Abstract
Mycoplasma bovis is an important cause of disease in cattle and bison. Because the bacterium requires specialized growth conditions, many diagnostic laboratories routinely use PCR to replace or complement conventional isolation and identification methods. A frequently used target of such assays is the uvrC gene, which has been shown to be highly conserved among isolates. We discovered that a previously described PCR putatively targeting the uvrC gene amplifies a fragment from an adjacent gene predicted to encode a lipoprotein. Comparison of the lipoprotein gene sequence from 211 isolates revealed several single nucleotide polymorphisms, 1 of which falls within a primer-binding sequence. Additionally, 3 isolates from this group were found to have a 1,658-bp transposase gene insertion within the amplified region that leads to a false-negative result. The insertion was not detected in a further 164 isolates. We found no evidence that the nucleotide substitution within the primer-binding region affects the assay sensitivity, performance, or limit of detection. Nonetheless, laboratories utilizing this method for identification of M. bovis should be aware that the region amplified may be prone to nucleotide substitutions and/or insertions relative to the sequence used for its design and that occasional false-negative results may be obtained.
Keywords
Mycoplasma bovis is an important respiratory pathogen of cattle worldwide.6,9 Although occasionally identified as a primary cause of pneumonia, it is most frequently encountered in beef cattle as a contributor to the polymicrobial syndrome bovine respiratory disease complex. Additional clinical manifestations of disease caused by M. bovis commonly observed in cattle include arthritis, conjunctivitis, otitis, and mastitis.6,9 M. bovis has also recently emerged as a pathogen of bison in which it is a primary cause of pneumonia, polyarthritis, necrotic pharyngitis, pleuritis, dystocia, and abortion, often with significant mortality.3–5,10
Isolation and identification of M. bovis is challenging given the fastidious nature of the bacterium and the need for specialized growth medium and incubation conditions.6,9 Many veterinary laboratories routinely use PCR to achieve rapid and accurate identification of the bacterium.6,9 Among the targets most frequently selected for diagnostic PCR assays is the uvrC gene, which encodes a protein involved in DNA excision and repair. The suitability of this gene for PCR-based identification of M. bovis was demonstrated using a conventional PCR method that amplifies 1,626 bp of the 1,716-bp open reading frame (ORF). 11 Other investigators confirmed the assay to be highly specific and sensitive.1,12 A comparison of the DNA sequence from the uvrC amplicon of 20 U.S. and European isolates identified only a single base polymorphism, found in a single, highly passaged laboratory strain. 12 Alternative uvrC-based PCRs have been developed, using primers placed more closely to one another so as to adapt the method for real-time (rt)PCR applications.2,8
We have routinely used the primers Mbov F2024 and Mbov R2135, originally designed for a rtPCR, 2 in a conventional PCR format to identify isolates of M. bovis acquired in our laboratory or to confirm the identity of isolates provided to us by others. However, in a few instances, isolates identified as M. bovis in other laboratories either tested negatively or gave inconsistent results depending on the reaction and cycling conditions used. An unexpectedly large amplicon of >1.5 Kb was obtained with 3 of the isolates we evaluated. All isolates with negative or questionable PCR results were confirmed to be M. bovis on the basis of DNA sequence from a 1,479-bp PCR amplicon from the 16S rRNA gene, obtained and sequenced using primers F-C (5’-GGCTGTGTGCCTAATACATGC-3’) 7 and 16S-6 (5’-AGAAAGGAGGTGATCCATCCC-3’). Top-scoring hits from a BLAST search of the National Center for Biotechnology Information nonredundant nucleotide database, all 100% identical to the amplicon sequences, arise exclusively from the 16S rRNA genes of other isolates of M. bovis. Contained within the F-C/16S-6 amplicon is a 770-bp region that has been shown to be sufficient for species assignment. 7 Furthermore, all isolates tested positively using an M. bovis–specific PCR that amplifies 1,626 bp of the uvrC gene. 11 Additional investigation revealed that the rtPCR primers F2024 and Mbov R2135 2 do not amplify a portion of the uvrC gene, as stated in the original study. Rather, they are positioned within a 405-bp ORF immediately downstream of uvrC and in the opposite orientation (locus tag MBOVPG45_0313 for the type strain, PG45; Fig. 1). The GenBank annotation for MBOVPG45_0313 is “putative lipoprotein.” Because we noted that several polymorphisms are present in corresponding sequences found in GenBank from other isolates, and considering our prior false-negative results, these observations raised concerns as to whether the PCR target is suitable for identification of all strains.
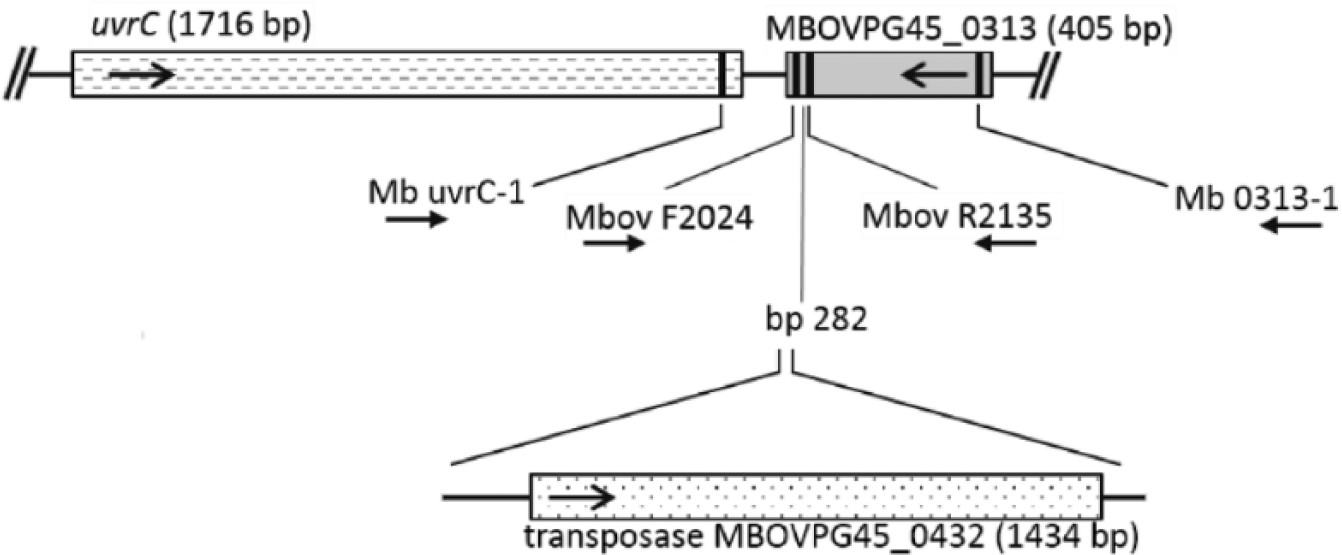
Schematic representation of the uvrC gene and the predicted lipoprotein gene immediately adjacent (locus tag MBOVPG45_0313), as found in the type strain, PG45, with the positions of PCR primers used in our study indicated by vertical bars. Primers Mbov F2024 and Mbov R2135 are those used for the rtPCR of reference 2. Arrows specify the direction of transcription (for genes) or amplification (for primers). The position at which a transposase insertion occurs in some isolates is indicated (bp 282 of the lipoprotein gene open reading frame, relative to the first nucleotide of the start codon).
To examine in detail the DNA sequence heterogeneity within the Mbov F2024/Mbov R2135 amplicon and, in particular, the probe- and primer-binding sites, we used PCR primers Mb uvrC-1 (5’-CTATGACGCTTCAGTTGAAGA-3’) and Mb 0313-1 (5’-GCTTTTGTCAATTGCTTCAA-3’), ob-tained commercially (Integrated DNA Technologies, Coralville, IA), to amplify and sequence a 486-bp region encompassing the Mbov F2024/Mbov R2135 amplicon (Fig. 1) from 54 M. bovis isolates (37 from cattle and 17 from bison). The M. bovis isolates were grown in PPLO broth (pleuropneumonia-like organism broth; Becton Dickinson, Sparks, MD) for 24–48 h at 37°C with 5% CO2. A 1-mL aliquot of each culture was centrifuged for 10 min at 10,000 × g. Pelleted cells were resuspended in 200 µL of PBS, and DNA was extracted (QIAamp DNA blood mini kit, Qiagen, Valencia, CA) according to the protocol for blood or body fluids. The concentration of DNA in each preparation was determined fluorometrically (Qubit dsDNA HS assay kit, Thermo Fisher Scientific, Waltham, MA). Each PCR contained ~2 ng of purified DNA, 200 mM of each dNTP, 0.5 µM of each primer, 1.5 mM of MgCl2, 2 U of polymerase (AmpliTaq DNA polymerase, Applied Biosystems, Carlsbad, CA), and 5.0 µL of 10× Buffer II in a final volume of 50 µL. Cycling conditions were 94°C for 5 min and 35 cycles of 94°C for 10 s, 55°C for 30 s and 72°C for 1 min, followed by a final elongation step of 72°C for 5 min. PCR amplicons were treated to remove unincorporated primers and dNTPs (ExoSAP-IT, USB Products, Affymetrix, Cleveland, OH) and sequenced in both directions with primers Mb uvrC-1 and Mb 0313-1 at the Iowa State University DNA Facility (Ames, IA; 3730xl genetic analyzer, BigDye terminator v.3.1 cycle sequencing kit, Applied Biosystems). Final consensus sequences used for subsequent analyses represent a minimum of 2 high-quality reads, with at least 1 from each strand. DNA sequences corresponding to the Mb uvrC-1/Mb 0313-1 amplicon were retrieved from privately held draft genome sequences, assembled using Illumina MiSeq 2 × 300-bp paired-end reads, for an additional 131 isolates (117 from cattle, 13 from bison, and 1 from a white-tailed deer) and from publically available GenBank submissions for a further 26 cattle isolates (Supplementary Table 1). These 211 sequences collectively represent isolates from the United States, Canada, Europe, Israel, China, and Australia.
The Vector NTI Advance 11 software package (Thermo Fisher Scientific) was used for DNA sequence analysis, editing, and alignment. An alignment of the Mb uvrC-1/Mb 0313-1 amplicon sequences from the 211 isolates included in our study revealed 6 unique variants with 2–8 nucleotide substitutions and a large insertion compared to the type strain, PG45, whose sequence was designated as variant 1 (Fig. 2, Table 1). All variants have sequences identical to the rtPCR probe (Mbov uvrC) and the primer Mbov R2135, 2 but variants 2–6 have a single nucleotide substitution in the 3’ portion of the primer Mbov F2024. Variant 6 additionally has an insertion of 1,658 bp positioned between Mbov F2024 and Mbov R2135, thus explaining the larger-than-expected amplicon occasionally obtained with these primers, as noted above. Additional primers used to sequence the insertion included Mb tpase-1 (5’-TGAAACACCTGTCCCAGTA-3’), Mb tpase-4 (5’-CATCCTCTAATGAAAGGTCAACA-3’), Mb tpase-5 (5’-GGCAAAGAAAGTGTTGGAAAC-3’), and Mb tpase-6 (5’-GGATAAGCACATCTAAGAATAGCA-3’). Top-scoring BLAST matches with the 1,658-bp insertion are 99% identical to a region found in M. bovis isolates PG45 and CQ-W70 comprised of a 1,434-bp transposase (locus tags MBOVPG45_0432 and K668_02140, respectively) and short stretches of adjacent 5’ and 3’ sequence (Fig. 1).

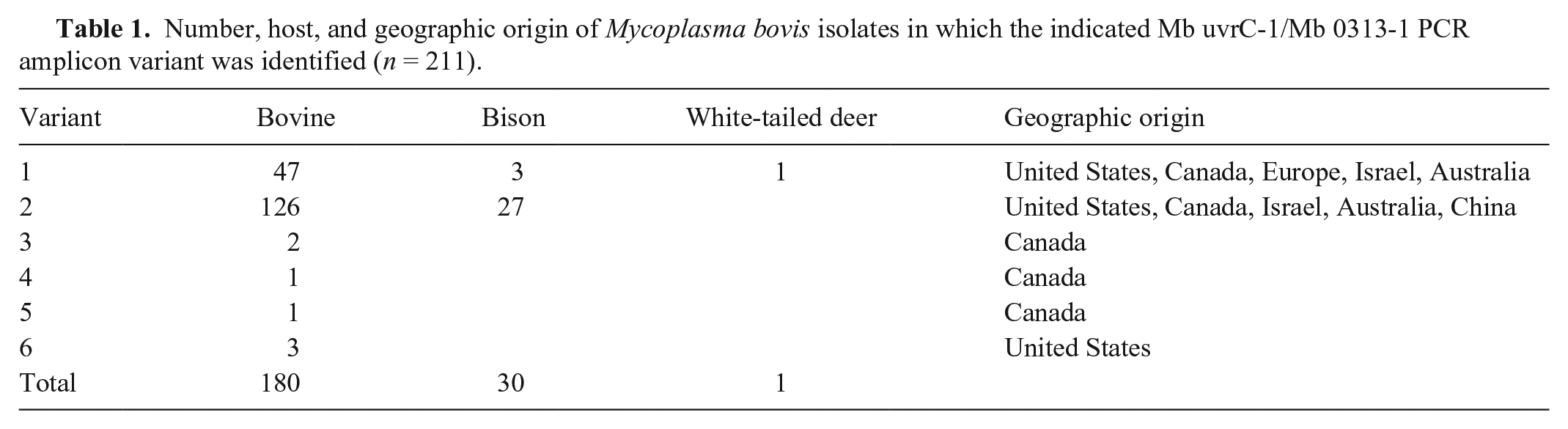
Nucleotide alignment of M. bovis uvrC-1/Mb 0313-1 amplicon variants identified in our study. Variant 1 is found in the type strain, PG45, used to design the PCR assay reported by reference 2. PCR primer sequences and the rtPCR probe are in bold and underlined; open reading frames (ORFs) are shaded in gray. Arrows specify the direction of amplification (for primers) or transcription (for ORFs). The position at which a transposase insertion occurs in some isolates is indicated by an arrowhead. Coordinates for the uvrC and lipoprotein gene ORFs are relative to the first nucleotide of their start codons.
Number, host, and geographic origin of Mycoplasma bovis isolates in which the indicated Mb uvrC-1/Mb 0313-1 PCR amplicon variant was identified (n = 211).

The design of the original rtPCR 2 was based on the mistaken assumption that the amplified target is within the uvrC gene, known to be stable and highly conserved among isolates. The discovery that the target falls within a nearby lipoprotein gene in which some isolates have numerous sequence polymorphisms and a large insertion raises concern as to whether false-negative results might be obtained with such isolates. To address this question, we tested 2 isolates with the G to A substitution found in the Mbov F2024 primer-binding site (NADC1 and NADC15) and 2 isolates with both the Mbov F2024 substitution and the 1,658-bp transposase insertion (Cs432 and Cs6213-2). Two isolates with sequences identical to that used for the assay design (PG45 and M23) were included as positive controls. Nine reactions were run for each isolate. PCR was carried out (ABI7300 real-time PCR system, Applied Biosystems) as previously reported 2 except that 0.5 ng of purified DNA was used as the template (reference 2 does not specify the amount of DNA used). Data outputs were analyzed and threshold cycle (Ct) values were determined (ABI7300 system SDS software v.1.3.1). In addition to the positive control isolates PG45 and M23, isolates NADC1 and NAC15 tested positively with little difference in average Ct values (19.18–19.84, depending on the isolate). In contrast, negative results were obtained from both isolates with the transposase insertion, Cs432 and Cs6213-2. Similar results were obtained when broth cultures of each isolate were submitted to an AAVLD-accredited veterinary diagnostic laboratory in which the method of reference 2 is used for identification of M. bovis. Depending on the isolate, cultures contained 0.7–5.2 × 109 colony forming units/mL based on colony counts from serial dilutions, plated in triplicate on PPLO agar (Becton Dickinson). Isolates PG45, M23, NADC1, and NADC15 were reported as positive, with Ct values of 13.0–14.9; Cs432 and Cs6213-2 were both reported as negative (Ct values >40).
To assess the potential effect of the Mbov F2024 substitution on the kinetics of the rtPCR, 10-fold serial dilutions of purified DNA from PG45, M23, NADC1, and NADC15 were prepared in nuclease-free water and used to generate a standard curve for each isolate (7 dilutions per isolate), with the amount of DNA added per PCR ranging from 10 ng to 0.01 pg. Amplification efficiency was calculated as (10−1/slope − 1) × 100 (https://goo.gl/WgHoY7). The statistical features of the standard curves indicate that the substitution has little effect on the performance of the primer set or on the amplification efficiency (Table 2). The limit of detection (LOD) appears to be similarly unaffected by the Mbov F2024 substitution, based on a comparison of the proportion of positive results from 9 replicate reactions per isolate for each of 4 DNA dilutions (0.5 ng–0.5 fg of DNA per reaction; Table 2). These data indicate that, under the conditions used in our study, the LOD is 5–50 fg per reaction, regardless of whether an isolate has the substitution. This corresponds to ~4.7–47 genome equivalents assuming a genome size of 0.983 Mb, based on the average size of the complete genome sequences available to date for M. bovis (GenBank accessions CP002188.1, CP002058.1, CP019639.1, CP011348.1, CP005933.1, and CP002513.1).
Kinetics and limit of detection of the real-time PCR from reference 2, using DNA extracted from broth cultures of Mycoplasma bovis.
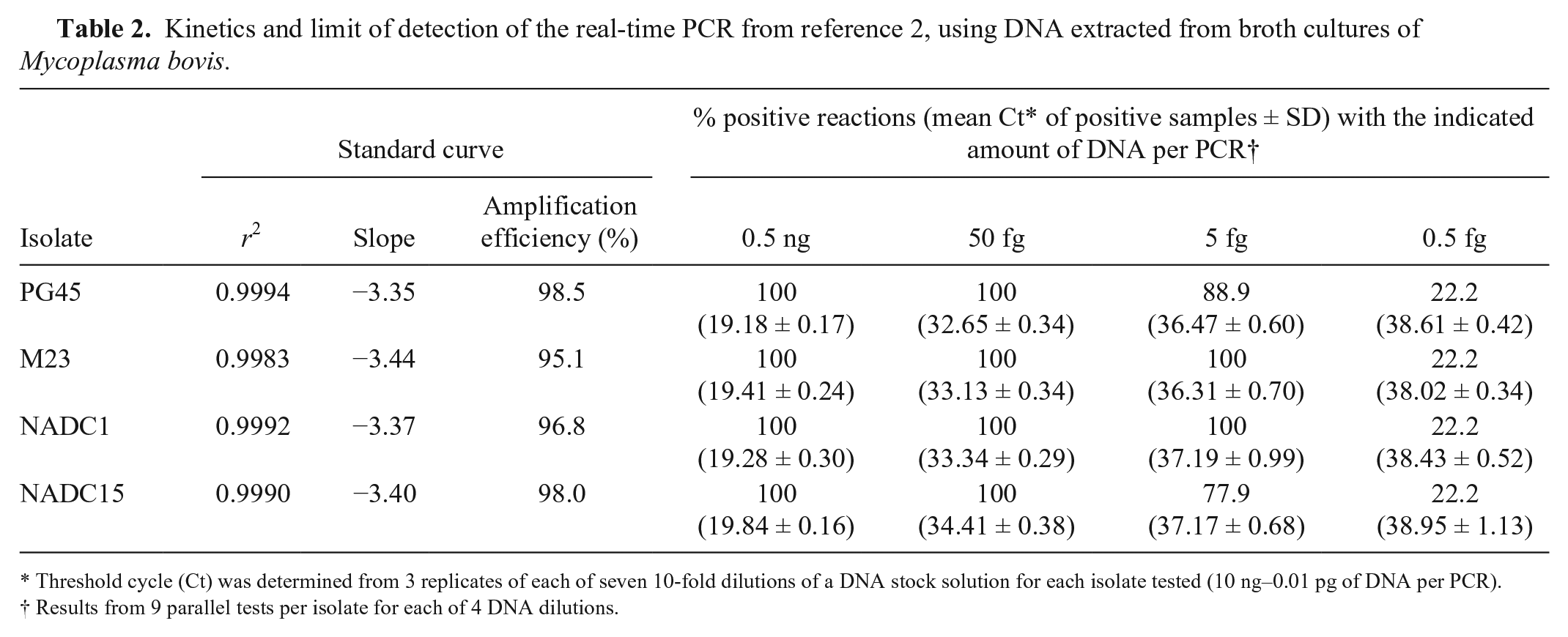
Threshold cycle (Ct) was determined from 3 replicates of each of seven 10-fold dilutions of a DNA stock solution for each isolate tested (10 ng–0.01 pg of DNA per PCR).
Results from 9 parallel tests per isolate for each of 4 DNA dilutions.
Together with the 211 isolates of M. bovis evaluated in our study, an additional 164 isolates obtained for other purposes have also been tested in our laboratory in rtPCR and/or conventional PCR using primers Mbov F2024 and Mbov R2135. Of those 375 isolates, false-negative results were obtained only with the 3 reported herein, suggesting these primers are highly sensitive with respect to the detection of current or historical isolates. Nonetheless, these false-negative results are of concern given the importance of early recognition in controlling disease and because the lesions associated with bovine mycoplasmosis can be mistaken for lesions of tuberculosis. 9 Data reported herein further demonstrate that 160 of 211 isolates from which we obtained DNA sequences for the region including the corresponding amplicon possess one or more nucleotide substitutions compared to the sequence of isolate PG45, used to design the PCR, one of which falls within a primer-binding region. Although we found no evidence that the nucleotide substitution in the primer-binding region affects the assay sensitivity, performance, or LOD, laboratories utilizing the method of reference 2 for identification of M. bovis, or the primers described therein, should be aware that the targeted sequence may be prone to nucleotide substitutions and/or insertions and that occasional false-negative results may be obtained.
Supplemental Material
DS1_JVDI_10.1177_1040638718764799 – Supplemental material for Mistaken identity of an open reading frame proposed for PCR-based identification of Mycoplasma bovis and the effect of polymorphisms and insertions on assay performance
Supplemental material, DS1_JVDI_10.1177_1040638718764799 for Mistaken identity of an open reading frame proposed for PCR-based identification of Mycoplasma bovis and the effect of polymorphisms and insertions on assay performance by Karen B. Register, William D. Boatwright, Karen M. Gesy, Tyler C. Thacker and Murray D. Jelinski in Journal of Veterinary Diagnostic Investigation
Footnotes
Authors’ note
Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
