Abstract
Detection of bovine Babesia spp. and Anaplasma marginale is based on the reading of Giemsa-stained blood or organ smears, which can have low sensitivity. Our aim was to improve the detection of bovine Babesia spp. and A. marginale by validating a multiplex PCR (mPCR). We used 466 samples of blood and/or organs of animals with signs and presumptive autopsy findings of babesiosis or anaplasmosis. The primers in our mPCR amplified the rap-1a gene region of Babesia bovis and B. bigemina, and the msp-5 region of A. marginale. We used a Bayesian model with a non-informative priori distribution for the prevalence estimate and informative priori distribution for estimation of sensitivity and specificity. The sensitivity and specificity for smear detection of Babesia spp. were 68.6% and 99.1%, and for A. marginale 85.6% and 98.8%, respectively. Sensitivity and specificity for mPCR detection for Babesia spp. were 94.2% and 97.1%, and for A. marginale 95.2% and 92.7%, respectively. Our mPCR had good accuracy in detecting Babesia spp. and A. marginale, and would be a reliable test for veterinarians to choose the correct treatment for each agent.
Introduction
The etiologic agents of bovine babesiosis and anaplasmosis in Uruguay are 2 protozoa, Babesia bovis and B. bigemina, and a rickettsia Anaplasma marginale, respectively. 36 These pathogens are intraerythrocytic parasites that can act alone or in combination. The main clinical signs of babesiosis and anaplasmosis are similar, and include fever, anemia, weakness, ataxia, hemoglobinuria, and jaundice, and infections can lead to death. 2 These diseases have high prevalence and cause large economic losses in livestock production in Uruguay and worldwide.4,30,31,36,37 Rhipicephalus microplus is the only competent vector for bovine babesiosis in South America. 21 Anaplasma can be transmitted by a wide range of hematophagous arthropods (horseflies, stable flies, mosquitoes, ticks), as well as iatrogenically.22,24
Bovine babesiosis and anaplasmosis can be detected indirectly or directly. The indirect methods evaluate antibodies generated from prior contact with the agents. The most commonly used techniques are ELISA, indirect immunofluorescence, and agglutination.19,29,40 These techniques are useful for gathering epidemiologic data such as the prevalence of these agents in a herd; however, they cannot be used for detection of acute disease. Direct methods are based either on visualization of the parasites within erythrocytes in smears stained with Giemsa, or DNA detection by PCR. 3
Smear reading is the traditional technique for detection of Babesia spp. and A. marginale. Parasites are found easily in bovine erythrocytes when animals are in the acute phase of infection (high parasitemia). 1 However, the smear technique has lower sensitivity in detecting carrier animals or animals in the early stages of the disease. Additionally, the stain used can generate artifacts, giving false-positive results, and requires trained personnel. 17 Although it is possible to distinguish between A. marginale and Babesia spp., it is not always possible to distinguish B. bovis from B. bigemina given their morphologic similarities. 11
To overcome these problems, molecular biology tools such as PCR can be useful, and have higher sensitivity and specificity. 9 There are PCR methods available that can identify the presence of both species of Babesia and A. marginale, even at very low parasitic loads.9,15,39 Although PCR assays designed to detect individual species are very sensitive, they are time-consuming and expensive when testing a large number of samples. Furthermore, the probability that some animals may be coinfected with more than one species of hemoparasites is likely. Therefore, multiplex PCR (mPCR) for the simultaneous detection of Babesia spp. and A. marginale offers a significant advantage when analyzing a large number of samples. In fact, mPCR has been shown to be a very valuable tool in epidemiologic studies on hemoparasites.10,16 To date, mPCR assays are nested, but they consume time and materials, and nonspecific amplification is a common issue that hinders interpretation of results. Therefore, a simple mPCR that has good sensitivity and is validated for routine use would be advantageous.
Validation of a new test is usually assessed using a gold standard (reference test). The gold standard should have high accuracy.14,20 Despite its low sensitivity, a Giemsa-stained blood smear is taken as the gold standard for the detection of Babesia spp. and A. marginale. 1 In situations in which the reference test is not perfect and the true disease status is unknown, a Bayesian analysis method is applied to estimate the sensitivity and specificity of the test used and, in turn, disease prevalence.8,14,34 Researchers who have previously developed PCR assays for these hemoparasites estimated the sensitivity and specificity by comparison with the smear technique without using a Bayesian approach,1,3,17 which may have led to inaccuracy in estimation of these parameters.
Improving the detection of Babesia spp. and A. marginale is of paramount importance in Uruguay. Our aim was to improve the detection of these agents by applying and validating a mPCR. We estimated mPCR sensitivity and specificity using a Bayesian model in the absence of a gold standard test.
Materials and methods
Sampling
Convenience sampling was carried out between August 2016 and October 2018. We used 466 samples (340 peripheral blood with K3-EDTA and 126 refrigerated organs [kidney, spleen, liver, heart, brain]) from sick cattle and autopsy of clinical cases with presumptive diagnoses of babesiosis or anaplasmosis (n = 152 suspect outbreaks). These samples were sent by veterinary practitioners to the laboratory of the ‘División Laboratorios Veterinarios’ (DILAVE), Northwest Region of the ‘Ministerio de Ganadería, Agricultura y Pesca’ (MGAP), Uruguay. Samples were obtained from 122 farms distributed in 10 departments of Uruguay: Artigas, Salto, Paysandú, Rio Negro, Tacuarembó, Rivera, Flores, Lavalleja, Maldonado, and Soriano.
Presumptive cases were defined as animals with at least one clinical sign or autopsy finding compatible with babesiosis or anaplasmosis. The signs and/or autopsy findings were fever (>39.9°C), anemia (microhematocrit <0.26 L/L; centrifuged at 11,800 × g for 5 min), weakness, aggressiveness, jaundice, and splenomegaly. The average morbidity and mortality (minimum–maximum) registered in the farms from which samples were collected were 5.4% (0.2–32%) and 2.8% (0–25.7%), respectively.
Multiplex PCR for detection of Babesia spp. and A. marginale
Samples
Multiplex PCR was developed and set up with the first 30 samples received: 5 blood samples for each agent, B. bovis, B. bigemina, and A. marginale; 5 organ samples (liver, heart, kidney, spleen, brain) for each agent from autopsies of 3 animals naturally infected by B. bovis, B. bigemina, and A. marginale. These agents were positively identified based on smears, which had been fixed with methyl alcohol for 5 min and stained with Giemsa for 1 h. 35 All smear reads were by the same trained technician. At least 100 fields of each smear were read under a light microscope at 1,000× magnification with oil immersion. A sample was considered smear-positive if >2%, 1%, and >5% of parasitized erythrocytes were observed for B. bigemina, B. bovis, and A. marginale, respectively. 28 The morphologic criteria used for parasite identification were those used in previous reports.24,26
DNA extraction
DNA extraction from blood and organs was carried out (PureLink genomic DNA mini kit; Invitrogen) following the manufacturer’s instructions. Aliquots of 200 µL of fresh blood and 25 mg of tissue were used. The DNA concentration of aliquots was quantified (NanoDrop 2000 spectrophotometer; Thermo Scientific).
Multiplex PCR for amplification of Babesia spp. and A. marginale
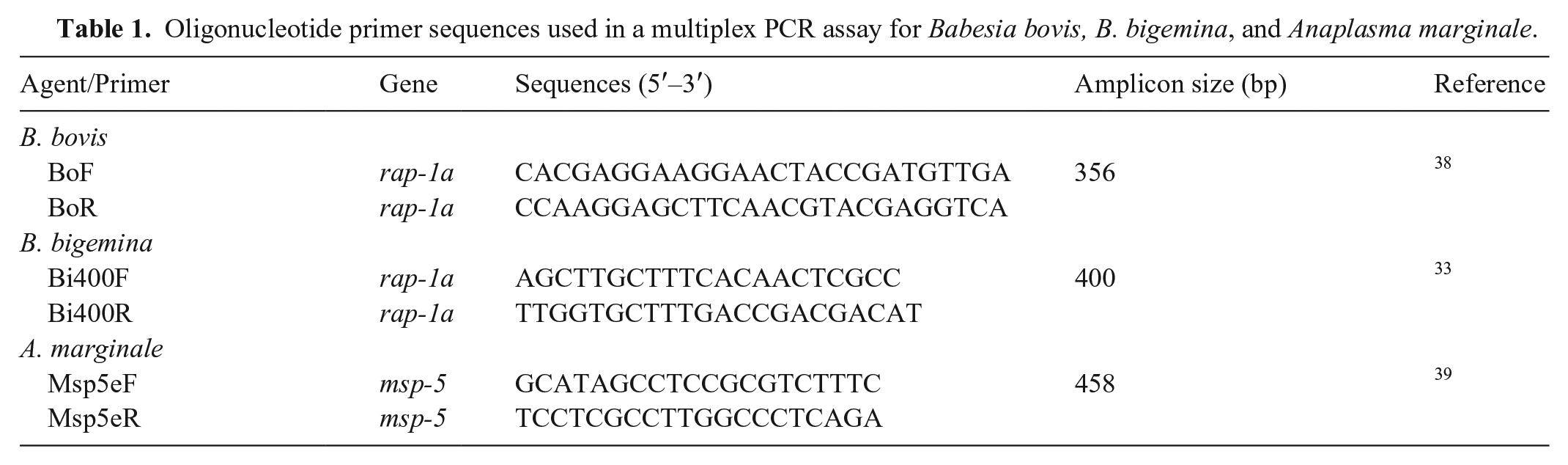
To develop the mPCR, the 30 blood and tissue samples mentioned above were used. A pair of specific primers for each parasite was selected, based on previous studies (Table 1). We used primers that amplified the rap-1a gene region of B. bovis and B. bigemina, and the msp-5 region of A. marginale. The selection criteria were based on similar annealing temperatures and differences in amplification sizes (>40 bp; Table 1). The reactions were performed in a final volume of 25 µL, consisting of 12.5 µL of MangoMix (Bioline; Meridian Bioscience), 2.5 µL of ultrapure water, 4 µL of sample DNA solution, and 1 µL (10 pmol) of each primer. The amplification protocol was as follows: initial denaturation at 95°C for 5 min, followed by 35 cycles at 95°C for 1 min, annealing at 58°C for 1 min, extension at 72°C for 1 min, and a final step at 72°C for 10 min. A negative control of ultrapure water and 3 previously sequenced positive controls (B. bovis, B, bigemina, and A. marginale) were included in each run of the assay. The amplification products from each mPCR were run on 1.5% agarose gel, which was stained (GoodView nucleic acid stain; Beijing SBS Genetech) and visualized under a UV transilluminator.
Oligonucleotide primer sequences used in a multiplex PCR assay for Babesia bovis, B. bigemina, and Anaplasma marginale.
Amplicons of the expected size were purified (PureLink quick PCR purification kit; Invitrogen) and sent for sequencing (Macrogen, Seoul, South Korea). The identities of the sequences were confirmed by BLASTn (http://www.ncbi.nlm.nih.gov/BLAST).
Limit of detection of single and mPCR from blood samples
The detection limit of single and mPCR was determined for each parasitic species using K3-EDTA blood parasitized with B. bovis, B. bigemina, or A. marginale. These positive samples were obtained from 3 acute clinical cases (based on clinical signs, hematocrit <0.20 L/L, agent identification in Giemsa-stained smears). For each agent, the number of parasitized erythrocytes (PE) based on 1,000 erythrocytes was observed (1,000×) in thin blood smears; the initial counts were: 8 (0.8% PE), 3 (0.3% PE), and 5 (0.5% PE) for B. bovis, B. bigemina, and A. marginale, respectively.
An initial blood dilution was prepared for each agent as follows: 40 µL of parasitized blood in 360 µL of blood extracted from healthy animals from tick-, Babesia-, and Anaplasma-free areas, located in the department of Colonia, Uruguay (33°59′42′′S, 57°56′33′′W). The hemoparasite-free blood was evaluated by single PCR to confirm the absence of hemoparasite DNA. Immediately, these dilutions were used as the base for 4 additional serial blood dilutions (10-1, 10-2, 10-3, 10-4, 10-5). From each dilution (total volume = 400 μL), DNA was extracted from 200 μL. Simple PCR assays for each species (using species-specific primers) and mPCR for Babesia spp. and A. marginale were performed under the conditions described above.
Statistical analysis
Statistical analysis was carried out with the detection data for 466 (340 blood and 126 organs) samples smeared and stained with Giemsa then subjected to the mPCR that we developed. The percent agreement between the techniques (smear and mPCR) and kappa coefficient for the detection of Babesia spp. and A. marginale were calculated using the package Classification and Regression Training of the software R v.3.6.3 (https://www.r-project.org/). 25
Sensitivity (Se) and specificity (Sp) of the smear and mPCR assays for the detection of Babesia spp. and A. marginale were estimated. The Bayesian approach was used,8,12 assuming that the true disease status in the population was unknown and that there was covariance between the tests. The unit of analysis was the biological sample (blood and/or organ; n = 466 samples). Test results were classified as follows: 1) Babesia spp. “positive” when parasites compatible with these species were found in the smears or amplified in the PCR reaction, otherwise “negative”; 2) A. marginale “positive” when a parasite compatible with this species was found in the smears or amplified in the PCR reaction, otherwise “negative.” Coinfection was counted as “positive” for either species in the analysis.
The Bayesian model had 7 parameters: Se and Sp for each of 2 tests, prevalence, and 2 covariance measures between tests (positive and negative results). As recommended, 8 4 of the prior distributions need to be informative; Se and Sp of the mPCR and blood smear were informative, and the prevalence (π) had a non-informative prior [π ~ Beta (1, 1)]. The prior distributions for the tests were obtained from previous reports, 17 which used experimentally infected animals. The parameters used to define the test prior distribution were: 1) Babesia spp.: Se t1 (mPCR) ~ Beta (30,3), Sp t1 (mPCR) ~ Beta (32,1); Se t2 (blood smear) ~ Beta (23,10), Sp t2 ~ (blood smear) Beta (32,1), which is the worst scenario from the 2 species of Babesia reported 20 ; 2) A. marginale: Se t1 (mPCR) ~ Beta (16,2), Sp t1 (mPCR) ~ Beta (17,1); Se t2 (blood smear) ~ Beta (16,2), Sp t2 ~ (blood smear) Beta (17,1).
Three simulations were run with different initial values, discarding the first 1,000 interactions of each as burn-in, using the OpenBUGS and BRugs packages in R software. Only one interaction of each 100 was saved for analysis. We used 12,000 interactions for analysis. Convergence testing was verified using trace plots, autocorrelation plots, and Gelman and Rubin statistics obtained with the coda package of the software R 18 (Supplementary material).
Results
The combination of the 3 pairs of primers for the detection of Babesia spp. and A. marginale successfully amplified all samples (n = 30). The band sizes were 356 bp, 400 bp, and 458 bp for B. bovis, B. bigemina, and A. marginale, respectively. The difference between the sizes of the amplicons was >40 bp; therefore, it was possible to differentiate each band produced by the DNA of each species, as well as the ability to amplify coinfected samples (Fig. 1).
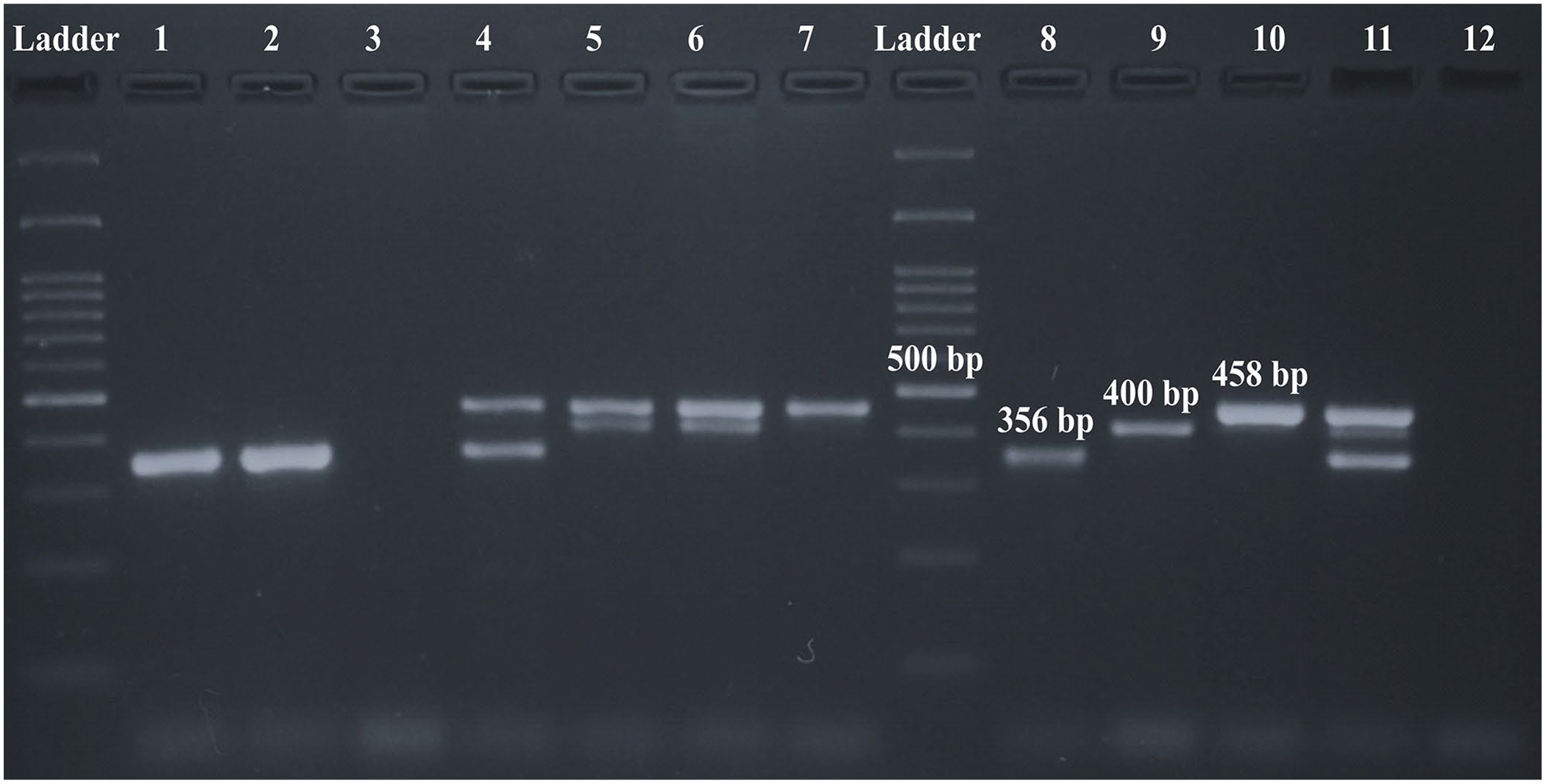
Multiplex PCR (mPCR) to detect Babesia spp. and Anaplasma marginale. Electrophoresis of mPCR products on 1.5% agar. Lanes: 1, 2 = positive samples for Babesia bovis; 3 = negative sample; 4 = positive sample for coinfection of B. bovis/A. marginale; 5, 6 = positive samples for coinfection of B. bigemina/A. marginale; 7 = positive sample for A. marginale; 8 = positive control for B. bovis (356 bp); 9 = positive control for B. bigemina (400 bp); 10 = positive control for A. marginale (458 bp); 11 = multiple positive control for B. bovis, B. bigemina, and A. marginale; 12 = negative control.
A sequence homology study was conducted to corroborate the identities of the 30 samples analyzed by mPCR. The results were as follows: 99–100% and 91–99% for samples of A. marginale (n = 10) and B. bovis (n = 10), and for samples of B. bigemina (n = 10), respectively (GenBank MK188829.1, AF030056.2, MK345485.1).
Limit of detection of single and mPCR with blood samples
Single PCR assays carried out with each species-specific primer were able to amplify the 10-3 dilutions of B. bovis (0.0008% PE) and B. bigemina (0.0003% PE), and the 10-4 dilution of A. marginale (0.00005% PE). The mPCR detected the 10-2 dilutions for all 3 agents: B. bovis (0.008% PE), B. bigemina (0.003% PE), and A. marginale (0.005% PE).
Descriptive results, agreement, and kappa coefficient
Based on smear reading, 24 samples were positive for a single infection (Babesia spp.) and 14 samples were positive for coinfection (Babesia spp. + A. marginale; Table 2). The species of Babesia could not be identified. A few samples (n = 8) that were not properly preserved affected the blood smear reading. The percent agreement between the smear and mPCR techniques were 89.5% (95% CI: 86.3–92.1%) and 91.6% (95% CI: 88.7–93.9%) for detection of Babesia spp. and A. marginale, respectively. The kappa coefficients were 0.72 (p < 0.001) and 0.79 (p < 0.001; moderate level of agreement) for detection of Babesia spp. and A. marginale, respectively.
Positive sample, by test, for Babesia bovis, B. bigemina, and Anaplasma marginale.
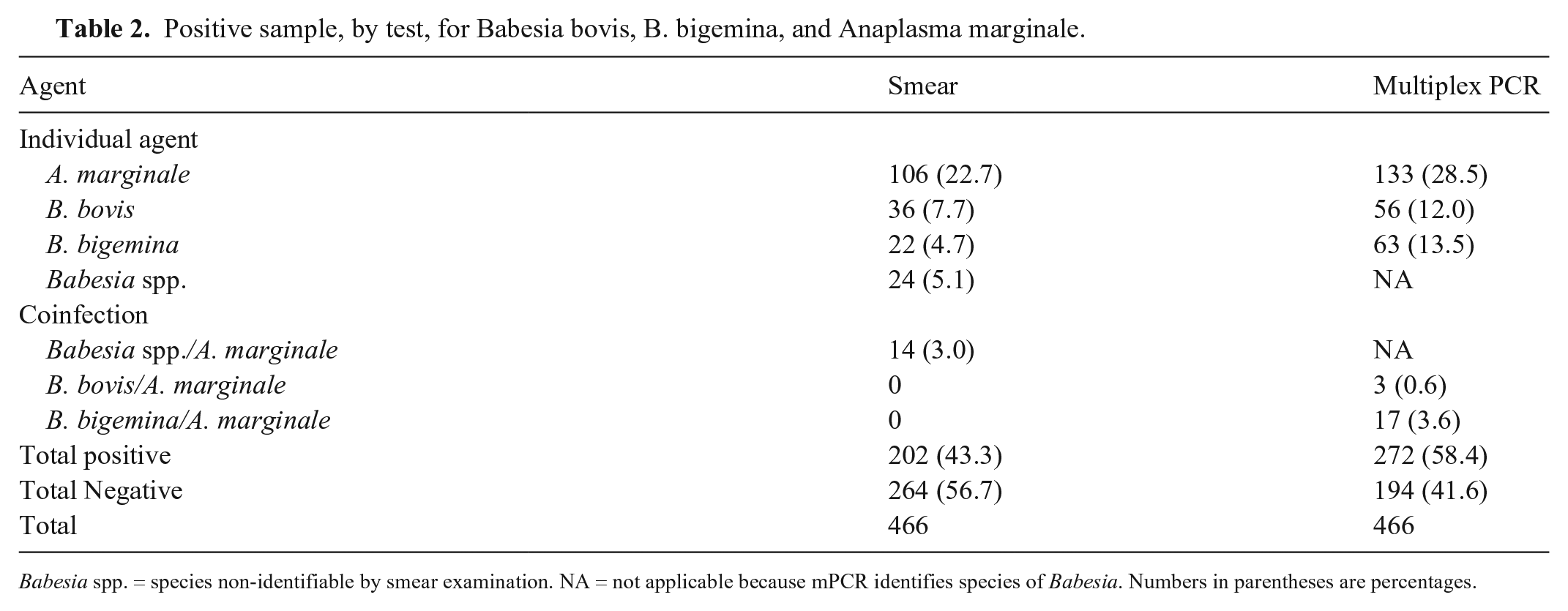

Babesia spp. = species non-identifiable by smear examination. NA = not applicable because mPCR identifies species of Babesia. Numbers in parentheses are percentages.
Sensitivity and specificity of the smear and mPCR techniques
The medians of Se and Sp estimated by the Bayesian analysis for the smear technique were 68.6% (95% CI: 58.8–79.6%) and 99.1% (95% CI: 95.9–100%) for Babesia spp., and 85.6% (95% CI: 71.7–97.1%) and 98.8% (95% CI: 93.1–99.9%) for A. marginale, respectively (Table 3). The median Se and Sp for the mPCR were 94.2% (95% CI: 84.8–98.5%) and 97.1% (95% CI: 90.5–99.9%), for Babesia spp., and 95.2% (95% CI: 85.2–99.1%) and 92.7% (95% CI: 85.6–99.2%) for A. marginale, respectively.
Results of the Bayesian analysis, sensitivity, and specificity for multiplex PCR (mPCR) and smear examination (smear) for Babesia bovis, B. bigemina, and Anaplasma marginale, in the absence of a gold standard test.
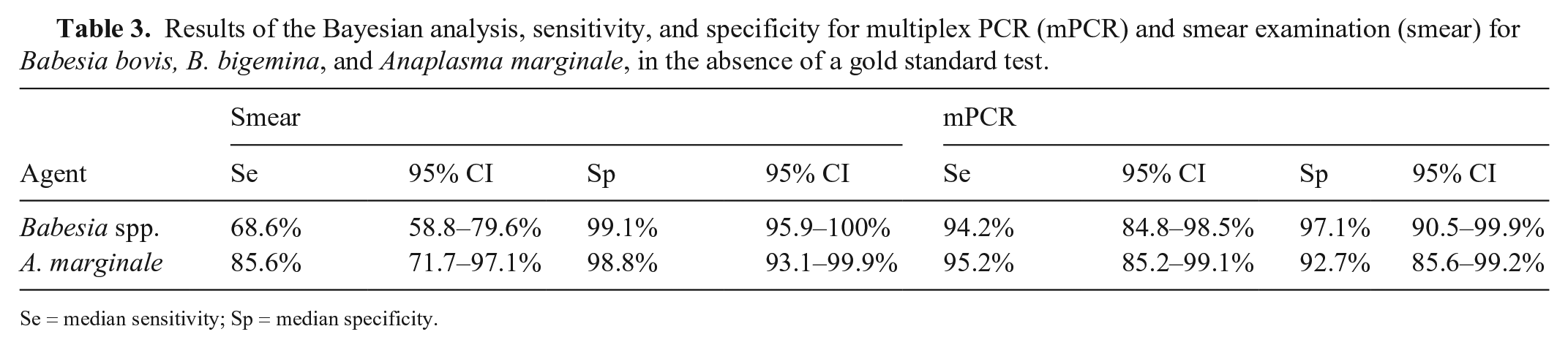
Se = median sensitivity; Sp = median specificity.
Discussion
A drawback in smear reading is the difficulty in differentiating Babesia species when immature forms are present, which requires skilled personnel for accurate detection.10,11,17 In our study, taking into account single infection and coinfection, discrimination could not be achieved in 38 Babesia spp.–positive smear samples (24 Babesia spp.; 14 Babesia spp.+ A. marginale). Most of the samples in our study were submitted under proper conditions; a few samples had deteriorated, which clearly affected smear reading (data not shown). Our data agree with other reports,1,10 in which the sensitivity of smears reading was reduced when samples were not optimal.
PCR protocols have been designed for detection of babesiosis and anaplasmosis agents, either individually or in combination, but most of these protocols are nested PCR.3,10.15,16,39 The main advantage of conventional mPCR is the ability to simultaneously amplify several pathogens without losing sensitivity or specificity. Our mPCR technique overcame this drawback as a single-round approach that was less time-consuming, minimized amplicon manipulation, and thus reduced the risk of false-positive results.
Previous studies noted that the sensitivity of mPCR could be affected by the amount of template DNA used as well as competition for dNTPs and Taq polymerase by different primers during the cycles.5,7,13 Although the sensitivity of the combined primers in our mPCR technique decreased, the concentration detected was sufficient to detect a minimum DNA dilution of 10-2, which allowed detection of coinfected samples and potential carrier animals.
Given that the treatments for anaplasmosis and babesiosis differ,6,24 the sensitivity and specificity of each test (smear and mPCR) for Babesia spp. and A. marginale was estimated separately. The results of the Bayesian model for the smear technique showed lower sensitivity than the mPCR technique. This result agreed with other studies that reported that PCR techniques are more sensitive for detecting Babesia spp. and A. marginale than the smear technique.1,3,10,16 Other reports indicate that blood smear reading has low sensitivity and does not visualize the parasite in early stages or carrier animals. 32 It should be noted that, in the reports mention above, sensitivity was calculated using the smear as the gold standard. Smear reading is a test that does not meet all conditions of a gold standard; therefore, it is likely that the sensitivity calculated by these authors is not accurate. For this reason, we used the Bayesian model to estimate parameters of sensitivity and specificity using a priori information, which allows these measures to be adjusted.23,27
The specificity of our mPCR technique was slightly lower than that of the smear technique. This slight decrease in specificity could be explained by the amount of DNA required for a positive result in a mPCR run. For instance, positive results could be obtained from a “healthy” or asymptomatic animal (carrier). These results are in accordance with previous reports. 32 In order to not interpret incorrectly a positive mPCR result (false-positive), clinicians must consider the clinical signs and or autopsy findings consistent with a diagnosis of babesiosis or anaplasmosis. Smears prepared with fresh and well-preserved samples and read by trained personnel reduce the chance of false-positive results. This could partially explain the higher specificity observed in our study.
Supplemental Material
sj-pdf-1-jvd-10.1177_1040638720975742 – Supplemental material for Validation of a multiplex PCR assay to detect Babesia spp. and Anaplasma marginale in cattle in Uruguay in the absence of a gold standard test
Supplemental material, sj-pdf-1-jvd-10.1177_1040638720975742 for Validation of a multiplex PCR assay to detect Babesia spp. and Anaplasma marginale in cattle in Uruguay in the absence of a gold standard test by Pablo Parodi, Luis G. Corbellini, Vanessa B. Leotti, Rodolfo Rivero, Cecilia Miraballes, Franklin Riet-Correa, José M. Venzal and María T. Armúa-Fernández in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dra. María Angélica Solari (DILAVE-Montevideo), and the veterinarians and farmers who submitted samples.
Declaration of conflicting interests
The authors declared no potential conflict of interest with respect to the research, authorship and/or publication of this article.
Funding
Our study received funds from the Instituto Nacional de Investigación Agropecuaria (INIA), Uruguay, project “Determination of the current situation of Rhipicephalus microplus and tick fever and integrated control of both diseases”, Animal Health Platform.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
