Abstract
We developed a multiplex reverse-transcription real-time PCR (RT-rtPCR) assay for the simultaneous detection of the main equine respiratory viruses: equid alphaherpesviruses 1 and 4 (EHV-1, -4) and equine influenza virus (EIV; species Influenza A virus). The primers and probes amplified only the targeted viruses, and there were no inter-assay cross-amplifications or nonspecific interactions. The multiplex assay efficiencies were 92.5%, 97%, and 90% for EHV-1, EHV-4, and EIV, respectively. The R2 values of the monoplex and multiplex assays were ⩾0.990, and the slopes were −3.37 to −3.59. The performance of the assay was evaluated by analyzing 152 samples from clinically infected horses. EHV-1 DNA was detected in 12 samples, EHV-4 DNA in 9 samples, and both EHV-1 and EHV-4 in 4 samples. The accuracy of the assay was confirmed by comparing these results using commercial rtPCR and RT-rtPCR kits. Our multiplex RT-rtPCR was a sensitive, specific, accurate, and cost-effective method for the detection of the target viruses whether they occur alone or as part of coinfections.
Equine viral respiratory disease is a major problem for the equine industry and causes severe economic losses. The most important viruses that cause respiratory diseases are equid alphaherpesviruses 1 and 4 (EHV-1, -4), equine influenza virus (EIV; species Influenza A virus), equine arteritis virus, equine rhinitis A virus, erbovirus A (formerly equine rhinitis B virus), and equine mastadenoviruses A and B (formerly equine adenovirus 1 and 2, respectively). The major viruses that contribute to equine viral respiratory disease are EHV-1, EHV-4, and EIV. 14 In Egypt, both EHV-1 and EHV-4 have been isolated. 1 Additionally, EIV has been reported and isolated in 2 outbreaks: H7N7 in 1989 9 and H3N8 in 2008. 7
Real-time PCR (rtPCR), one of the most commonly applied molecular detection techniques, is characterized by accurate and rapid detection of infectious agents while reducing the risk of cross-contamination that commonly occurs in conventional PCR assays. Initially, rtPCR was dependent on the use of intercalating dyes, which were used as the detection method. 18 SYBR Green I has been used in the development of rtPCR for the detection of EHVs 3 and in development of reverse-transcription rtPCR (RT-rtPCR) assays for EIV. 15 However, these dyes are not sequence-specific and bind nonspecifically to any double-stranded DNA. Fluorescent-labeled probes were introduced as specific detection systems for rtPCR in 1996. 5 Additionally, hydrolysis probes were used to target the glycoprotein B gene of EHV-1 8 or the minor groove binding (MGB) probe of the same gene. 3 On the other hand, a RT-rtPCR hydrolysis probe assay that targets the matrix (M) gene of influenza A viruses was developed and used as the main molecular detection technique during the Sydney 2007 equine influenza outbreak. 10 In addition, other specific RT-rtPCR hydrolysis probe assays have been applied to target the hemagglutinin (HA) gene of the H3N8 subtype 4 or a wide range of EIV strains, including H3N8 and H7N7 subtypes. 13
Multiplexing in rtPCR using labeled probes with fluorescent dyes is achieved by different dyes that have sufficiently different excitation and emission wavelengths to detect multiple targets. 18 A multiplex rtPCR for detection and genotyping of EHV-1 and EHV-4 has been developed. 3 Multiplex rtPCR assays have been developed for the detection of viruses that cause bovine respiratory diseases, 17 neurologic pathogens that cause meningoencephalitis in dogs, 6 and pathogens that cause respiratory disease in small ruminants. 16
We aimed to develop and optimize a multiplex RT-rtPCR assay for the detection of EHV-1, EHV-4, and EIV. We evaluated the sensitivity and specificity of the assay and its application in the detection of single or dual infections.
Isolates of EHV-1 (AIV) and EHV-4 (V01-3-13) were provided by the Institute of Virology (Berlin, Germany). EIV (A/Equine/Egypt/VRLCU/2008) was obtained from the Virology Department, Faculty of Veterinary Medicine, Cairo University (Egypt). Nucleic acids of the isolates were extracted (Allprep DNA/RNA mini kit, Qiagen, Hilden, Germany) according to the manufacturer’s protocol. Purified DNA and RNA were stored at −80°C until use. Primers and probes for EHV-1and EHV-4 3 were selected to target the glycoprotein B gene, which is a highly conserved region in equine herpesviruses and enables differentiation of viral strains. The selected primers and probe for EIV 13 target the conserved region of the M gene. Probes were used with modification of reporter and quencher dyes. Hexachlorofluorescein (HEX), cyanine 5 (Cy5), and 6-carboxyfluorescein (FAM) fluorescent dyes were selected for labeling of EHV-1, EHV-4, and EIV probes, respectively. The spectra of the selected dyes are sufficiently different to detect them separately, reducing the crosstalk phenomenon in the multiplex assay. Quenchers were also modified to be compatible with reporter dyes. Primers and probes were manufactured (Metabion International, Planegg, Germany; Table 1). The sequence specificities of the selected primers and probes were confirmed by nucleotide–nucleotide BLAST searches in GenBank (http://www.ncbi.nlm.nih.gov/BLAST/).
Sequences of primers and probes used in the multiplex RT-rtPCR for equine respiratory viruses.
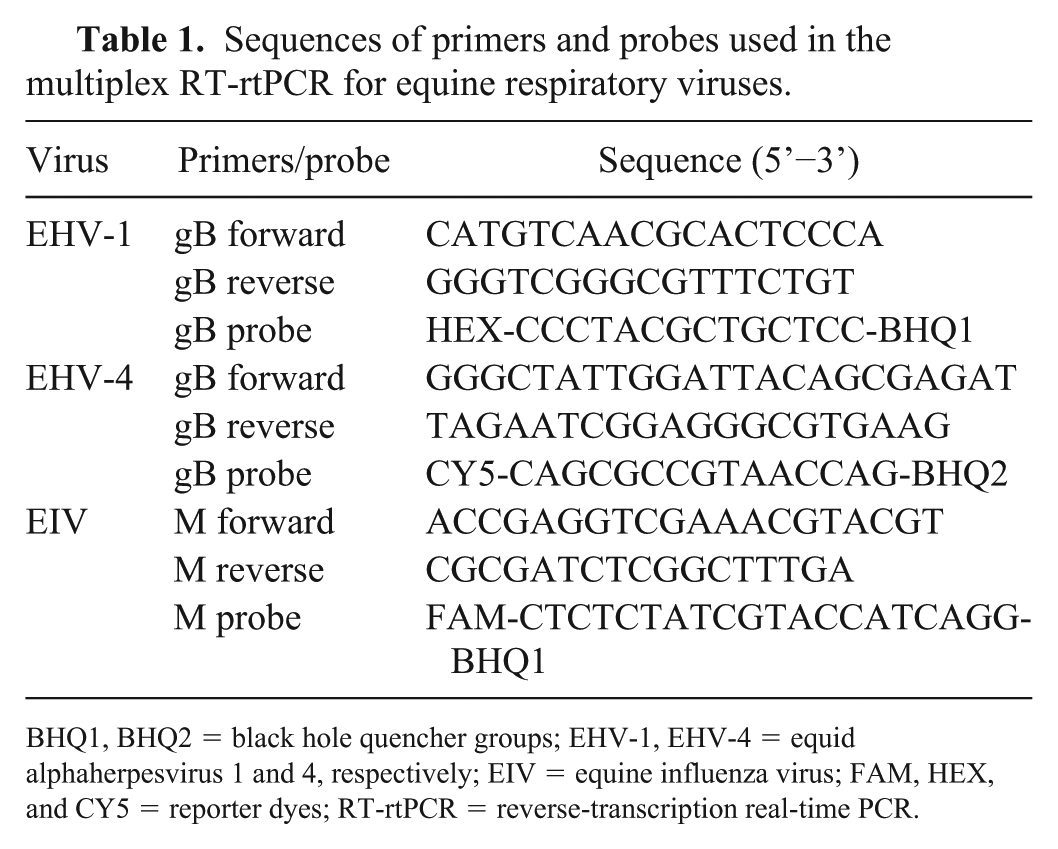
BHQ1, BHQ2 = black hole quencher groups; EHV-1, EHV-4 = equid alphaherpesvirus 1 and 4, respectively; EIV = equine influenza virus; FAM, HEX, and CY5 = reporter dyes; RT-rtPCR = reverse-transcription real-time PCR.
Primer annealing temperature and probe concentration were both initially optimized in monoplex assays specific for each virus. Monoplex and multiplex RT-rtPCR assays were conducted (QuantiFast multiplex RT-PCR kit, Qiagen). Briefly, the optimized multiplex reactions were performed in a 25-μL reaction volume containing 2× QuantiFast multiplex RT-PCR master mix (HotStarTaq Plus DNA polymerase, QuantiFast multiplex RT-PCR buffer, and ROX passive reference dye), a final concentration of 500 nM of each primer, 125 nM of each probe, and 5 μL of DNA or RNA (template). All reactions were conducted (Stratagene Mx3005P QPCR, Agilent Technologies, Santa Clara, CA) with the following cycling parameters: 20 min at 50°C (RT reaction), 10 min at 95°C (hot start), 40 cycles of 15 s at 95°C (denaturation), and 30 s at 60°C (annealing and extension).
The specificity of the developed assays was evaluated using isolates of EHV-1, EHV-4, and EIV in monoplex and multiplex forms. Each isolate reacted only with its specific primers and probe generating specific amplification curves without inter-assay cross-amplifications or nonspecific interactions, confirming the specificity of the utilized primers and probes (Fig. 1). The developed multiplex assay was able to detect and differentiate the 2 closely related viruses EHV-1 and EHV-4. Furthermore, the multiplex assay detected the EIV genome after the addition of EIV-specific primers and probe. There was no preferential amplification of 1 of the targeted viruses, which ensured the capabilities of the developed assay as an upgraded method for the detection of 3 viruses that cause respiratory diseases in equids. The cycle threshold (Ct) values for each isolate were similar in monoplex and multiplex forms (Fig. 1), confirming that there is no inhibition, competition, or cross-reactivity between primers and probes in the multiplex assay.
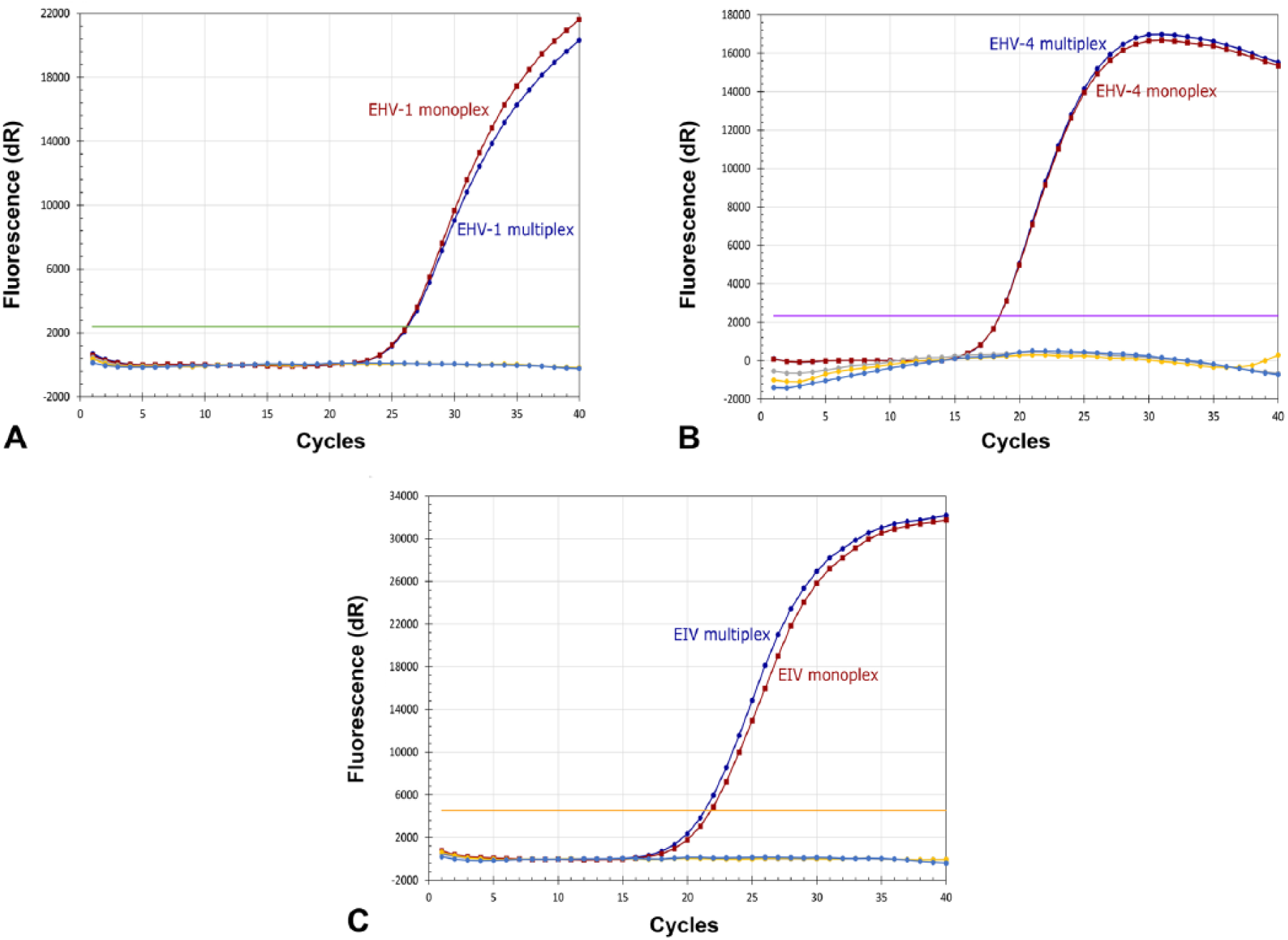
Amplification curves obtained from monoplex (red) and multiplex forms (blue) of
Streptococcus equi, which causes respiratory diseases in equids, foot-and-mouth disease virus, bovine viral diarrhea virus, lumpy skin disease virus, and sheeppox virus were used to confirm the absence of nonspecific interactions in the multiplex RT-rtPCR. S. equi was obtained from the Department of Bacteriology, Animal Health Research Institute (AHRI; Dokki, Egypt). Negative results were obtained, confirming that the primers and probes used in our assay were highly specific (Supplementary Fig. 1).
Infections by both types of EHVs have been recorded previously in equine respiratory samples. 2 Therefore, EHV-1 and EHV-4 DNA were mixed in vitro and used in a reaction. Both EHV-1 and EHV-4 DNA were detected (Fig. 2); the presence of more than one nucleic acid in the same clinical sample did not affect the assay specificity, confirming the ability of the developed assay to detect coinfections.
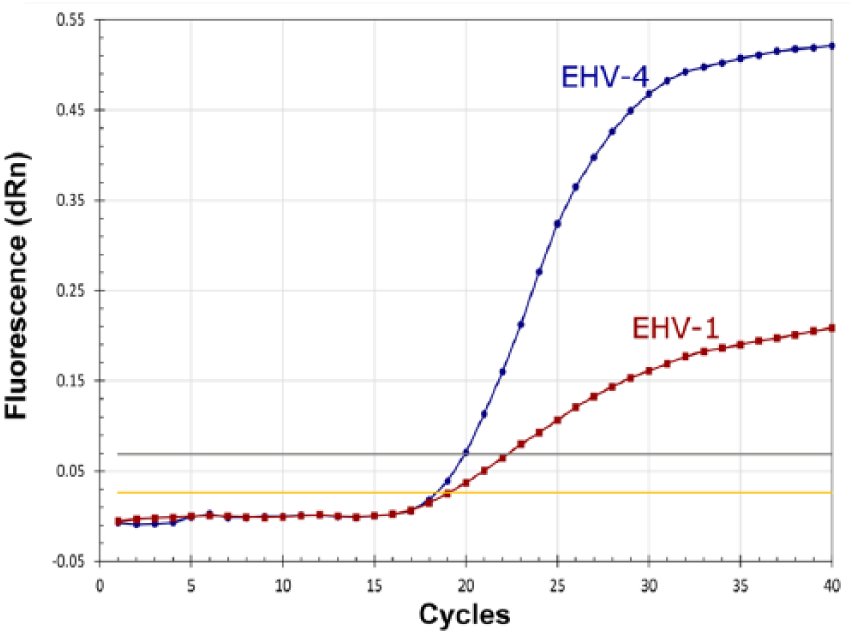
Amplification curves obtained from the multiplex RT-rtPCR for the detection of EHV-1 and EHV-4 in mixed reaction. Amplification curves for EHV-1 (red) and EHV-4 (blue) indicated that both can be detected in samples from coinfection cases.
To determine the sensitivity of our multiplex RT-rtPCR assay, EHV-1 and EHV-4 DNA and EIV complementary DNA (cDNA) were amplified, purified, and quantified as described previously 12 (QIAquick gel extraction kit, Qiagen; Invitrogen Qubit dsDNA HS assay kit, Thermo Fisher Scientific, Carlsbad, CA) for determination of the DNA and cDNA concentration, as recommended by the manufacturers. Numbers of copies for DNA and cDNA were calculated. 11 The amplified DNA of EHV-1 and EHV-4 and cDNA of EIV were serially diluted, and five 10-fold serial dilutions (105–101 copies per reaction) were amplified in triplicate to generate standard curves in monoplex and multiplex. Linear regression analysis was performed (PRISM v.7.0 for Mac, GraphPad Software, San Diego, CA). Standard curves of both forms were generated by plotting the log values of the copy numbers against the Ct values for each dilution (Fig. 3). The mean Ct values for each copy number of both forms were similar, and the obtained curves were linear in both forms for each virus. The multiplex assay could detect 101 viral copy numbers for all targeted viruses, with Ct values of 30−33 (Table 2). The monoplex assay efficiencies were 97.4%, 98.2%, and 90.7% for EHV-1, EHV-4, and EIV, respectively, and the multiplex assay efficiencies were 92.5%, 97%, and 90% for EHV-1, EHV-4, and EIV, respectively, indicating the optimal doubling of each amplicon in each cycle. The R2 in both forms were >0.990, confirming that the amplification efficiency was consistent at different template concentrations. Also, the slopes in both forms of −3.37 to −3.59 are well within the acceptable range of −3.1 to −3.6. The sensitivity of the multiplex assay was similar to the monoplex assays, which proved that multiplexing the reactions did not affect the efficiencies of the monoplex assays. 16
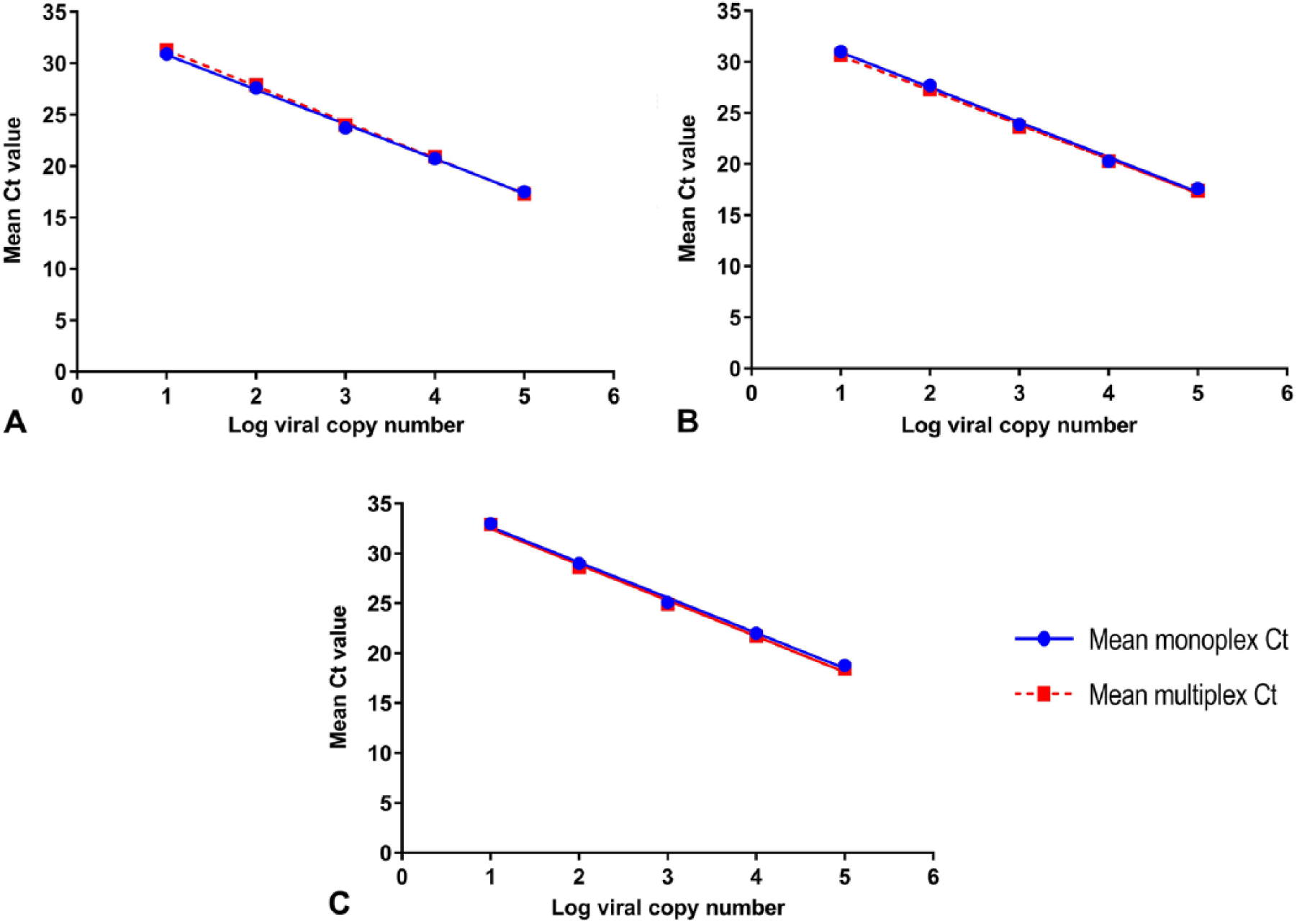
Comparison between standard curves of the monoplex and multiplex forms of
Mean Ct values obtained from monoplex and multiplex reactions in relation to the number of copies for each equine respiratory virus.
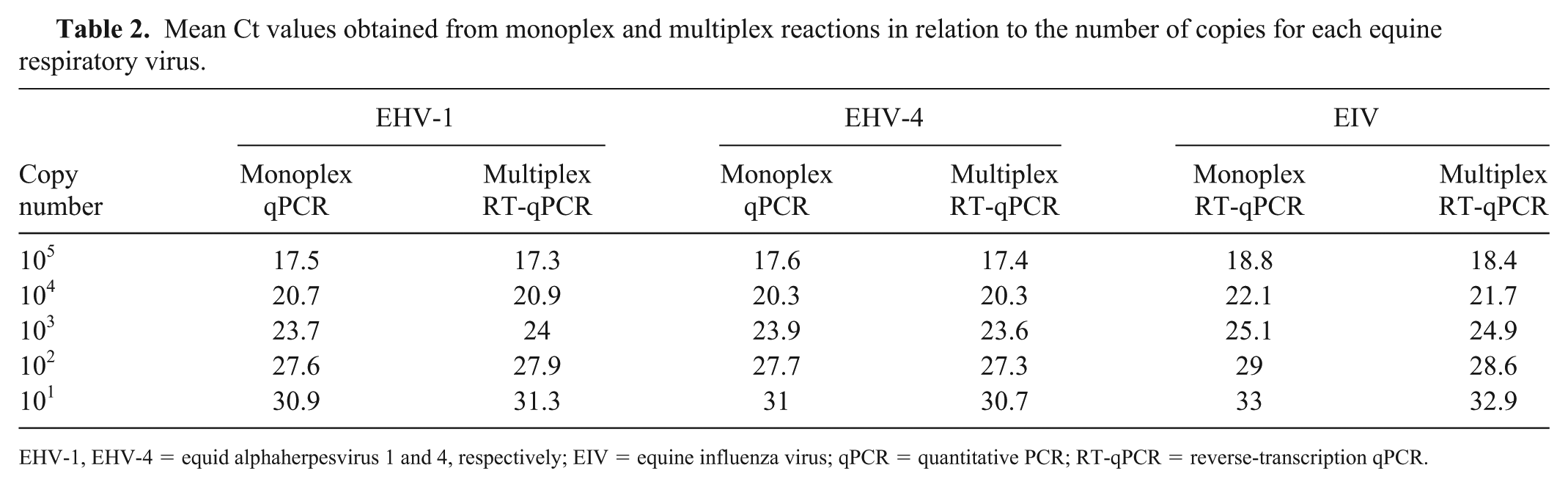
EHV-1, EHV-4 = equid alphaherpesvirus 1 and 4, respectively; EIV = equine influenza virus; qPCR = quantitative PCR; RT-qPCR = reverse-transcription qPCR.
The developed multiplex RT-rtPCR was applied to different types of clinical samples to evaluate assay efficacy. All procedures involving animals were reviewed and approved by the Institutional Animal Care and Use Committee of AHRI. Clinical samples (n = 152; nasal swabs collected from horses displaying respiratory signs; lungs and spleens of dead horses that had displayed respiratory signs; and liver of aborted fetuses) were collected from February 2014 to November 2016, from El Zahraa Stud for Arabian Horses (Cairo, Egypt) and Brooke Hospital for Animals Egypt. Additionally, 8 clinical samples known positive for EHV-1, and 3 clinical samples known positive for EIV, were tested to prove the validity of the developed assay. Nucleic acids of clinical samples were extracted (Allprep DNA/RNA mini kit, Qiagen). Commercial monoplex rtPCR and RT-rtPCR kits (Equid herpesvirus 1 genesig advanced kit, Equid herpesvirus 4 genesig advanced kit, and Equine/Canine influenza [H3N8 & H7N7] genesig standard kit, Primerdesign, Chandler’s Ford, UK) were compared with the developed multiplex RT-rtPCR assay. Reaction mixes and amplification protocols were conducted according to the manufacturer’s instructions. Of the 121 nasal swabs tested, 2 samples were positive for EHV-1, and 1 sample was positive for EHV-4 (Table 3). Of the 13 lung tissue samples tested, 8 samples were positive for EHV-4. Four of 8 spleen tissue samples and 6 of 10 fetal liver samples tested were positive for EHV-1. The developed multiplex assay detected both EHV-1 and EHV-4 in 4 samples, compared to 12 and 9 samples in which EHV-1 and EHV-4 were detected, respectively, as the single causative agent. Although EIV RNA was not detected during our study, the known-positive samples revealed the expected Ct value (Supplementary Fig. 2). These samples were analyzed using commercial monoplex rtPCR and RT-rtPCR kits; 16 samples were positive for EHV-1, and 13 samples were positive for EHV-4. Among the samples that tested positive for EHV-1, 4 samples were also positive for EHV-4. Dual infections with both EHV-1 and EHV-4 were reported (Table 3), which confirmed the prevalence of coinfections with EHVs 1 and 4 among this equine population in Egypt. 1 The multiplex RT-rtPCR assay was accurate, specific, and sensitive in the detection of the targeted viruses when compared to the results obtained with commercial rtPCR kits specific for each virus.
Results of the multiplex RT-rtPCR used for the detection of tested viruses in clinical samples.
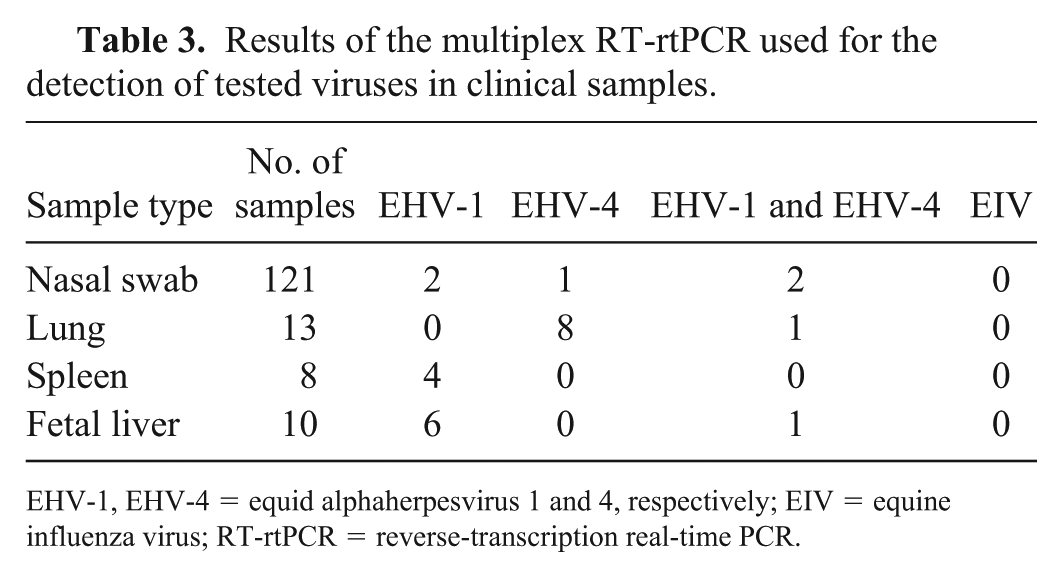

EHV-1, EHV-4 = equid alphaherpesvirus 1 and 4, respectively; EIV = equine influenza virus; RT-rtPCR = reverse-transcription real-time PCR.
The multiplex assay is more economical than performing multiple monoplex rtPCR reactions, and also reduces the time and effort spent in the laboratory. Overall, we proved the capability of the multiplex assay to detect single or coinfections by the targeted viruses. Obtaining rapid results allows prompt application of control measures to minimize the spread of disease. Our assay can be usefully applied in the surveillance of viral respiratory diseases in equids.
Supplemental Material
DS1_JVDI_10.1177_1040638718799388 – Supplemental material for Development and evaluation of a multiplex reverse-transcription real-time PCR assay for detection of equine respiratory disease viruses
Supplemental material, DS1_JVDI_10.1177_1040638718799388 for Development and evaluation of a multiplex reverse-transcription real-time PCR assay for detection of equine respiratory disease viruses by Shimaa M. Ghoniem, Ayman H. El Deeb, Mohammed G. Aggour and Hussein A. Hussein in Journal of Veterinary Diagnostic Investigation
Supplemental Material
DS2_JVDI_10.1177_1040638718799388 – Supplemental material for Development and evaluation of a multiplex reverse-transcription real-time PCR assay for detection of equine respiratory disease viruses
Supplemental material, DS2_JVDI_10.1177_1040638718799388 for Development and evaluation of a multiplex reverse-transcription real-time PCR assay for detection of equine respiratory disease viruses by Shimaa M. Ghoniem, Ayman H. El Deeb, Mohammed G. Aggour and Hussein A. Hussein in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by a grant from the Science and Technology Development Fund (STDF-RSG grant 12722).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
