Abstract
Nanoparticle-assisted PCR (nanoPCR) is a novel method for the simple, rapid, and specific detection of viruses. We developed a nanoPCR method to detect and differentiate canine coronavirus I (CCoV I) and II (CCoV II). Primer pairs were designed against the
Canine coronaviruses include canine enteric coronavirus (CCoV) and canine respiratory coronavirus (CRCoV), which belong to genus
CCoV is an enveloped, single-strand, positive-sense RNA virus with a large genome of 27–32 kb.
3
The genome contains 10 open reading frames (ORFs). The 5′ two-thirds of the genome comprises 2 large, partially overlapping ORFs (1a and 1b) encoding 2 polyproteins that form the viral replicase. Downstream of ORF1b, 8 small ORFs encode the structural proteins, including the spike (S), envelope (E), membrane (M), and nucleocapsid (N), and a number of other presumptive nonstructural proteins.
5
Sequence and phylogenetic analyses of the
Nanoparticle-assisted PCR (nanoPCR) is an advanced form of PCR in which solid gold nanometal particles (1–100 nm diameter) form colloidal nanofluids with increased thermal conductivity.6,14 Therefore, PCR assays using nanofluids reach the target temperature faster than PCR assays using original liquids; the time at non-target temperatures is reduced, which decreases nonspecific amplification and increases specific amplification.7,10 Additionally, higher sensitivity is achieved through improved heat transfer. Moreover, nanoPCR does not require expensive machines, such as real-time PCR instruments; nanoPCR is also not contaminated as easily as are LAMP assays. Thus, nanoPCR is a sensitive, effective, and time-saving technique that has been utilized successfully to detect diverse pathogens, including porcine parvovirus, porcine bocavirus, mink enteritis virus, porcine epidemic diarrhea virus, and bovine respiratory syncytial virus.2,9,17,18,20
Reports of infections by CCoV are increasing; therefore, we aimed to develop a nanoPCR system to detect and differentiate canine CCoV I and CCoV II, using the
Published sequences of CCoV I and Ⅱ strains were obtained from GenBank. Primers were designed based on conserved regions of
Sequences of the primers used in our study of canine coronavirus I and II (CCoV I, II).
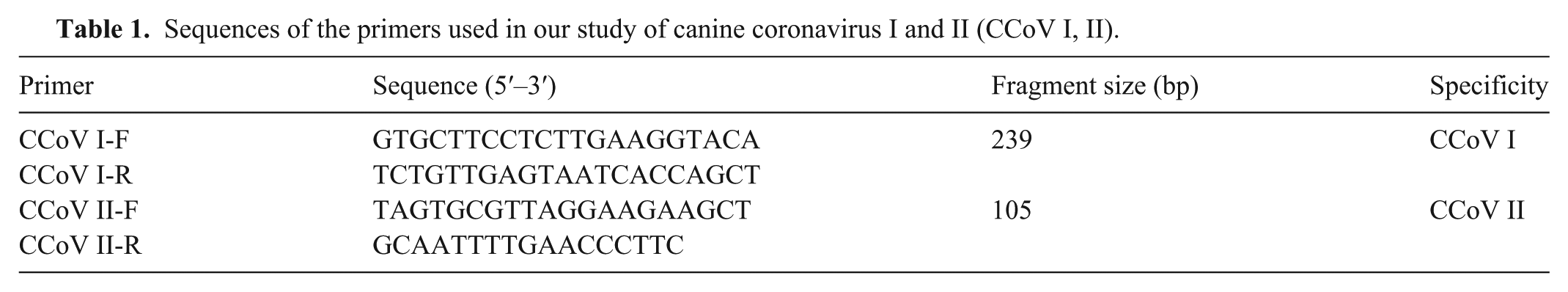

The CCoV nanoPCR system (NPK02; Weihai) was used according to the manufacturer’s instructions, as described in previous studies.18,21 The annealing temperature and primer concentration were optimized. The recombinant plasmids pMD-CCoV Ⅰ and pMD-CCoV II were used as templates in the optimization experiments. Annealing temperatures ranged from 46 to 53°C, and the amounts of primer (CCoV I-F, CCoV I-R, CCoV II-F, and CCoV II-R at 10 μM) ranged from 0.2 to 1.2 μL in increments of 0.2 μL. The optimal annealing temperature was 48°C (Fig. 1A); we used 48°C to determine the optimal primer concentrations. For all primers (CCoV I-F, CCoV I-R, CCoV II-F, and CCoV II-R at 10 μM), 1.0 μL was identified as the optimal amount of primer for the nanoPCR assay (Fig. 1B). Based on the optimized conditions, the nanoPCR assay was performed in a 20-μL reaction mixture comprising 0.5 μL of the pMD-CCoV I and pMD-CCoV II recombinant plasmids; 10 μL of 2× nanoPCR buffer; 1.0 μL of CCoV I-F/CCoV I-R or CCoV II-F/CCoV II-R (10 μM); 0.5 μL of Taq DNA polymerase (5 U/μL); and ddH2O up to 20 μL. The PCR reaction conditions were: 95°C for 3 min; 35 cycles of 95°C for 30 s, 48°C for 30 s, and 72°C for 30 s; and a final extension at 72°C for 10 min.
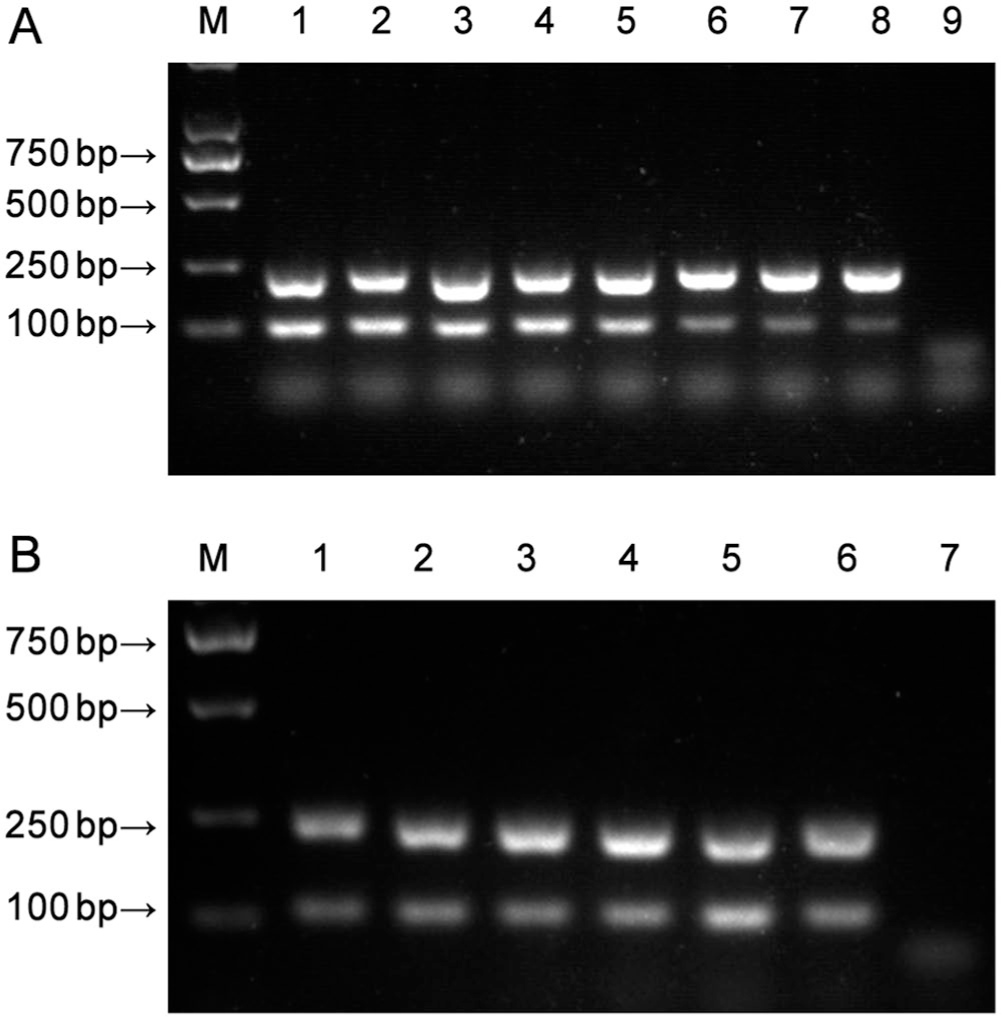
Optimization of the canine coronavirus nanoPCR.
The sensitivities of the CCoV nanoPCR assay and the conventional RT-PCR assay were compared using serial dilutions of the pMD-CCoV Ⅰ and pMD-CCoV II plasmids as templates. The concentrations of recombinant plasmid pMD-CCoV I and pMD-CCoV II were determined using UV spectroscopy (6.47 × 1010 copies/μL of pMD-CCoV I; 6.91 × 1010 copies/μL of pMD-CCoV II). The recombinant plasmids were diluted serially 10-fold from 6.47 × 108 to 6.47 × 100 copies/μL for pMD-CCoV I, and from 6.91 × 108 to 6.91 × 100 copies/μL for pMD-CCoV II. The nanoPCR assay was performed using the optimized reaction parameters. The RT-PCR assay was performed using the same primers and reaction parameters as used for the nanoPCR. Briefly, the conventional PCR assay was performed in a 20-μL system comprising 1.0 μL each of the recombinant plasmids pMD-CCoV I and pMD-CCoV II; 1.0 μL each of primers CCoV I-F, CCoV I-R, CCoV II-F, and CCoV II-R; 1.0 μL of dNTPs (2.5 mM); 0.5 μL of LA Taq (5 U/μL; Takara Bio); 2.0 μL of 10× LA Taq buffer Ⅱ (Mg2+ plus; Takara Bio); and ddH2O up to 20 μL. PCR products were analyzed by electrophoresis through 1.5% agarose gels. The detection limits of pMD-CCoV Ⅰ and pMD-CCoV II were 6.47 × 103 copies/μL and 6.91 × 104 copies/μL, respectively, using the conventional RT-PCR assay (Fig. 2A). By contrast, the detection limits were 6.47 × 101 copies/μL and 6.91 × 102 copies/μL, respectively, using the CCoV nanoPCR assay (Fig. 2B). Hence, the CCoV nanoPCR assay was 100 times more sensitive than the conventional RT-PCR assay.
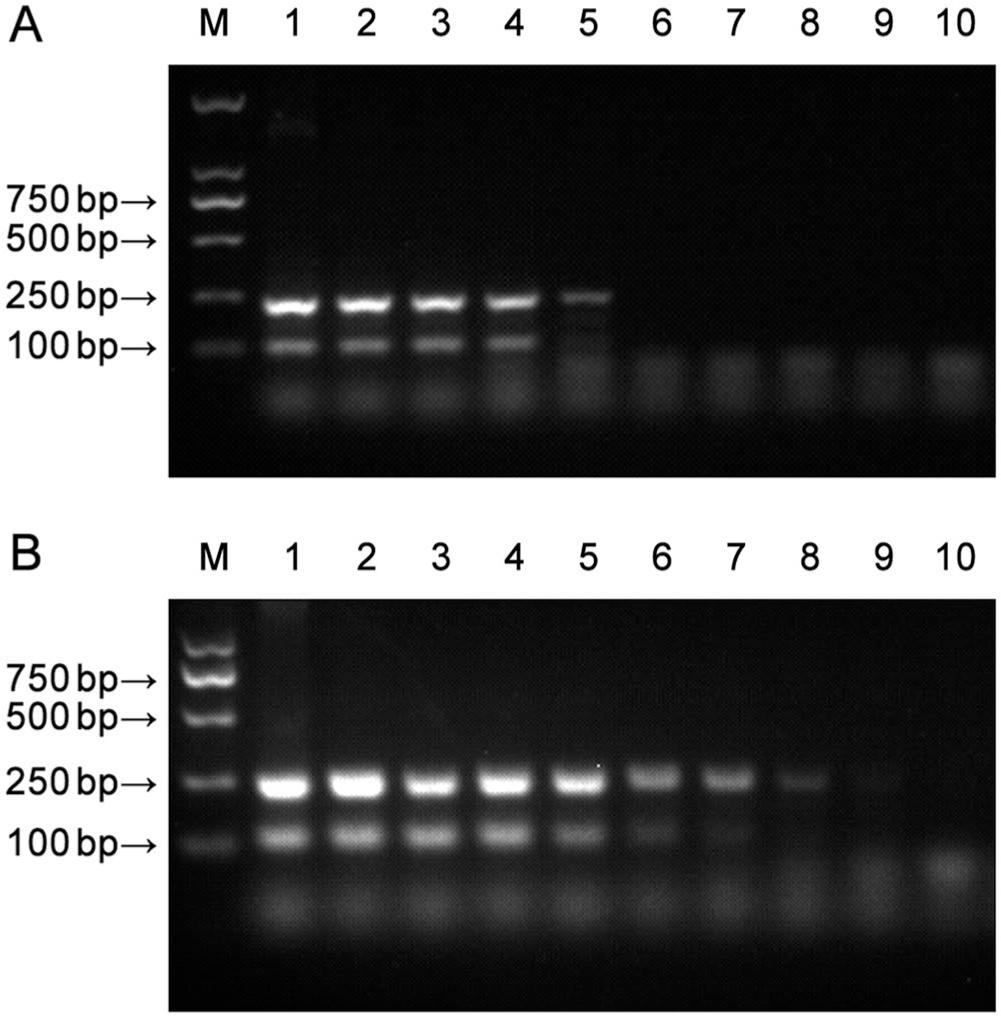
Sensitivity of the canine coronavirus (CCoV) RT-PCR (
The specificity of the CCoV nanoPCR assay was evaluated using other canine viruses including canine distemper virus, canine adenovirus 2, canine parainfluenza virus, and canine parvovirus, with ddH2O as a negative control. The nanoPCR assay did not amplify any of these 4 viruses (Fig. 3), indicating that the CCoV nanoPCR assay was highly specific.
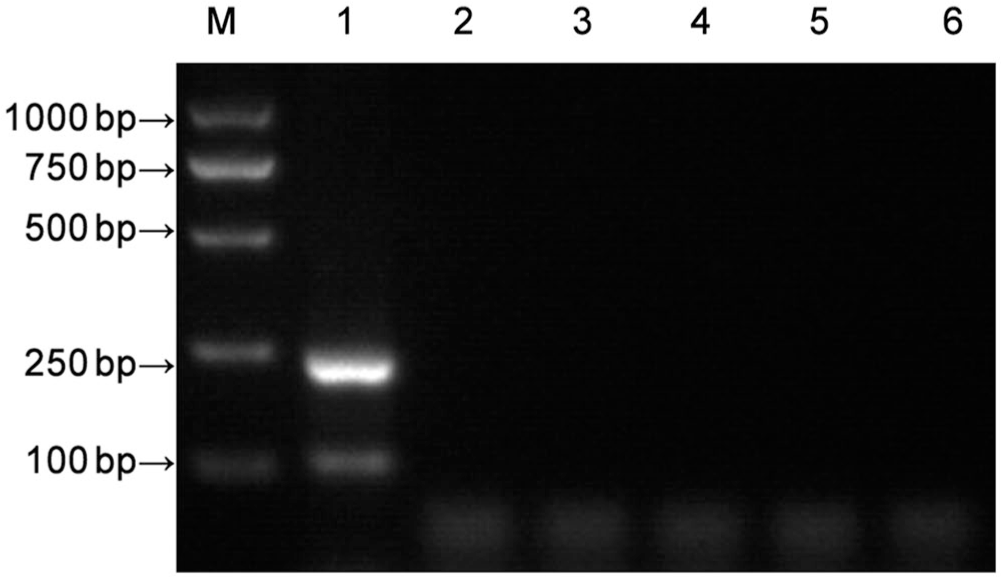
Specificity of the canine coronavirus (CCoV) nanoPCR. Lanes: M = Trans 2K plus DNA marker; 1 = CCoV Ⅰ and CCoV II; 2 = canine distemper virus; 3 = canine adenovirus 2; 4 = canine parainfluenza virus; 5 = canine parvovirus; 6 = negative control.
Feces from dogs with diarrhea were obtained from a pet hospital in Beijing. All samples were collected in accordance with the International Guiding Principles for Biomedical Research Involving Animals (https://olaw.nih.gov/sites/default/files/Guiding_Principles_2012.pdf). Fecal samples were diluted with 0.01 M (pH 7.6) phosphate-buffered saline and homogenized. Diluted fecal samples were repeatedly frozen and thawed, after which the samples were centrifuged at 12,000 ×
Detection rates of canine coronavirus I and II (CCoV Ⅰ, Ⅱ) in 60 clinical samples using conventional reverse-transcription PCR (RT-PCR) and CCoV nanoPCR.
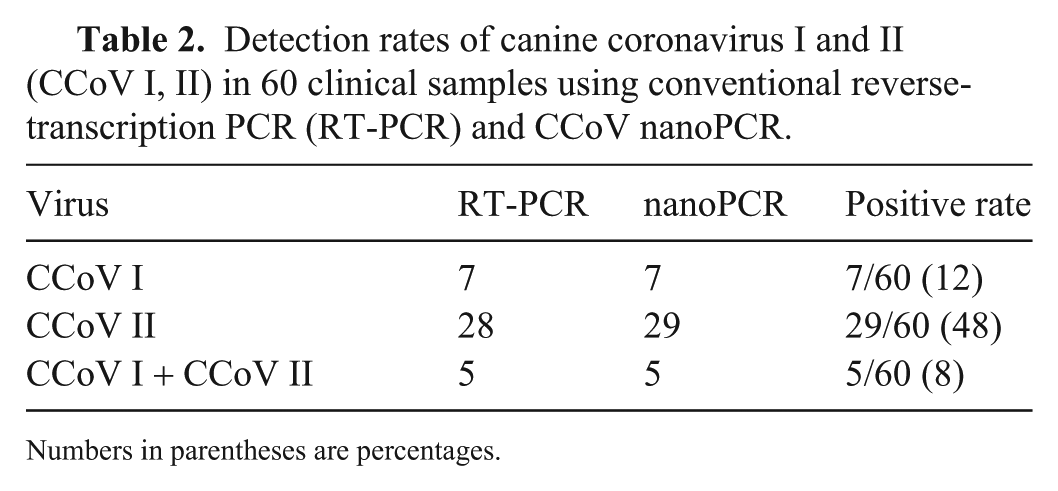
Numbers in parentheses are percentages.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Agricultural Science and Technology Innovation Program (grant ASTIP-IAS15), Scientific Observing and Experimental Station of Veterinary Drugs and Diagnostic Techniques Open Operating Fee, the Ministry of Agriculture (2019-YWF-ZX-10), and the National Key Research and Development Program of China (grant 2016YFD0501003).
