Abstract
We describe the molecular analysis of a wild-type field strain of bovine viral diarrhea virus (BVDV) identified in a mummified fetus from a small Brazilian dairy cattle herd. Nucleic acids extracted from samples of the lung, liver, heart, spleen, and kidney were tested by PCR assays for bovine alphaherpesvirus 1, Neospora caninum, Leptospira spp., Histophilus somni, and Brucella abortus, a nested PCR assay for Mycoplasma bovigenitalium and Ureaplasma diversum, and a RT-PCR assay for BVDV. Amplicons were only obtained in the RT-PCR assay for the partial amplification of the BVDV 5′UTR (288 bp) in kidney and spleen samples and the Npro (438 bp) gene in the kidney sample. Nucleotide sequencing of the amplified products and phylogenetic analyses based on the 2 BVDV genomic regions enabled the BVDV strain to be classified as subgenotype 1a.
Bovine viral diarrhea virus (BVDV) is a major pathogen that infects beef and dairy cattle worldwide. 11 Depending on the age, sex, and reproductive stage of the cattle, BVDV infection can be associated with a range of distinct clinical signs. In cows, BVDV infection can cause reproductive disorders, such as embryonic death, abortion, and fetal mummification if the infection occurs during the first trimester of gestation,3,7 although mummification may occur at 3–8 mo of gestation. 9 Mummification caused by BVDV infection in cattle was first described in 1973, and the authors employed an illustration to describe the occurrence of mummified fetuses after pregnant heifers were inoculated intravenously with 105 TCID50 of BVDV. 12 Potential causes of fetal mummification in cattle include infectious factors (bovine viral diarrhea, leptospirosis, neosporosis, molds), mechanical factors (compression or torsion of the umbilical cord, uterine torsion), and animal-specific factors (defective placentation, genetic anomalies, abnormal hormonal profiles, chromosomal abnormalities). 9
BVDV (Flaviviridae, Pestivirus) is classified as Pestivirus A (BVDV1), Pestivirus B (BVDV2), and Pestivirus H (HoBi-like pestivirus, or BVDV3). 8 Wild-type BVDV strains have been described worldwide, and at least 23 subgenotypes of BVDV1 (1a–1w) and 4 subgenotypes of BVDV2 (2a–2d) have been described.5,19
The detection of BVDV infection in aborted bovine fetuses has been described previously in Brazil.6,10 However, although this virus has been associated regularly with the occurrence of fetal mummification in cattle, only on rare occasions worldwide has this virus been identified in mummified fetuses.9,12 Most likely, this lower frequency of detection is attributable to the difficulty in obtaining mummified fetuses. Further possible reasons for the scarcity of these reports are that the virus may be present but noninfectious in the mummified fetus, and that the viral proteins and nucleic acids may degrade, particularly given that BVDV is an RNA virus, and RNA is more labile than DNA, which comprises the genomes of many other viruses, bacteria, and protozoa.
A few reports have described fetal mummification in cattle associated with BVDV infection, but to our knowledge, there are no studies describing the molecular characterization of the infecting viral field strain. We describe here the etiology of this phenomenon and our molecular analysis of the wild-type BVDV strain detected in a mummified bovine fetus from a Brazilian dairy cattle herd.
During a regular visit by the herd veterinarian for pregnancy diagnosis, an asymptomatic primiparous Holstein cow at 8-mo gestation spontaneously aborted a mummified fetus (Fig. 1). The fetus was ~45 cm long, which indicates that fetal death occurred at 6 mo gestation. The dairy cattle herd was located in the northern region of Paraná state, Brazil, and had 40 lactating cows with an average production of 19 L/cow/d. The Holstein and crossbred Holstein × Gir (Girolando) cows were maintained in a semi-confined system and fed corn silage, hay, crushed sugar cane, and pasture of mombaça grass (Panicum maximum; Brazilian: Quênia grass). The herd had a history of reproductive failure, including abortion, and aside from brucellosis vaccination, no other vaccinations were employed for the control and prevention of reproductive diseases.
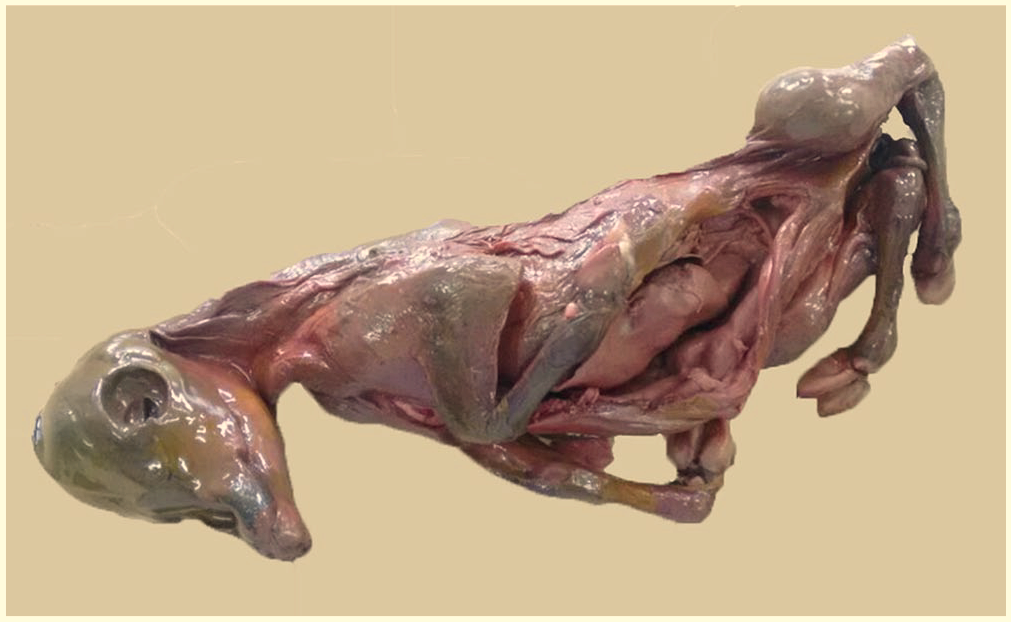
Mummified 6-mo gestation fetus from a Brazilian dairy cattle herd.
Given that fetal mummification is a relatively rare phenomenon in cattle, the veterinarian collected the mummified fetus for laboratory analysis. In the laboratory, samples of the lung, liver, heart, spleen, and kidney were collected and stored at −80°C until processed. The samples were mechanically disrupted, homogenized by vortexing with phosphate-buffered saline (10% w/v), and centrifuged at 1,500 × g for 5 min. Nucleic acids were extracted from 500-μL aliquots of organ supernatants, pretreated with sodium dodecyl sulfate (1% v/v) and proteinase K (0.2 mg/mL), and incubated at 56°C for 30 min. Nucleic acid extraction was performed using a combination of phenol-chloroform-isoamyl alcohol (25:24:1) and silica-guanidinium isothiocyanate extraction methods.1,2 Nucleic acid was eluted in 50 μL of ultrapure nuclease-free, diethylpyrocarbonate-treated sterile water and stored at −80°C until molecular analyses were performed. Ultrapure sterile water was used as a negative control.
The BVDV reverse-transcription PCR (RT-PCR) assay was performed using 324/326 primers to amplify a partial sequence in the 5′UTR BVDV genomic region. 15 To confirm the results of the phylogenetic analysis based on the 5′UTR fragment, BD1/BD3 primers were employed to amplify a fragment of the Npro region. 16 The cell culture (Madin–Darby bovine kidney, MDBK)-adapted BVDV Singer strain was used as positive control.
To rule out other infectious agents associated with abortion in cows, we performed PCR assays for bovine alphaherpesvirus 1 (BoHV1), Neospora caninum, Leptospira spp., Histophilus somni, Brucella abortus, as well as nested PCR for Mycoplasma bovigenitalium and Ureaplasma diversum (Suppl. Table 1). For positive controls, we used cell culture (MDBK)-adapted BoHV1 Los Angeles strain, as well DNA (kindly provided by the protozoology and bacteriology laboratories of Universidade Estadual de Londrina) extracted from N. caninum, L. interrogans serovar Hardjo, H. somni, B. abortus, M. bovigenitalium, and U. diversum samples. Test results of the mummified fetus for these agents were all negative.
The RT-PCR, PCR, and nested PCR products were analyzed by electrophoresis on 2% agarose gel in Tris-borate-EDTA buffer (89 mM Tris, 89 mM boric acid, 2 mM EDTA) at pH 8.4; the gel contained 0.5 μg/mL of ethidium bromide and was visualized under UV light. The BVDV RT-PCR products were purified (PureLink quick gel extraction and PCR purification combo kit; Invitrogen) and quantified (Qubit dsDNA BR assay kit, Qubit fluorometer quantitation; Invitrogen) as per the manufacturer’s recommendations. Direct sequencing was performed using the same forward and reverse primers used in the RT-PCR assays (BigDye Terminator v.3.1 cycle sequencing kit, 3500 genetic analyzer sequencer; Applied Biosystems). The quality of nucleotide (nt) sequences and consensus sequences were obtained using PHRED/CAP3 (http://asparagin.cenargen.embrapa.br/phph/), and similarity searches were performed with nt sequences deposited in GenBank using BLAST (https://blast.ncbi.nlm.nih.gov/). The phylogenetic trees were reconstructed using the neighbor-joining method with the Kimura 2-parameter model based on 1,000 bootstrapped datasets in MEGA v.7 (https://www.megasoftware.net/).
A 288-bp product from the 5′UTR region of the BVDV genome was detected in samples of the kidney and spleen. For the BVDV Npro gene, a 428-bp amplicon was obtained from the kidney sample. The RT-PCR products with the best quality, as determined through agarose gel electrophoresis, were selected for sequencing, and the gene sequences described here were deposited in GenBank as accessions MT786207 (5′UTR) and MT786208 (Npro).
The 5′UTR is the most conserved region of the BVDV genome and is the viral genomic region for which the most nt sequences are available in GenBank. Npro is a viral autoprotease, the first protein in the viral polyprotein, and the second most abundant sequence available in GenBank. 19 The quality of BVDV classification increases and the virus field strain is more strongly classified when it is performed in more than one region of the genome. 18
Several studies have described BVDV1a as the most prevalent subgenotype in Brazil. The BVDV strain found in the mummified bovine fetus described here was classified as BVDV subgenotype 1a (Suppl. Figs. 1, 2), which is consistent with other findings in Brazil. In one study, 26 of 38 (68%) BVDV1 isolates obtained from cattle were classified as subgenotype 1a. 13 Other studies in Brazil detected BVDV 1a in 8 of 11 (73%), 4 in 15 of 33 (45%), 17 and in 4 of 10 (40%) cases. 14 The preponderance of BVDV subgenotype 1a in Brazil is consistent with observations made in the United Kingdom, Ireland, and South Africa. 19
Supplemental Material
sj-pdf-1-jvd-10.1177_10406387211022012 – Supplemental material for Bovine viral diarrhea virus subgenotype 1a in a mummified fetus from a Brazilian dairy cattle herd
Supplemental material, sj-pdf-1-jvd-10.1177_10406387211022012 for Bovine viral diarrhea virus subgenotype 1a in a mummified fetus from a Brazilian dairy cattle herd by Juliana T. T. Fritzen, Arthur B. Morettin, Elis Lorenzetti, Alice F. Alfieri and Amauri A. Alfieri in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the following Brazilian Institutes for their financial support: National Council of Scientific and Technological Development (CNPq), the Brazilian Federal Agency for Support and Evaluation of Graduate Education (CAPES), and the Araucária Foundation (FAP/PR). A.A. Alfieri and A.F. Alfieri are recipients of CNPq fellowships. We thank the veterinarian, Dr. Idalino Sgobero Filho (deceased), for submitting the mummified fetus to our laboratory.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was funded by Instituto Nacional de Ciência e Tecnologia para a Cadeia Produtiva do Leite (grant 465725/2014–7).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
