Abstract
Calf diarrhea is a common cause of pre-weaning morbidity and mortality in cattle operations. We evaluated the role of Escherichia coli by assessing the frequency of genes encoding virulence factors (virotypes) in E. coli from feces or intestinal contents, and the association of these virotypes or other diarrheagenic pathogens with intestinal morphologic changes in calves with or without diarrhea. E. coli was isolated from 408 feces and 105 intestines of calves with diarrhea and compared to those isolated from 635 feces and 100 intestines of calves without diarrhea, from 2002 to 2016. Virotype EAST1:F17, in combination with minor virotypes, was the most commonly detected type, but without differences in frequency between the 2 groups of calves. No significant intestinal morphologic changes were observed with the different E. coli virotypes in either group of calves, except for bacterial attachment to enterocytes for virotype STa:F5, which was detected only in calves with diarrhea. These observations suggest that E. coli, excluding virotype STa:F5, is not a significant diarrhea-causing agent in calves. However, the intestinal lesions observed in ~82% of calves with diarrhea were attributed to other diarrheagenic pathogens that include bovine coronavirus, Clostridium perfringens, Cryptosporidium spp., Eimeria spp., rotavirus, and Salmonella spp.
Diarrhea in calves is a common clinical sign associated with pre-weaning morbidity and mortality in cattle operations worldwide. 5 Multiple factors associated with this diarrhea include pathogens such as Escherichia coli, Clostridium perfringens, Salmonella enterica ssp. enterica, rotavirus (RV), bovine coronavirus (BCoV; species Betacoronavirus 1), Cryptosporidium spp., and Eimeria spp. 7 Failure of passive colostral transfer to the calf and myriad environmental factors may have a role in development of calf diarrhea. 5
The role of E. coli has been recognized for many years, and neonatal colibacillosis is still commonly accepted as an important cause of diarrhea in calves.5,7 Multiple E. coli pathotypes have been associated with diarrhea in domestic animals; however, enterotoxigenic E. coli (ETEC) producing heat-stable enterotoxin A (STa) and expressing F5 fimbriae (i.e., virotype STa:F5) is considered the major cause of diarrhea in calves that occurs during the first week of life.9,14 The role of other E. coli virotypes in production of diarrhea has not been demonstrated clearly. 20 We, therefore, assessed the frequency of E. coli virotypes and their potential role in calf diarrhea.
We collected 1,248 feces or intestinal contents of distal ileum and proximal colon during calving seasons from newborn to 8-wk-old calves with (n = 513) or without (n = 735) diarrhea at different cow–calf operations in western Canada. For the first study, feces were collected from 2002 to 2013 and intestines collected as a complementary study from 2013 to 2016. Among these samples, 408 E. coli isolates were recovered from feces and 105 from fresh intestines of calves with diarrhea. For comparison, we used 635 isolates from feces and 100 from intestines of calves without diarrhea. Intestinal samples from calves without diarrhea were collected during calving seasons as well as from animals with conditions affecting body systems other than the intestinal tract (n = 100).
All feces and fresh pooled intestinal contents were inoculated onto 5% sheep blood agar plates (Oxoid, Nepean, Ontario, Canada) and incubated at 37°C for 18 h in an environment containing 5% CO2, and onto MacConkey agar plates (Oxoid) incubated as above under normal atmospheric conditions. E. coli was identified using standard biochemical tests for Enterobacteriaceae 6 prior to 2014, or by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS; Bruker Daltonics, Billerica, MA) for isolates collected after 2014. 13 All primary cultures on blood and MacConkey agar plates were sealed with a film and shipped to the OIE Reference Laboratory, the Escherichia coli Laboratory (EcL), for E. coli virotyping using colony hybridization or PCR, as described previously.3,15
In addition to E. coli isolation, all 205 fresh intestinal contents were subjected to 1) culture for C. perfringens and Salmonella spp.; 2) PCR for the major bovine enteric viruses (RV and BCoV); and 3) routine fecal flotation for the detection of Cryptosporidium spp. and Eimeria spp., using standard laboratory protocols.4,7
The detection of E. coli virotypes was carried out as follows: for feces, 5–10 E. coli colonies from the primary MacConkey agar plate cultures were examined, whereas, for intestines, confluent growth from primary cultures was examined. All isolates or confluent growth were suspended in trypticase soy broth (TSB; Difco Laboratories, Detroit, MI) containing 15% glycerol and stored at −80°C. An aliquot of the stored material was inoculated into 5 mL of lysogeny broth (Difco) and enriched overnight at 37°C and DNA templates prepared by heat lysis. 12 The boiled cell suspensions were centrifuged, and the resulting lysates were used for colony hybridization or PCR. Briefly, virotyping was carried out at the EcL by colony hybridization using radioactively labeled (32P) DNA probes or PCR for virulence genes that define the E. coli pathotypes commonly found in calves—ETEC (estA and f5), enteropathogenic E. coli (eae), shiga toxin–producing E. coli (stx1 and stx2)—and additional virulence genes astA, cdtB-1, cnf1/2, iucD, iroN, f17, papC, tsh, sfaA, afaD8, and paa associated with E. coli from cases of diarrhea and/or septicemia15,20 (Supplementary Table 1). Procedures for detection of these genes were performed according to the protocol of the EcL available on the APZEC website (http://apzec.ca/en/Protocols) and as described previously. 10 Water was used as the negative control.
Distal ileum and proximal colon of calves submitted live to the Prairie Diagnostic Services postmortem suite (University of Saskatchewan, Saskatoon, Canada) were collected and submitted fresh for bacteriology, parasitology, and virology tests and additional portions (duodenum, jejunum, ileum, and colon) fixed in formalin for histologic examination. Similarly, fresh and formalin-fixed tissue samples from field autopsies were collected within 6 h after death of animals to minimize autolysis and shipped to the laboratory soon after collection. Portions of the 205 intestines collected were fixed in 10% neutral-buffered formalin for 48 h before trimming and processed routinely, for histologic and bacteriologic examination. 11 Cases with intestinal tissues in advanced autolysis were excluded from the study.
In the first study, various E. coli virotypes were detected without discrimination in isolates from both groups of calves, regardless of geographic location. The main virotypes included the following: EAST1:F17; EAST1:CDT:CNF:F17; and EAST1:EAE, this last virotype associated or not with virotypes CDT, CNF, Stx1, Stx2, and F17 (Table 1).Virotype STa:F5, associated or not with EAST1 or F17, was detected only in feces from calves with diarrhea, suggesting that this virotype remains the most significant cause of E. coli diarrhea in calves.
Escherichia coli virotypes detected in isolates from feces of calves with or without diarrhea, 2002–2013.
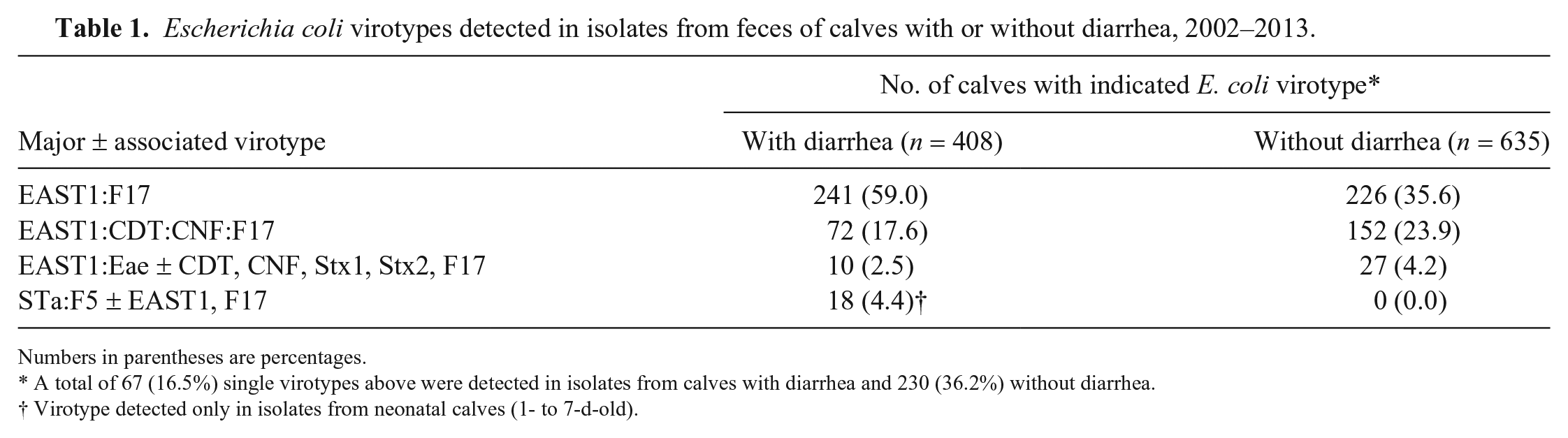
Numbers in parentheses are percentages.
A total of 67 (16.5%) single virotypes above were detected in isolates from calves with diarrhea and 230 (36.2%) without diarrhea.
Virotype detected only in isolates from neonatal calves (1- to 7-d-old).
In the subsequent study, we expanded the tests for virulence genes to include estA, astA, stx1a, stx2a, cdtB-1, cnf1/2, iucD, iroN, eae, f5, f17, papC, tsh, sfaA, afaD8, and paa. Similar to the findings of our first study, these virulence genes were also detected without discrimination in both groups of calves in the different geographic locations, with the exception of estA and f5 genes encoding virotype STa:F5, which were detected only in a sample from a 6-d-old calf with diarrhea (Table 2).
Distribution of virulence genes in Escherichia coli from intestinal contents of calves with or without diarrhea, 2013–2016.
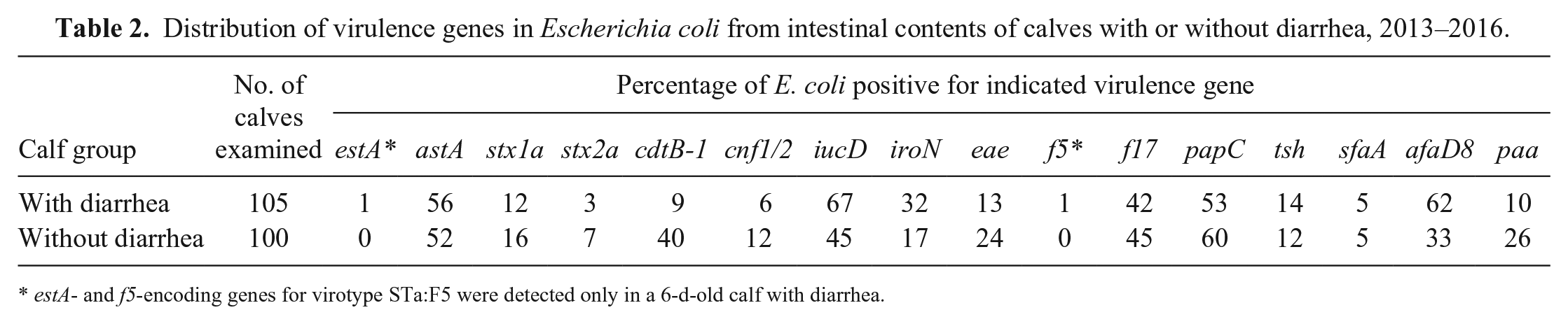
estA- and f5-encoding genes for virotype STa:F5 were detected only in a 6-d-old calf with diarrhea.
At autopsy, calves with diarrhea were dehydrated, and the intestinal tract was filled with watery contents, tainted with blood in some calves. In contrast, no significant abnormalities were observed in the intestinal tract of calves without diarrhea. Examination of histologic sections from the duodenum and jejunum of diarrheic and non-diarrheic animals did not reveal any significant findings. Most morphologic changes were observed in the ileum or colon of calves with diarrhea. Bacterial attachment to enterocytes of ileum, characteristic of ETEC, as described previously, 19 was only observed in a 6-d-old calf from which virotype STa:F5 was detected. In the remaining samples, no E. coli–like bacteria were seen attached to, or in close proximity to, enterocytes. The intestinal lesions observed in ~82% of calves with diarrhea were attributed to diarrheagenic pathogens other than E. coli (Table 3). No significant intestinal morphologic changes were observed in the intestines from 100 calves without diarrhea, despite isolation of E. coli from all of the samples, C. perfringens type A from 60% of samples, and detection of RV and BCoV in 40% of samples. These agents are highly likely to be endemic in bovine populations.7,17 Therefore, detection of one or more of these agents with current tests may not be sufficient evidence to ascribe disease status. In contrast, S. enterica ssp. enterica, Cryptosporidium spp., and Eimeria spp. were not isolated or detected in calves without diarrhea, suggesting that the presence of these pathogens as well as E. coli virotype STa:F5 may be considered for final diagnosis of the cause of diarrhea in young calves.
Morphologic diagnosis and associated diarrheagenic pathogens detected in the intestine of 105 calves with diarrhea, 2013–2016, by age range of calf.
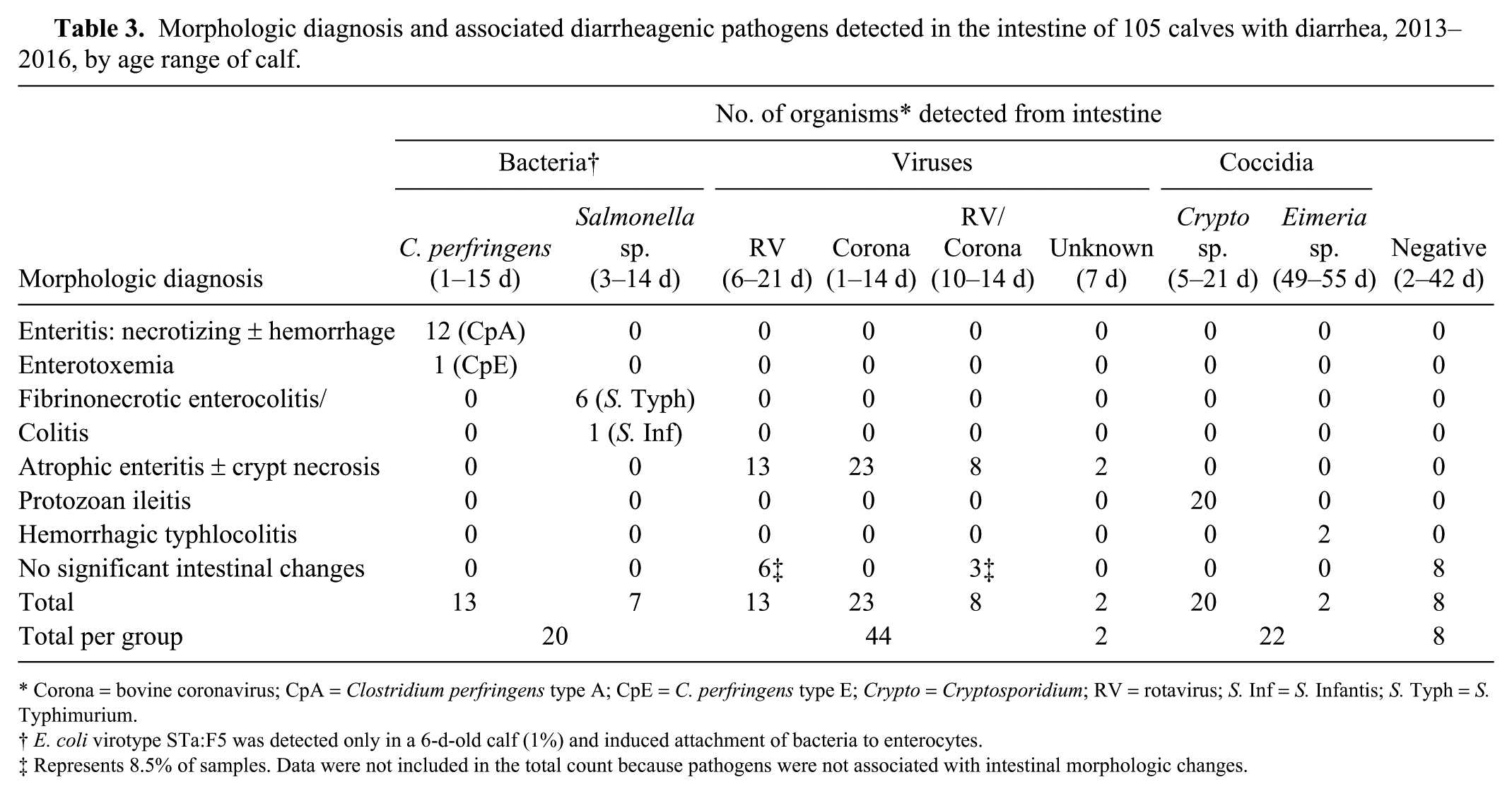
Corona = bovine coronavirus; CpA = Clostridium perfringens type A; CpE = C. perfringens type E; Crypto = Cryptosporidium; RV = rotavirus; S. Inf = S. Infantis; S. Typh = S. Typhimurium.
E. coli virotype STa:F5 was detected only in a 6-d-old calf (1%) and induced attachment of bacteria to enterocytes.
Represents 8.5% of samples. Data were not included in the total count because pathogens were not associated with intestinal morphologic changes.
E. coli virotype STa:F5 was detected only from calves with diarrhea (up to 1-wk-old). In one calf in which an autopsy was performed, typical bacterial attachment to enterocytes was observed, as commonly seen in infections with ETEC. 19 Apart from E. coli STa:F5, the other E. coli virotypes were detected at similar rates and appear randomly distributed in both groups of calves, including the most common virulence determinant combinations EAST1:F17 and EAST1:CDT:CNF:F17. Similarly, virulence genes encoding these determinants (astA, cdtB-1, cnf1/2, f17) and other genes reported in Table 2 were found with similar frequency in intestinal samples of both group of calves, and were not attached or in close proximity of the intestinal epithelium, suggesting that E. coli strains of these virotypes may not be primary pathogens in calf diarrhea.
Virulence genes cdtB-1, cnf1/2, iucD, iroN, papC, tsh, sfaA, afaD8, and paa were also detected without discrimination in both groups of calves. These genes have been reported earlier in E. coli isolates from calves with diarrhea and/or septicemia.9,20 However, there was no strong correlation between the presence of these genes and diarrhea or septicemia in the calves examined. 20 Based on our observations, it appears that E. coli virotypes other than STa:F5 may not be significant diarrhea-causing agents in calves, in contrast to what has commonly been suggested.2,18
In addition to the ETEC attachment to enterocytes seen with virotype STa:F5, another interesting finding in the calves with diarrhea was the association of intestinal lesions with other bovine diarrheagenic agents, including C. perfringens, Salmonella spp., RV, BCoV, Cryptosporidium spp., and Eimeria spp.
C. perfringens is commonly divided into 5 types (A–E). 16 We isolated C. perfringens type A in a large number of calves both with and without diarrhea. However, the organism colonized the small intestine as seen histologically and was associated with characteristic necrotizing enteritis, with or without hemorrhage, in only 12.4% of newborn to 15-d-old diarrheic animals. This suggests that C. perfringens type A can affect the gastrointestinal tract of neonatal calves and cause diarrhea, but that the diagnosis should be confirmed by histopathology.
Salmonella spp. can colonize the gastrointestinal tract of a wide range of hosts and induce a variety of clinical signs, ranging from very mild to severe enteritis. 1 Infection is most common in 1-wk- to 3-mo-old calves and manifests as acute diarrhea mainly caused by S. Typhimurium and septicemia associated with S. Dublin. 1 We isolated S. Typhimurium and S. Infantis on primary culture from 3- to 14-d-old diarrheic animals in association with characteristic fibrinonecrotic enteritis or colitis. These observations reinforce the idea that isolation of this organism on primary culture from calves with diarrhea may be sufficient to diagnose the disease.
Among the bovine enteric viruses, RV and BCoV are the major etiologic agents of diarrhea in calves. 7 RV usually causes peracute diarrhea in 1- to 2-wk-old calves. BCoV infection can manifest as 3 distinct clinical syndromes, including calf diarrhea in 1-to 2-wk-old calves; winter dysentery with hemorrhagic diarrhea in adult animals; and respiratory disease in both young and adult cattle. 7 Viral enteritis, with typical atrophic enteritis with or without crypt necrosis 19 was associated with RV and/or BCoV, in 44 of 105 cases (Table 3). However, no morphologic changes were observed on histopathology of 9 of 105 calves with diarrhea and also in 40% of calves without diarrhea, despite the detection of viruses in these animals. Hence, similar to the observations made for C. perfringens, it appears that detection of RV and BCoV alone may not be sufficient to determine disease status. Therefore, diagnosis of diarrhea caused by these viruses should also be confirmed by histopathology.
Cryptosporidium spp. and Eimeria spp. were detected only in calves with diarrhea. Cattle are commonly infected by C. parvum, C. bovis, C. ryanae, and C. andersoni as well as E. zuernii, E. bovis, and E. auburnensis.7,8 We detected Cryptosporidium spp. on routine flotation, on histology of the small intestine, and in association with protozoan ileitis, as described elsewhere, 19 in 19.0% of 1- to 3-wk-old calves. Eimeria spp. were also detected and were associated with hemorrhagic typhlocolitis in a few calves (1.9%) >1-mo-old. Although we did not speciate these parasites, our data suggest that their presence may be considered in the differential diagnosis of diarrhea in young calves.
In the few remaining cases of diarrhea examined (7.6%), there were no significant intestinal morphologic changes and no infectious agents detected, suggesting that diarrhea in this group of calves may be noninfectious and may be attributed to other factors as suggested elsewhere. 5
Overall, our data show that E. coli, excluding STa:F5, does not seem to play a significant role in diarrhea of calves, as has been commonly suggested. Testing or detection of the E. coli STa:F5 virotype may be sufficient for routine diagnosis to rule out neonatal colibacillosis in calves <7-d-old. For other diarrheagenic pathogens, such as C. perfringens, RV, and BCoV, simple detection of these organisms from calves with diarrhea may not be sufficient to ascribe disease status, but should be confirmed by histopathology.
Supplemental Material
DS1_JVDI_10.1177_1040638719857783 – Supplemental material for Frequency of Escherichia coli virotypes in calf diarrhea and intestinal morphologic changes associated with these virotypes or other diarrheagenic pathogens
Supplemental material, DS1_JVDI_10.1177_1040638719857783 for Frequency of Escherichia coli virotypes in calf diarrhea and intestinal morphologic changes associated with these virotypes or other diarrheagenic pathogens by Musangu Ngeleka, Dale Godson, Ghyslaine Vanier, Gabriel Desmarais, Chris Wojnarowicz, Soraya Sayi, Yanyun Huang, Rambod Movasseghi and John M. Fairbrother in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Dr. Brendan O’Connor and Marilyn Jonas for guidance with the research proposal, and the Bacteriology, Parasitology, Virology and Molecular Diagnostics staff of Prairie Diagnostic Services for technical assistance.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This project was funded in part by the Saskatchewan Agriculture Development Fund (ADF 20120271) and Alberta Livestock and Meat Agency (ALMA 2013R044R).
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
