoan Tritrichomonas foetus. 6 In addition, the viruses bovine viral diarrhea virus (BVDV; Flaviviridae, Pestivirus A) and bovine herpesvirus 1 (BoHV1; Herpesviridae, Varicellovirus, Bovine alphaherpesvirus 1) have an important role in bovine reproductive losses. 38
Bulls can be subclinical chronic carriers of the bacterial and protozoal agents involved in the infertility of cows, harboring the pathogens in their preputial crypts, which enables transmission of agents to cows by natural breeding or via stored semen.10,15,22,27 Therefore, epidemiologic surveillance and disease control are based, primarily, on identifying bulls carrying the infectious agents. In cows, the agents commonly colonize the genital mucosa, mainly in the vagina and uterus, initiating inflammatory reactions that result in an earlier return to estrus, infertility, and sporadic abortion.5,28
To obtain an overview of the main bacterial and protozoal etiologic agents that could be carried by bulls and cause possible poor reproductive performance in beef cattle, we aimed to identify C. fetus subsp. venerealis, M. bovis, U. diversum, M. bovigenitalium, and T. foetus in the preputial mucus of bulls from farms with high infertility rates.
Materials and methods
Selection of samples and DNA extraction
We invited farms with a history of poor reproductive performance, specifically high rates of reduced pregnancy and extended calving intervals (as determined by the staff of each farm), that submitted samples for diagnostic purposes of genital campylobacteriosis to our laboratory, to consent to the search for additional infectious agents in their bulls. The recruitment of convenience samples was during the winter of 2019–2020. The sampled bulls were randomly selected by either the farmer or the farm staff. Thus, we included 210 bulls from 18 beef cattle farms (A–R) in our study. The included farms were from the 2 largest beef cattle production regions in Brazil, the South Region (Rio Grande do Sul [RS] state) and the Midwest Region (Mato Grosso [MT] and Mato Grosso do Sul [MS] states).
Samples were collected after bulls had been sexually rested for at least 7 d. During the collection of biological material, the bulls were restrained carefully in a squeeze chute, and the preputial ostium was shaved, cleaned with water, and dried with paper towels. A soft brush coupled to a pipette was inserted into the animal’s prepuce, and mucus was collected by scraping the preputial and penile mucosa, as described previously. 9 After collection, the mucus and brush were placed in 1.5-mL tubes with 1 mL of buffered PBS, pH 7.0. The samples were stored in a cool box with ice and sent to the laboratory within 48 h. Samples collected in farms far away from the laboratory were frozen at −20°C for up to 10 d, and then sent to the laboratory as standardized previously by our group. 9 DNA extraction was performed directly from the collected mucus (Biopur mini spin plus; Biometrix Diagnóstica), according to the manufacturer’s instructions. DNA was stored at −20°C until use.
Molecular detection of infectious agents in preputial mucus samples
Each extracted DNA sample was tested by singleplex PCR for C. fetus subsp. venerealis and T. foetus, and by multiplex PCR for M. bovis, U. diversum, and M. bovigenitalium, with PCR reactions that had been validated previously (Table 1). C. fetus subsp. venerealis ATCC 19438, and in-house isolates of U. diversum, M. bovigenitalium, M. bovis, and T. foetus, were used as positive controls for the respective PCR reactions. To provide confirmation of assay specificity, the PCR amplicons from each positive control were sequenced by the Sanger method, and further assembled using Geneious v.10.2.3 (Biomatters) and analyzed in the NCBI platform (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Primers used in PCR assays for detection of infectious agents in samples of preputial mucus from Brazilian bulls from herds with a history of poor reproductive performance, 2019–2020.
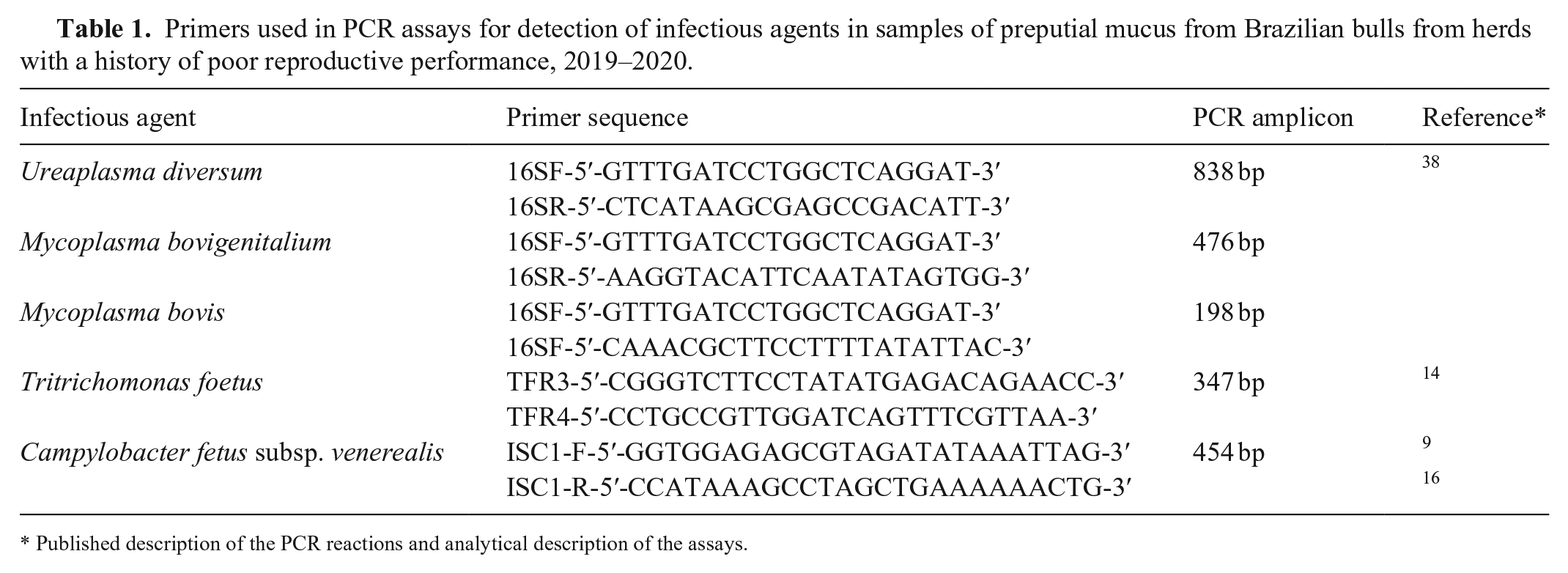

Published description of the PCR reactions and analytical description of the assays.
The singleplex PCRs carried out for C. fetus subsp. venerealis and T. foetus each contained 0.5 U of Taq DNA polymerase (Quatro G P&D), 1× reaction buffer, 1.25 μL (1.5 mM) of MgCl2, 0.5 µL (0.4 µM) of each primer, 2.5 µL (0.2 mM) of dNTPs, and 10 ng of template DNA. Conditions for amplification of C. fetus subsp. venerealis included initial denaturation at 95°C for 3 min, followed by 35 cycles of 95°C for 20 s, annealing at 52°C for 20 s, and extension at 72°C for 2 min, with a final extension of 72°C for 10 min. Conditions for T. foetus amplification were: initial denaturation at 94°C for 5 min, followed by 40 cycles of 94°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 90 s, with a final extension at 72°C for 5 min.
The multiplex PCR assays used to detect U. diversum, M. bovigenitalium, and M. bovis contained 1 U of Taq DNA polymerase (Quatro G P&D), 1× reaction buffer, 1.25 μL (1.5 mM) of MgCl2, 0.5 µL (0.4 µM) of each primer, 2.5 µL (0.2 mM) of dNTPs, and 10 ng of template DNA. The optimal PCR conditions were: initial denaturation at 95°C for 15 min, followed by 40 cycles of 95°C for 1 min, 52°C for 1 min, and 72°C for 1 min, with a final extension at 72°C for 10 min.
All PCR assays were carried out in a thermocycler (MiniAmp thermal cycler; Applied Biosystems). Subsequently, the amplification confirmation was visualized by electrophoresis on 1% agarose gels stained with UniSafe dye (Uniscience).
Statistical analyses
The PCR results were compiled and analyzed (SPSS v.26.0; IBM). Possible associations among etiologic agents (coinfection) were assessed by Pearson chi-square and Fisher exact tests. All estimates were 2-tailed with a pre-established significance level for the alpha error of 5% (p ≤ 0.05). The 95% CIs were calculated for the percentages of the infectious agents detected in bulls and farms.
Results
Molecular detection of 5 reproductive infectious agents
The generated sequences from positive controls were submitted to a BLASTn search and showed identity of 99.3–100% with the sequences available in GenBank. The nucleotide sequences were deposited in GenBank as accessions MW725075, MW725076, MW725077, and MW726033, for M. bovis, M. bovigenitalium, U. diversum, and T. foetus, respectively. From the 18 Brazilian cattle farms tested, 16 had at least 1 of the etiologic agents analyzed (Table 2). Analyzing the distribution of the microorganisms in the 16 positive farms, C. fetus subsp. venerealis was present in 14 farms (78%; 95% CI: 58–98%), followed by M. bovis in 13 (72%; 95% CI: 51–94%), U. diversum in 11 (61%; 95% CI: 38–84%), M. bovigenitalium in 8 (44%; 95% CI: 21–68%), and finally, T. foetus in 1 farm (5%; 95% CI: –5–16%). No agents were identified in 2 of the analyzed farms (B and H) from the South Region (Table 2).
Molecular detection of 5 reproductive etiologic agents in preputial mucus samples of bulls from 18 Brazilian cattle herds with poor reproductive performance between 2019 and 2020.
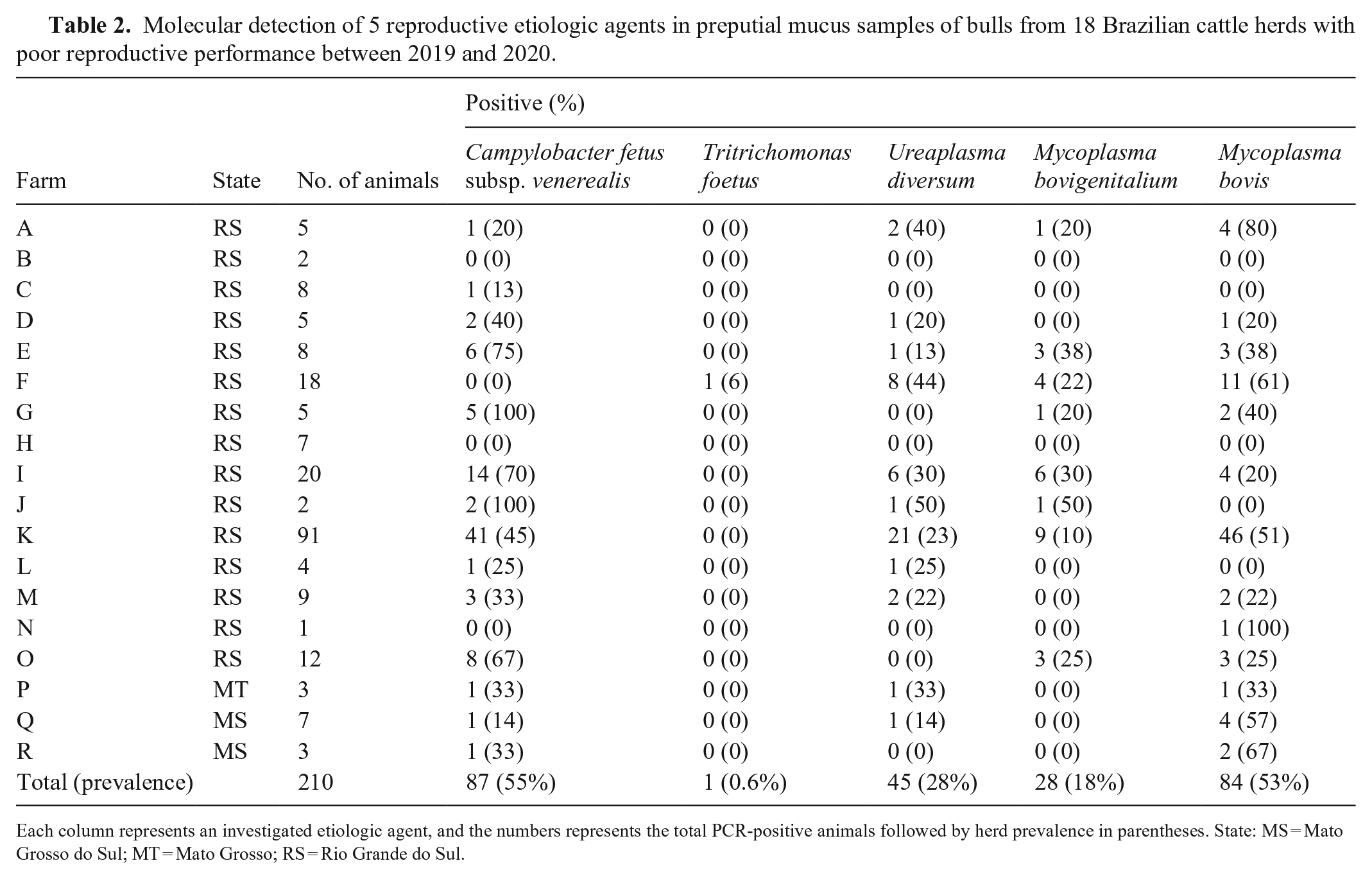
Each column represents an investigated etiologic agent, and the numbers represents the total PCR-positive animals followed by herd prevalence in parentheses. State: MS = Mato Grosso do Sul; MT = Mato Grosso; RS = Rio Grande do Sul.
PCR assay results showed that 159 of 210 bulls (76%; 95% CI: 70–81%) had at least 1 infectious agent identified in the preputial mucus, including C. fetus subsp. venerealis, M. bovis, U. diversum, and M. bovigenitalium, in 87 (55%; 95% CI: 47–63%), 84 (53%; 95% CI: 45–61%), 45 (28%; 95% CI: 21–35%), and 28 (17%; 95% CI: 12–24%) of 159 bulls, respectively (Table 2).
Codetection of agents in the preputial mucosa of bulls
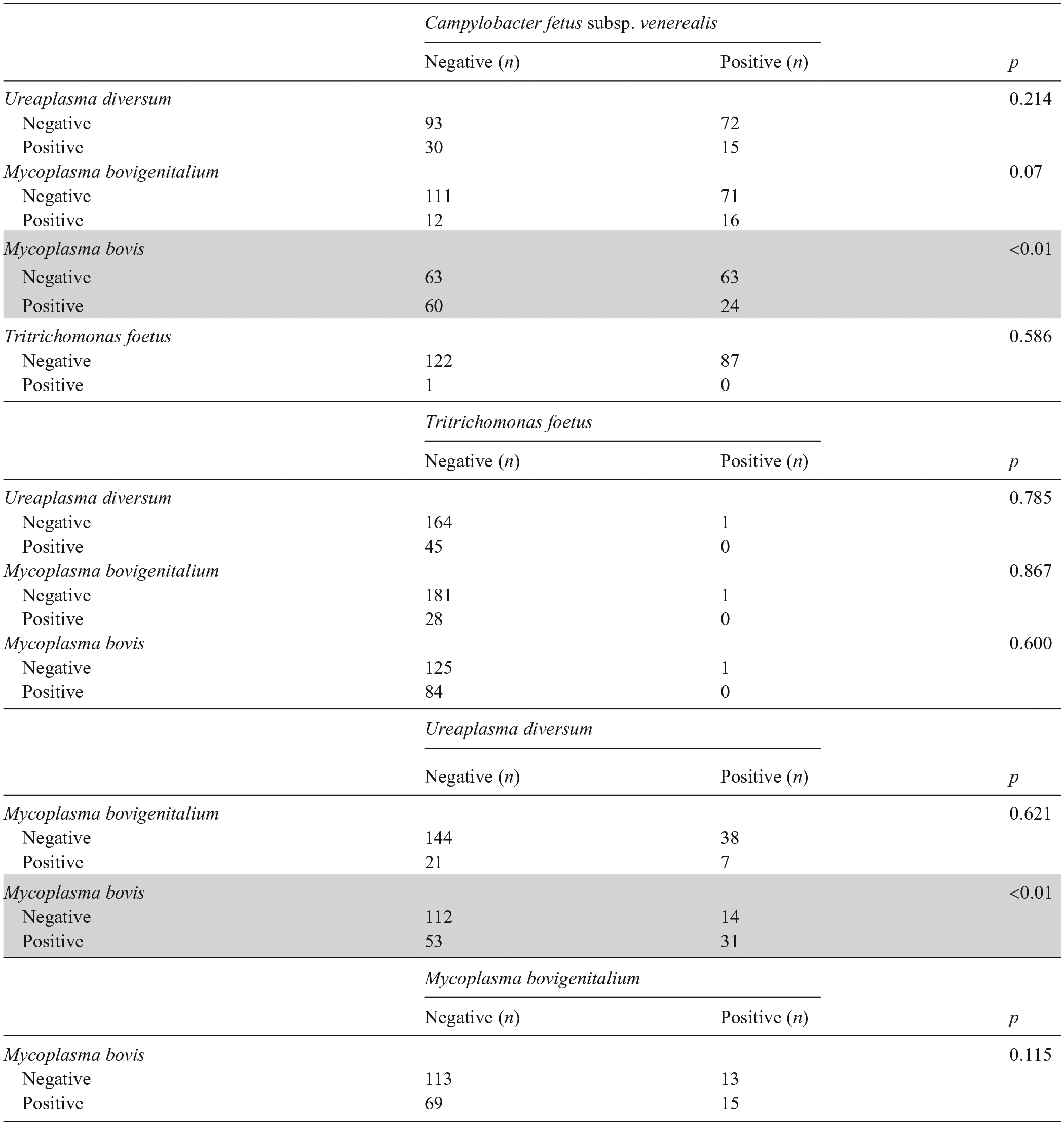
We found that 95 of 210 bulls (60%; 95% CI: 52–67%) were positive for only 1 etiologic agent (single infection); 64 bulls (40%; 95% CI: 33–48%) carried multiple agents. Among these codetection cases, 42 of 64 bulls (66%; 95% CI: 54–78%) harbored 2 of the screened agents; 22 bulls (34%; 95% CI: 23–46%) carried 3 of the screened agents.
From the 87 bulls positive for C. fetus subsp. venerealis, 55 bulls (63%; 95% CI: 53–73%) had codetections (Table 3). In addition to C. fetus subsp. venerealis infection, 24 bulls (28%; 95% CI: 18–37%) were also positive for M. bovis, 16 bulls (18%; 95% CI: 10–27%) were positive for M. bovigenitalium, and 15 bulls (17%; 95% CI: 9–25%) were positive for U. diversum. Moreover, a statistically significant association was identified between C. fetus subsp. venerealis and M. bovis (p < 0.01; Table 3).
Codetections of reproductive etiologic agents identified by PCR in the preputial mucus of Brazilian bulls.
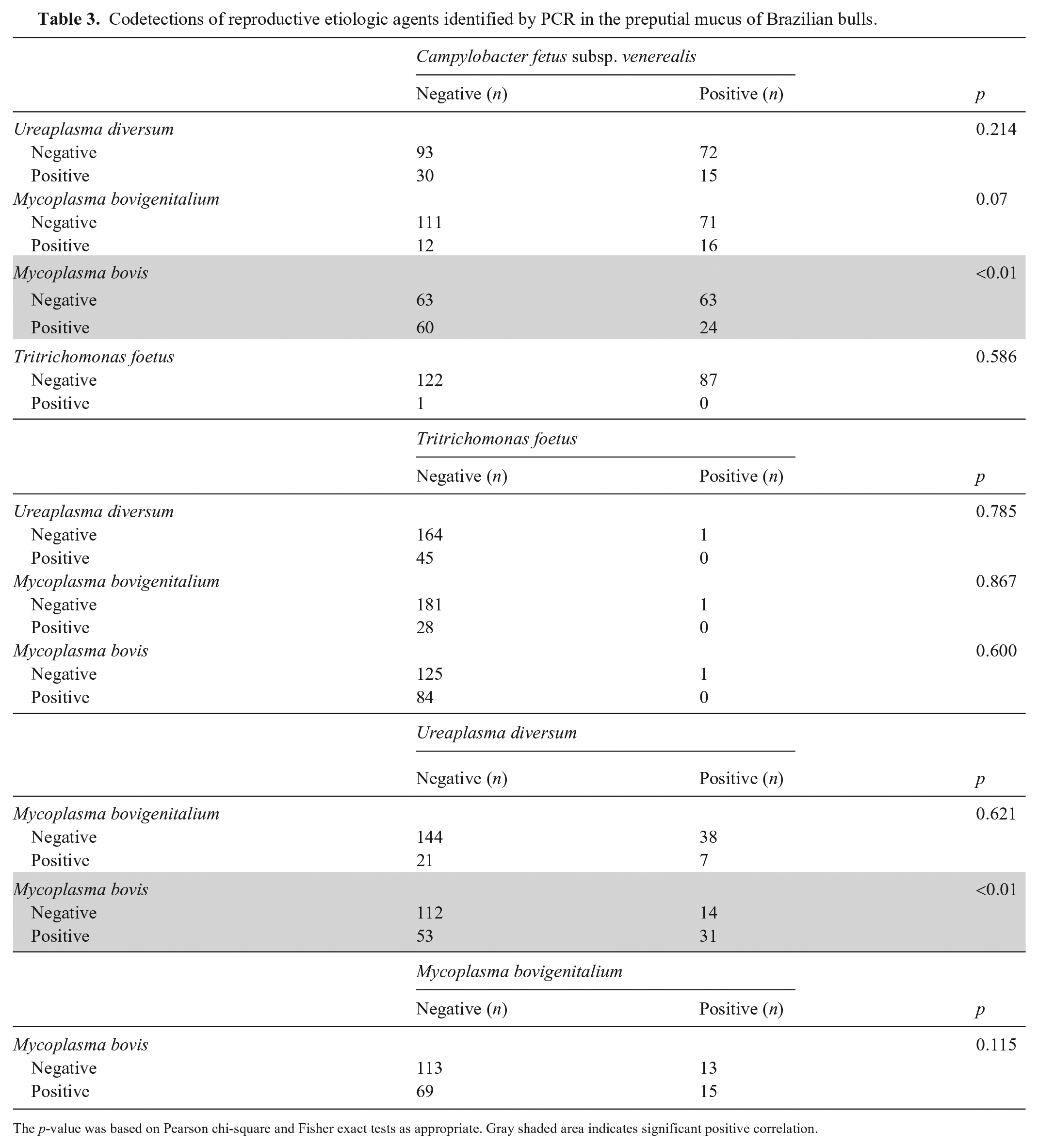
The p-value was based on Pearson chi-square and Fisher exact tests as appropriate. Gray shaded area indicates significant positive correlation.
M. bovis was identified in 84 bulls, with 14 (17%; 95% CI: 9–25%) harboring the bacterium as a single pathogen. Among the M. bovis–positive bulls, 31 (37%; 95% CI: 26–47%) also carried U. diversum, with a statistically significant association between the bacteria (p < 0.01; Table 3).
Discussion
Our analyses reveal a high frequency and diversity of reproductive bacterial and protozoal infectious agents circulating in Brazilian cattle herds with a history of poor reproductive performance. Altogether, 159 of the 210 bulls (76%) analyzed in our study were carriers of at least 1 of the agents investigated by PCR assay: C. fetus subsp. venerealis, M. bovis, M. bovigenitalium, T. foetus, and/or U. diversum. In addition, 16 of 18 farms were identified as having at least 1 of the investigated agents. Our study provides prevalence information that may influence the management of infectious agents carried by bulls.
C. fetus subsp. venerealis was the infectious agent identified most frequently in the bulls tested (55%). This bacterium is considered an important cause of reproductive loss in beef herds; however, previous studies have reported occurrences of 1.8–51.7% among Brazilian cattle. 36 The variability in the frequencies of identification might be related to the difficulty in detecting C. fetus subsp. venerealis by both culture and fluorescence methods,24,35 which were the most common approaches in the past. Molecular assays are now the most useful methods to identify C. fetus subsp. venerealis because they are independent of bacterial viability.9,16 Moreover, the frequency of C. fetus subsp. venerealis that we observed is higher than previous reports from herds with an infertility history in other South American countries, such as Uruguay and Argentina.13,23 In North American and Asian countries, C. fetus subsp. venerealis is rare among breeding bulls from herds with an infertility history, with frequencies of 1.1% and 8.8%, respectively.18,39 The described level of infected bulls in herds with low pregnancy rates was 28.8% in Oceania. 34 Despite the absence of reliable data from many countries, campylobacteriosis is known to be distributed worldwide. 25
M. bovis, M. bovigenitalium, and U. diversum are members of the Mycoplasmataceae family. Interestingly, although Mycoplasma spp. can produce energy by metabolizing carbohydrates and amino acids as an alternative to urea, Ureaplasma spp. rely strictly on urea for their growth. 20 An increase of the blood urea concentration has a relationship with the presence of U. diversum in the vagina of cows; the authors hypothesized that the urea could be a substrate for U. diversum growth in the vagina. 33 The significant association that we found between U. diversum and M. bovis (19.5% of bulls were carriers of both pathogens) in preputial crypts might be the result of a synergistic effect between these bacteria. We hypothesize that M. bovis would promote a favorable colonization environment for U. diversum in the prepuce of bulls, and we intend to study this hypothesis in future in vitro research.
Even though M. bovis has been demonstrated to colonize the reproductive tract of cows, venereal transmission of M. bovis has not been defined. 17 As well, the reproductive consequences of M. bovis and other Mycoplasma spp. are poorly studied in cattle. 8 We found that 84 bulls (43%) harbored M. bovis in preputial crypts. The need for more study of how and if M. bovis interacts with other pathogens should be highlighted, given that we observed this pathogen in association with C. fetus subsp. venerealis (co-harbored in 15% of the animals) and with U. diversum (co-harbored in 19.5% of the animals).
The occurrence of U. diversum in 21% of the animals tested is very similar to results of an Australian study, 1 in which 21.2% of bulls with a history of breeding problems (including genital lesions) were positive. On the other hand, the frequency of U. diversum in subclinical bulls was even larger in Argentina, with 11 of 17 bulls (65%) harboring the agent in their crypts. 37 Ureaplasma has also been described as inhabiting the reproductive tract of bulls in the United States. 40 To our knowledge, few reports have investigated the epidemiology of U. diversum in bulls; our study demonstrated the presence of U. diversum in Brazilian bulls from herds with poor reproductive performance.
We found that 18% of bulls were positive for M. bovigenitalium. The clinical signs of M. bovigenitalium infections can be observed in cows that are infected from subclinical bulls.7,32 Information on the distribution of M. bovigenitalium is scarce, although this bacterium is associated with abortions, lesions in cow reproductive tracts, including granulo-pustular vulvovaginitis, and infertility problems in cattle.32,38
Bovine trichomoniasis depends on carrier bulls transmitting the agent to cows, consequently causing low fertility rates in herds.14,29,31 According to our molecular screening, T. foetus was identified in only one bull (0.6%). Our results agree with the literature, which indicates a low prevalence of T. foetus in cattle in American countries.11,19,23,26,30 Prevalence studies associating molecular approaches with protozoan culture showed the absence of T. foetus in bulls from Tennessee as well as the Eastern Caribbean.11,26 A study using protozoan culture in Costa Rican bulls showed a prevalence of ~4%. 30 Prevalence of T. foetus in Switzerland and Spain, according to parallel analyses of culture and PCR reactions, was 0–32%.4,21 Reports from Spain demonstrated a re-emergence of trichomoniasis in bulls, of 13–32%.12,21 Interestingly, the recurrence of trichomoniasis occurred simultaneously with the return of natural mating as the predominant reproduction type in some places within Europe. 12
Our data illustrate the presence of C. fetus subsp. venerealis, M. bovis, M. bovigenitalium, T. foetus, and U. diversum in the prepuce of bulls from farms with poor reproductive performance, specifically infertility. The limitation of our convenience sampling in cattle herds with poor reproductive performance precludes conclusions about whether the infectious agents identified were directly responsible for reproductive losses, especially by the lack of information on pregnancy rates, calving intervals, and diagnostic surveillance of the studied herds, as well the absence of tests for other infectious agents that are potentially involved in reproductive diseases. In addition, the codetections identified should be interpreted with caution because, although we demonstrated a statistically significant association among bacteria, these data do not prove synergy.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was supported by the Coordination of Improvement of Higher Educational Personnel (CAPES)—Finance Code 001; Propesq/UFRGS (Process 001/2019); and the National Council of Technological and Scientific Development–CNPq (Process 420590/2018-8–CNPq-Universal/2018).