Abstract
Schmallenberg virus (SBV), discovered in Germany in 2011, causes congenital malformations in ruminants. Reverse-transcription real-time PCR (RT-rtPCR) assays based on various segments of SBV have been developed for molecular detection. We developed alternative RT-rtPCR assays for SBV detection to avoid earlier reported mutations and hypervariable regions of the S and M segments of the viral genome. For SYBR Green-based detection of the S segment, the R2 value and efficiency of the developed assay were 0.99 and 99%, respectively. For probe-based S segment detection, 2 assays were developed; the first had an R2 value of 0.99 and 102% efficiency, and the second had a R2 value of 0.98 and 86% efficiency. The probe-based M segment assay had an R2 value of 1.00 and 103% efficiency. Detection limits of the RT-rtPCR assays with new primer sets were 102 and 101 copies/µL for the S and M segments, respectively. Field samples from cattle and sheep were also used for primary validation of the developed assays. Our assays should be suitable for SBV detection in ruminants and for in vitro studies of various SBV strains.
Schmallenberg virus (SBV; Bunyavirales, Peribunyaviridae, Orthobunyavirus) was discovered in 2011 in Germany, 13 and epidemiologic studies reveal that SBV is present in many European countries, including Turkey.4,19 SBV has a 3-segment RNA genome consisting of small (S; 830 nt), medium (M; 4,415 nt), and large (L; 6,864 nt) segments. 13
Studies have reported some mutations and hypervariable regions in SBV segments since the discovery of SBV.7-9,14 The M segment contains a hypervariable region between 1,483–1,864 nt in German SBV isolates. 9 The M segments of Belgian SBV isolates from sheep include the hypervariable region between 1,394–2,562 nt 7 (Suppl. Fig. 1). The N-terminal region of the Gc protein, which is encoded by the M segment of SBV, harbors mutations, although other regions of the M segment of SBV are conserved. Although the S segment of SBV was found to be conserved, 14 analysis of the S segment sequences reveals that the nonstructural protein coding region (57–332 nt) has many nucleotide substitutions. 8 The L segment was found to contain point mutations, but all were silent. 14
Reverse-transcription real-time PCR (RT-rtPCR) is commonly used to detect genome segments of various agents 11 including SBV. 5 We developed SYBR Green- and probe-based alternative RT-rtPCR assays for detection of the S and/or M segments of SBV by using primers that were designed to avoid the reported mutations and hypervariable regions7–9,14 of the S and M segments. We optimized and validated the selected primer sets in SYBR Green- and probe-based RT-rtPCR assays by using plasmids and field samples and compared efficiency, coefficient of correlation (R2), slope, and reproducibility or repeatability of the assays.
SBV (strain F6; GenBank accessions KC355457–KC355459 represent sequences from the 3 segments of SBV), from the laboratory of Dr. Wim H. M. van der Poel (Wageningen Bioveterinary Research), was propagated in Vero cells at 37°C in 5% humidified CO2. SBV propagations were confirmed by RT-PCR with 382-469 primers 5 and a plaque titration 1 assay (Suppl. Fig. 2).
Primer sets specific for the S and M segments of SBV were designed (LightCycler probe design software 2.0; Roche) to avoid possible mutation regions on the segments that have been reported (Suppl. Fig. 1).7-9 Available SBV isolates in GenBank were used (n = 41 for S segment, n = 104 for M segment), and the most conserved regions in the S and M segments were selected as primer-binding sites. All primers and probes were synthesized by Ella Biotech (Planegg, Germany).
RNA samples were isolated (High Pure viral RNA kit; Roche), and reverse transcription was carried out as described earlier. 2 The S and M segments of SBV were amplified by conventional RT-PCR (Table 1). To amplify the full S segment (with AKASBV primers), cycling parameters were as follows: 95°C for 2 min, followed by 35 cycles of 95°C for 30 s, 54°C for 30 s, 72°C for 30 s, and final extension at 72°C for 6 min. To amplify the partial M segment (with M1-1107 primers), cycling parameters were as follows: 95°C for 2 min, 36 cycles of 95°C for 30 s, 54°C for 60 s, 72°C for 60 s, and final extension at 72°C for 10 min.
Primer sets used for amplification of Schmallenberg virus (SBV) S and M segments and optimization of SYBR Green- and probe-based assays.

F = forward primer; P = probe; R = reverse primer.
Whole S (702 bp) and partial M (1,107 bp) segments that were amplified by conventional RT-PCR were cloned into the pCR2.1 cloning vector (TA Cloning kit; Invitrogen) according to the manufacturer’s instructions. Selection of the S and M segment positive clones was carried out by PCR screening 3 with the aforementioned segment-specific primers, and plasmid DNA was isolated (Invitrogen PureLink HiPure plasmid midiprep kit; Thermo Fisher Scientific). DNA concentrations of positive plasmids were measured by spectrophotometer (MSP touch-200; Inovia Teknoloji) and calculated using the formula described earlier. 18 Concentrations were calculated as 1010 copies/µL for both the S and M segment plasmids. The S and M plasmids were 10-fold diluted, ranging from 101 copies/µL to 109 copies/µL, in molecular-grade water.
The positive plasmid dilutions were used as standards in rtPCR assays that were performed with SYBR Green I master mix (Roche) and probes master mix (Roche) in a LightCycler 96 rtPCR instrument (Roche).
The SYBR Green-based assay was optimized in 20 µL of total reaction volume consisting of 1 µL of template, 2× SYBR Green I master mix, a final concentration of 500 nM of each primer, and PCR-grade water up to the final volume. The probe-based assays were performed with 1 µL of template, 2× probe master mix, a final concentration of 500 nM of each primer, a final concentration of 150 nM of probe, and PCR-grade water up to the final volume of 20 µL.
For detection of the S segment in the SYBR Green-based assay, the following profile was used: 10 min at 95°C, followed by 38 cycles of 10 s at 95°C, 10 s at 56°C, 10 s at 72°C. The profile for detection of the S segment in probe-based assays was: 10 min at 95°C, followed by 40 cycles of 15 s at 95°C, 20 s at 54°C, 30 s at 72°C. The M segment was detected using the profile of 10 min at 95°C pre-incubation, 40 cycles of 10 s at 95°C, 10 s at 55°C, 10 s at 72°C. Temperature profiles and melting curve analysis were evaluated using LightCycler 96 SW1.1 software (Roche).
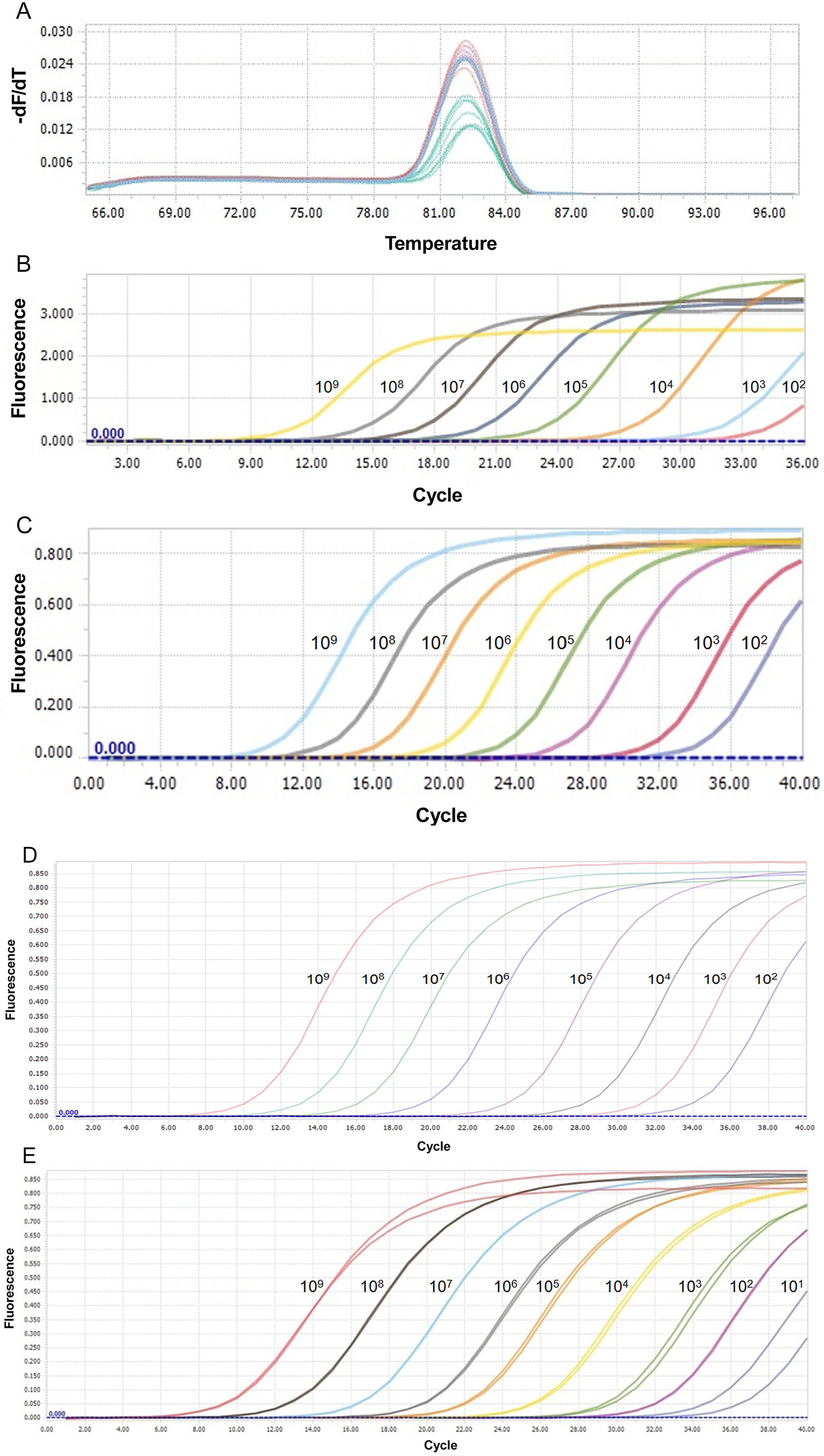
The optimal annealing temperature of the 304-415 primers, 376-508-P436 and 368-533-P419 primers for S segment, and 49-213-P82 primers for M segment (Table 1) were 56°C, 54°C, and 55°C, respectively. The melting curve of 304-415 primers (Fig. 1A) and the amplification curves of SYBR Green- and probe-based assays (Fig. 1B–E) were generated using the same software. The detection limit of both SYBR Green- and probe-based assays for the S segment was 102 copies/µL. The detection limit of the probe-based assay for the M segment was 101 copies/µL.
Efficiency, R2 value, and slope of primers-probes were determined (Fig. 2A–D). The efficiency of the SYBR Green-based assay with the 304-415 primers were 99%, and the R2 of the standard curve was 0.99 (Fig. 2A). The efficiency and R2 value of the probe-based assay with the 376-508-P436 primers-probe, which was designed for the S segment, were 102% and 0.99, respectively (Fig. 2B). The 368-533-P419 primers-probe for probe-based S segment detection had 86% efficiency and R2 value of 0.98 (Fig. 2C). The R2 value was 1.00, and the efficiency was 103% for the rtPCR assay for the M segment (Fig. 2D).
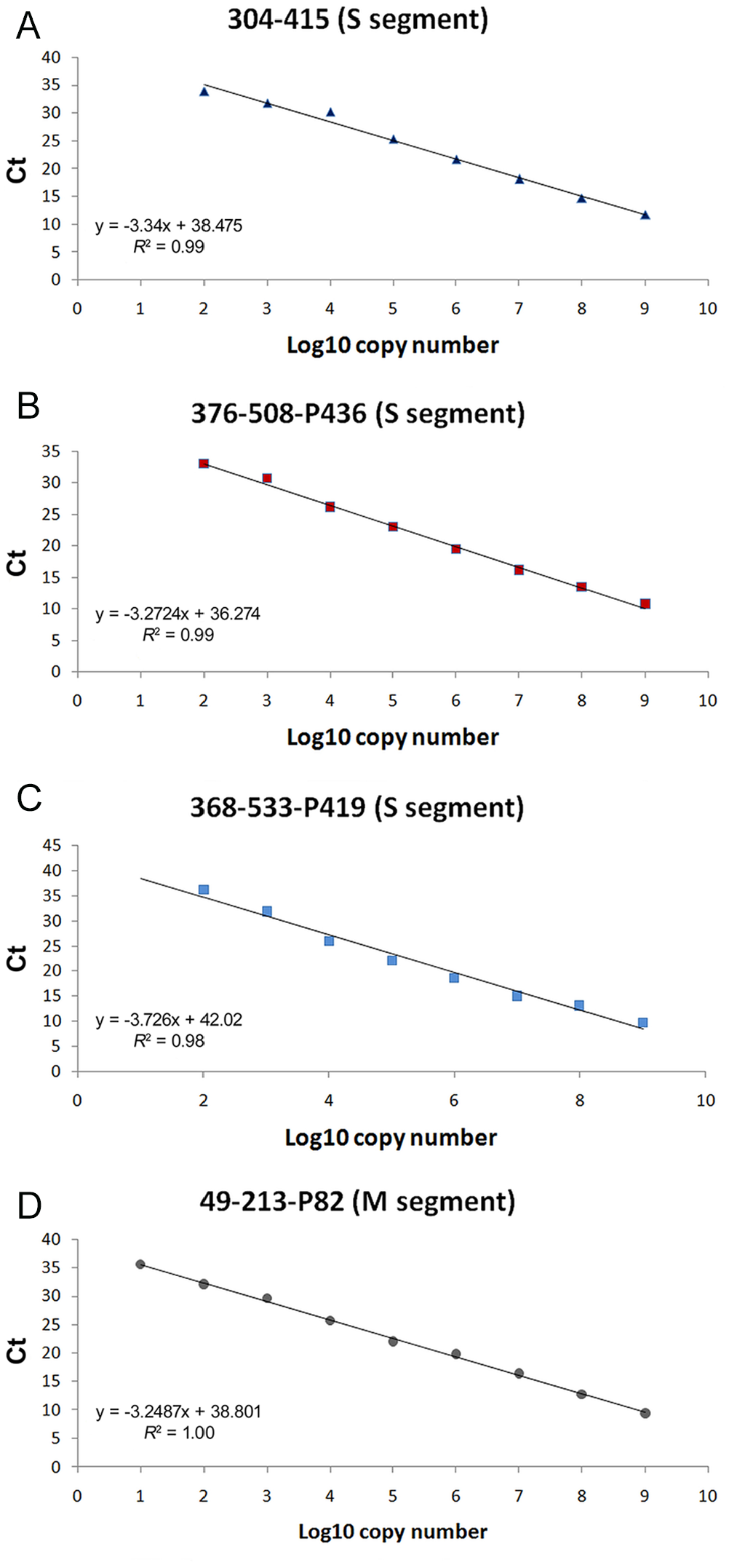
Standard curves of the newly designed primers with plasmid dilutions.
In a well-optimized rtPCR assay, the slope of the standard curve should be −3.2 to −3.5, and the coefficient of correlation should be close to 1 (i.e., R2 ≥ 0.98). 17 Based on these results and according to plasmid standards, the assay for M segment detection was the most sensitive detection method for SBV.
To determine the sensitivity of the assays, 10-fold dilutions of standard plasmids of 109–101 copies/µL were used. The intra-assay test was conducted with duplicates within the same run. The inter-assay test was performed as 3 different runs on different days. The mean, standard deviation (SD), and coefficient of variation (CV) were estimated for the intra- and inter-assay analyses. The R2 of the standard curve and efficiency were calculated 6 and multiplied by 100% to obtain the efficiency percentage (E%) using mean cycle threshold (Ct) values. 15 Intra- and inter-assay variations were performed to assess the reproducibility and repeatability with the plasmid standards (107,106, 105, 104 copies/µL; Table 2).
Reproducibility and repeatability of SYBR Green- and probe-based RT-rtPCR assays for the S and M segments of Schmallenberg virus. The intra-assay test variability was conducted with duplicates within the same run. The inter-assay test was performed as 3 runs on different days.
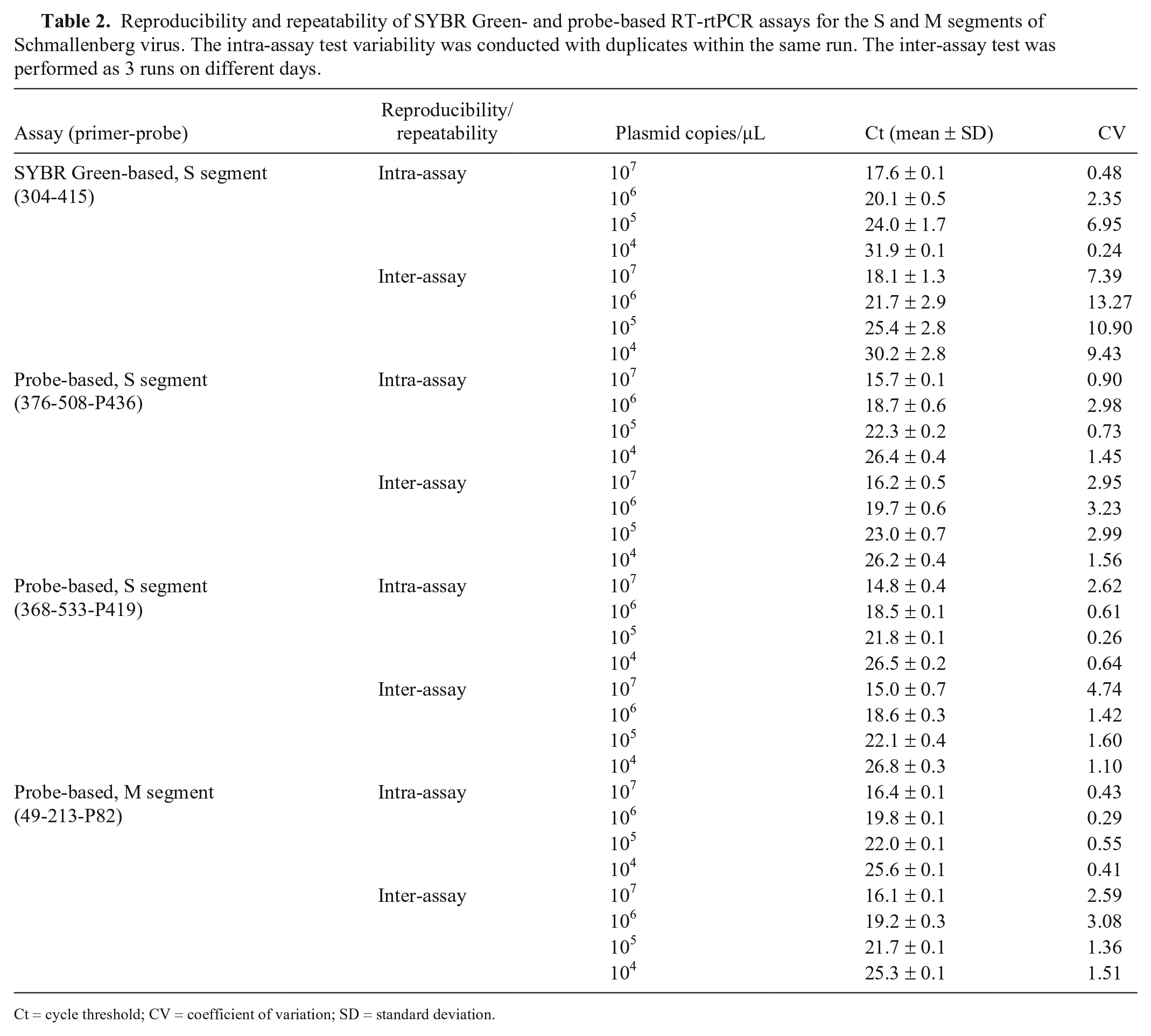
Ct = cycle threshold; CV = coefficient of variation; SD = standard deviation.
The CVs of both probe-based assays for the S segment were lower than the SYBR Green-based assay, indicating that the probe-based RT-rtPCR is more reproducible (Table 2). Among the newly developed assays, the probe-based assay for the M segment had the highest reproducibility and repeatability (Table 2).
Following optimization of the assays, SBV-positive field samples (4 bovine sera, 8 sheep sera, and 8 sheep brain samples) obtained from the laboratory of Dr. van der Poel (Wageningen Bioveterinary Research) were tested for validation of the assays. The samples were collected from farms in which animals had acute clinical signs associated with SBV infection. Serum samples tested positive by virus isolation and/or demonstration of seroconversion using a virus neutralization test. 16 Viral complementary DNA (cDNA) from SBV-positive field samples were further confirmed by the 382-469 primers 5 and tested as positive (Suppl. Table 1). All primers-probes developed in our study were tested with samples of bovine rotavirus, bovine coronavirus, Epstein-Barr virus (positive human sera), Akabane virus cDNA, and Hazara virus S segment plasmid, and results were negative (data not shown).
The SYBR Green-based assay scored all 4 bovine and 16 ovine samples as positive (20 of 20) by using the 304-415 primers for detection of the S segment (Table 3). The results of the 304-415 primers were compatible with the results of the 382-469 5 primers (Table 3; Suppl. Table 1). The Ct value of one sheep brain sample could not be determined in the 376-508-P436 assay but was detected at a Ct of 38.0 in the 368-533-P419 assay and was considered as suspicious (Table 3). The 49-213-P82 primers-probe for the M segment detected 19 samples as positive; 1 was considered as suspicious (Table 3). It was hypothesized that the 49-213-P82 primers-probe did not miss a positive signal because the primers-probe did not match any mutation region in the M segment. Consistent with the findings of others, 12 we detected the lowest Ct values in brain samples among sera and brain field samples (Table 3).
Cycle threshold (Ct) values of Schmallenberg virus genome positive bovine and ovine field samples tested with newly designed assays. Ct ≥37 and Ct ≥38 are considered as suspicious for probe-based S and M segment assays, respectively.
ND = not determined.
In a previous study, the 382-469-P408 primers 5 for the S segment were found to be more sensitive than primers-probe that are specific for the M and L segments. 10 The reason for this may be that the researchers 10 designed the primers for the M segment for position 1,690–1,827 nt of the M segment, which has been determined to be a hypervariable region7,9 (Suppl. Fig. 1). In our study, the 49-213-P82 primers-probe, specific for the M segment, does not target the positions that have been reported to be a hypervariable region (Suppl. Fig. 1). This may be the reason for the higher sensitivity of the M segment primers-probe in our study; mismatches between the primer and the M segment may reduce assay sensitivity. 10 The 368-533-P419 assay for S segment detection and the 49-213-P82 assay for M segment detection could be useful and alternative tools for detection of SBV infections in ruminants and for detection of the SBV genome in in vitro assays.
Supplemental Material
Supplemental_material – Supplemental material for Development and validation of SYBR Green- and probe-based reverse-transcription real-time PCR assays for detection of the S and M segments of Schmallenberg virus
Supplemental material, Supplemental_material for Development and validation of SYBR Green- and probe-based reverse-transcription real-time PCR assays for detection of the S and M segments of Schmallenberg virus by Ahmet Kursat Azkur, Wim H. M. van der Poel, Emel Aksoy, Renate Hakze-van der Honing, Murat Yildirim and Kader Yıldız in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dr. Mehmet Ziya Doymaz (Department of Medical Microbiology, Faculty of Medicine, Bezmialem Vakif University, Istanbul, Turkey) for his critical reading of the manuscript. Hazara virus S segment plasmid and Akabane virus cDNA were kindly provided by Drs. Mehmet Ziya Doymaz and Harun Albayrak (Department of Virology, Faculty of Veterinary Medicine, Ondokuz Mayis University, Samsun, Turkey), respectively.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study is a part of a project supported by the General Directorate of Agricultural Research and Policies, Ministry of Agriculture and Forestry, Republic of Turkey (grant TAGEM15/AR-GE48).
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
