Abstract
Ornithobacterium rhinotracheale (ORT) is an infectious respiratory pathogen of chickens, turkeys, and wild birds. There are 18 serotypes of ORT reported worldwide. In this study, enterobacterial repetitive intergenic consensus (ERIC) polymerase chain reaction and random amplified polymorphic DNA assay with Universal M13 primer-based fingerprinting techniques were investigated for their ability to differentiate ORT isolates. The authors examined 50 field isolates and 8 reference strains of ORT for their genetic differences. The fingerprint patterns were compared with serotyping results of ORT by the agar gel precipitation test. M13 fingerprinting revealed different patterns for 6 reference serotypes of ORT that were tested, namely, C, D, E, I, J, and K. Ornithobacterium rhinotracheale reference serotypes A and F yielded indistinguishable fingerprints with M13 fingerprinting. The ERIC 1R technique discerned only 5 of the 8 reference serotypes of ORT. Distinct fingerprints were also found within the ORT serotypes with both techniques. From 58 isolates of ORT that were fingerprinted belonging to 8 ORT serotypes, 10 different fingerprints were obtained with M13 fingerprinting and 6 different fingerprints were obtained with ERIC 1R fingerprinting. M13 fingerprinting technique was found to be more discriminative in differentiating ORT isolates than the ERIC 1R fingerprinting technique. These results suggest that fingerprinting techniques may be a more discerning tool for characterizing ORT isolates than the serological test using the agar gel precipitation test. This fingerprinting technique could potentially be a valuable tool in identifying an isolate from a clinical outbreak of ORT infection for development of an autogenous vaccine.
Keywords
Ornithobacterium rhinotracheale (ORT) is a gram-negative bacterium that has been identified as an emerging respiratory pathogen in turkeys and chickens. 2–4 The organism has been reported worldwide and has also been isolated from wild birds. 2 The disease caused by ORT is characterized by tracheitis, airsacculitis, and fibrinous pneumonia in severely affected birds. 3,9 Mortality has been reported to be up to 10% in ORT-infected flocks, and the high carcass condemnation rates due to airsacculitis at processing plants lead to marked economic loss to producers. 11 There are 18 serotypes of ORT, designated by the letters A through R. 4 At the present time, there is no commercial ORT vaccine available in the United States, but autogenous vaccines are being used by many turkey producers. The current method of subgrouping ORT field isolates for epidemiological studies and selection of candidates for autogenous vaccines is by serotyping based on the agar gel precipitation (AGP) test. This test uses monovalent antisera against the corresponding serotypes. However, the nonavailability of the antisera and cross-reactions between ORT serotypes 13 render this test more subjective and inefficient in subgrouping ORT field isolates. The distribution of enterobacterial repetitive intergenic consensus (ERIC) elements on prokaryotic genomes has been examined by polymerase chain reaction (PCR). 14 ERIC-PCR was shown to be an efficient technique to differentiate species and strains of many gram-negative bacteria. 6–8,10 For characterization of Actinobacillus pleuropneumoniae serotypes, the Universal M13 primer-based PCR was previously used and produced discriminatory DNA fingerprints among unrelated strains. 5 A repetitive-PCR (rep-PCR) fingerprinting technique using ERIC 1R and ERIC 2 primers with 55 isolates of ORT from wild birds and poultry revealed different rep-PCR patterns. 1 However, in that study, no comparison between or within the serotypes of ORT was made. Polymerase chain reaction fingerprinting helps to detect different molecular targets of bacterial isolates for diagnostic and monitoring purposes and is found to be a faster, more reliable, and more economical technique than the AGP test. 7,12 A PCR-based fingerprint analysis of ORT isolates may be more discriminative than the AGP test and could be potentially useful in selecting autogenous vaccine candidates.
The objective of this study was to address the question as to whether the PCR fingerprinting technique could be used to differentiate ORT isolates. For this study, 2 PCR fingerprinting techniques were investigated, one using a repetitive primer (ERIC 1R) and another with the Universal M13 primer to differentiate field isolates of ORT.
Fifty-eight isolates of ORT were used in this study. Eight of them were reference isolates of ORT for serotypes A, C, D, E, F, I, J, and K obtained from the Netherlands. a The other 50 isolates were isolated from clinically ill turkeys in the Midwest region, including Minnesota, between 2003 and 2006. Primary isolation was performed by streaking tracheal swabs onto 5% sheep blood agar plates supplemented with gentamycin (5 μg/ml) and incubating the plates under microaerophilic conditions (10% CO2)at 37°C for 48 hours. Suspected small, round, gray to gray-white, butyrous colonies were subcultured and tested for their catalase and oxidase reactions. 13 Catalase-negative and oxidase-positive colonies were then confirmed to be ORT by using the api-ZYM b kit. 4 Prior to fingerprinting, all ORT isolates were serotyped by the AGP test as previously described. 13 Briefly, boiled extract antigens (BEAs) were prepared by washing ORT cultures from sheep blood agar with 0.02 M of phosphate buffer containing 8.5% sodium chloride and 0.3% formaldehyde. Subsequently, the suspensions were boiled for 60 minutes at 100°C. After centrifugation, the supernatants were used as antigens in the AGP test. Agar gel precipitation glass slides were prepared using 4 ml of preheated, sterile 1.5% Noble agar and 8.5% sodium chloride on standard glass slides. Six wells with a diameter of 2 mm were punched around a center well at a distance of 5 mm on agar glass slides. Type specific sera against different ORT serotypes were placed in the center well and BEAs in the peripheral wells. The slides were incubated for 24 hours in a moist chamber at 37°C and observed for line of precipitation under ultraviolet light.
For PCR, a single colony per isolate was taken from the respective plate with a sterile loop and mixed with 300 μl of phosphate-buffered saline (PBS). The bacterial suspension was then vortexed for 1 minute and centrifuged at 14,000 × g for 3 minutes. The supernatant was completely discarded. The pellet was resuspended in 200 μl of Prepman-Ultra, c boiled at 100°C for 10 minutes, and again centrifuged at 15,000 × g for 3 minutes. Fifty microliters of the supernatant containing bacterial DNA was diluted with 50 μl of sterile water. The DNA concentration was measured by spectrophotometry d at 260 nm. Stock solutions of bacterial DNA were adjusted to a concentration of 100 ng/5 μl for PCR. The primer ERIC 1R (5'ATG TAA GCT CCT GGG GAT TCA C) 9 and the Universal M13 primer (5'T TAT GTA AAA CGA CGG CCA GT) 10 used in this study have been described previously. Both of these primers were obtained e and used to amplify 100 ng of ORT DNA for each PCR fingerprinting reaction. The PCR was performed using a modified protocol. 1 Twenty-five micro-liters of PCR reaction was prepared in sterile water using 5 μl of PCR Buffer I, c 3.5 mM of MgCl2, 4 μM of primer ERIC 1R or M13, 200 μM each of deoxy-ribo-nucleotide phosphates, c 0.25 IU of AmpliTaq Gold DNA polymerase, c and 5 μl (100 ng) of template DNA. The reactions were carried out in a thermal cycler. f The PCR conditions included an initial denaturation at 95°C for 10 minutes followed by 35 cycles of denaturation at 94°C for 45 seconds, annealing at 40°C for 3 minutes, and extension of 72°C for 4 minutes, with a final extension step at 72°C for 10 minutes. An 8-μl volume of PCR products was mixed with 2 μl of loading buffer, and the mixture was electrophoresed at 65 V in a 2% agarose gel with 0.5 μg/ml of ethidium bromide. Gels were photographed under ultraviolet light using a gel imaging system. g To confirm the reproducibility of the technique, 2 independent reactions were conducted for each isolate. The images obtained from each fingerprint were subjected to visual inspection and interpretation.
The reference isolates were confirmed by the AGP test as ORT serotypes A, C, D, E, F, I, J, and K. Of the 50 ORT field isolates, 32 isolates were serotyped as ORT serotype A, 11 as serotype C, and 7 as ORT serotype I. Ornithobacterium rhinotracheale serotypes A, C, and I are the most prevalent serotypes in turkeys in Midwest states (Nagaraja, unpublished data).
Reference ORT serotypes C, D, E, I, J, and K produced different fingerprints with the M13 PCR technique, whereas ORT serotype A and F yielded indistinguishable fingerprints (Fig. 1). On the other hand, the ERIC 1R PCR for the 8 reference serotypes yielded only 5 different fingerprints. The ERIC 1R fingerprints were distinct for ORT serotypes C, D, E, and K, respectively. However, ORT serotypes A, F, I, and J yielded indistinguishable fingerprints (Fig. 2). From the 58 ORT isolates fingerprinted, 9 different fingerprint patterns were obtained with M13 fingerprinting. Only 6 different fingerprinting patterns were obtained with ERIC 1R fingerprinting. It appears, therefore, that the M13 fingerprinting technique is more useful than the ERIC 1R fingerprinting for discriminating ORT isolates. Unfortunately, neither method was completely serotype discriminative, as all ORT serotypes could not be differentiated. This is in contradiction with previous study results, in which both M13 and ERIC 1R fingerprinting were found to be effective in differentiating all the ORT serotypes tested. (Hafez HM, Beyer W: 1997, Proceedings of 11th International Congress of the World Poultry Association, Budapest, Hungary). This discrepancy could be due to the variation in the source and number of ORT isolates tested.
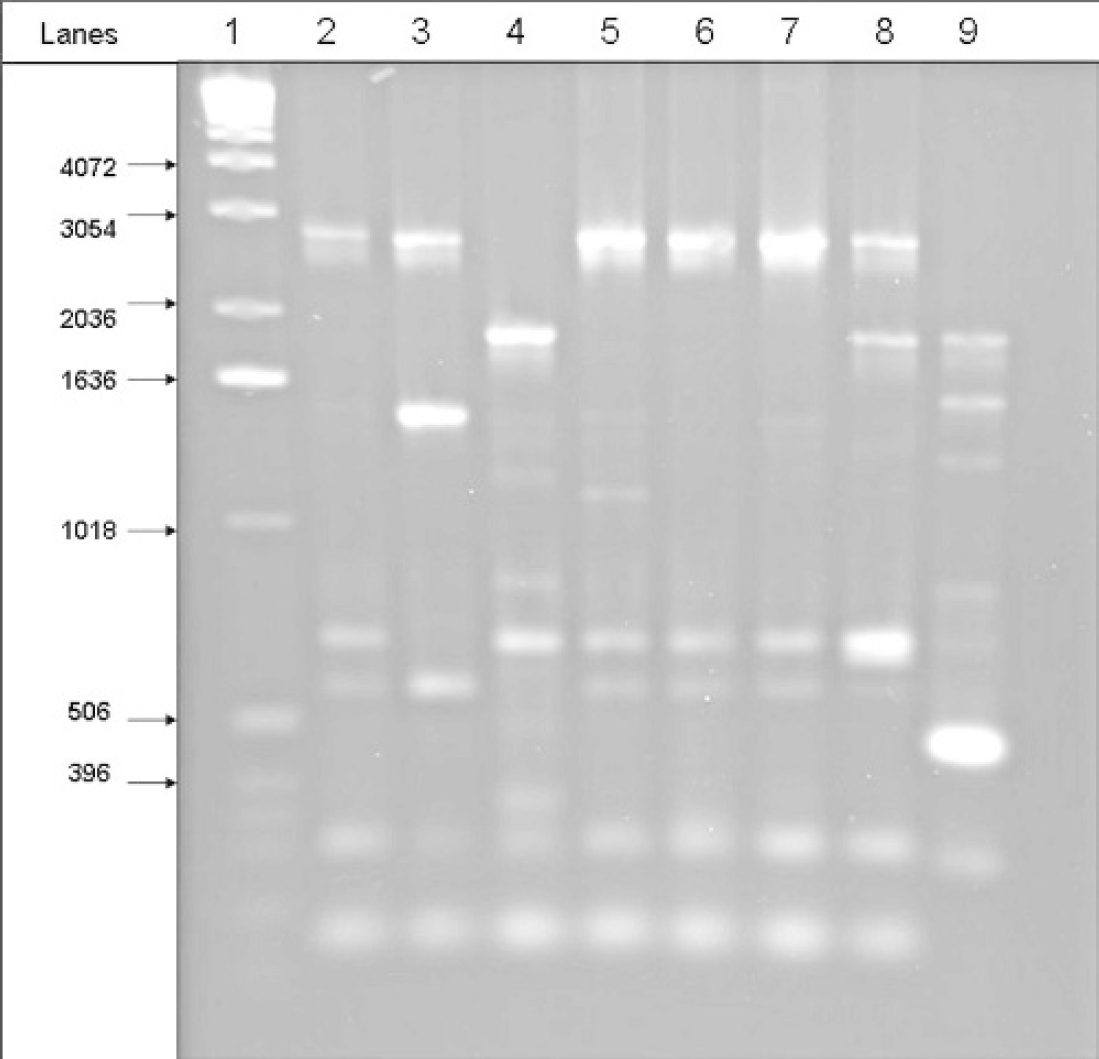
M13 fingerprints of ORT reference serotypes. Lane 1 is a 1-Kb DNA ladder; lanes 2–9 represent ORT reference serotypes A, C, D, E, F, I, J, and K, respectively.
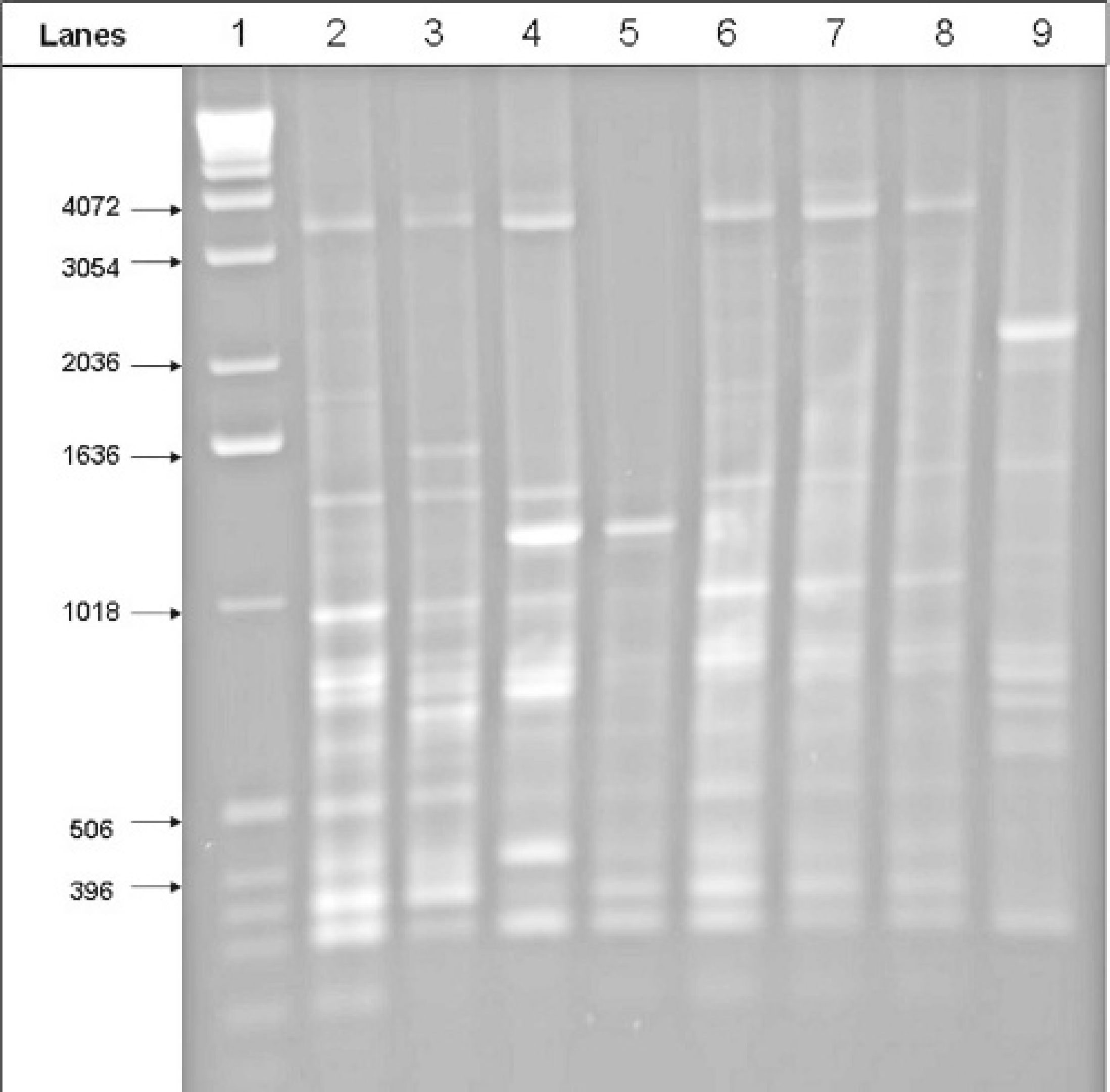
ERIC 1R fingerprints of ORT reference serotypes. Lane 1 is a 1-Kb DNA ladder; lanes 2–9 represent ORT reference serotypes A, C, D, E, F, I, J, and K, respectively.
One of the most important results of this study was the detection of distinct fingerprint patterns within each ORT serotype. The M13 PCR of the ORT serotype A isolates revealed 3 different types of fingerprint patterns. Thirty isolates of ORT A generated the reference ORT serotype A pattern, whereas 2 isolates of ORT A gave patterns that were different from all other patterns (Fig. 3). Interestingly, ERIC 1R fingerprints obtained were indistinguishable for all the isolates of ORT serotype A. It was only with M13 fingerprinting that different fingerprint patterns for ORT serotype A isolates were obtained. Among the 11 isolates of ORT serotype C tested, only 1 isolate generated the reference ORT serotype C pattern, both in the ERIC 1R and the M13 PCR (Table 1). The ERIC 1R and M13 PCR of the remaining 10 isolates of ORT serotype C generated a pattern indistinguishable from the reference ORT serotype A pattern.
Of the 7 isolates of ORT serotype I examined with ERIC 1R PCR, 6 isolates produced a fingerprint indistinguishable from the ORT serotype A pattern. However, the remaining 1 serotype I isolate gave an ERIC 1R fingerprint pattern different from all other patterns. With the M13 PCR technique, all ORT serotype I isolates produced reference ORT serotype I fingerprint patterns. These results thus suggest the existence of genetic differences among the isolates of the same ORT serotype.
Once the strain involved in an ORT outbreak has been identified by serotyping, it is common practice to use the standard reference strain to produce an autogenous vaccine. However, the results reported herein indicate that genetic differences within the same serotypes may exist. Genetic differences may also contribute to antigenic differences, and an ORT isolates that have indistinguishable fingerprints within a serotype would be ideal autogenous vaccine candidates.
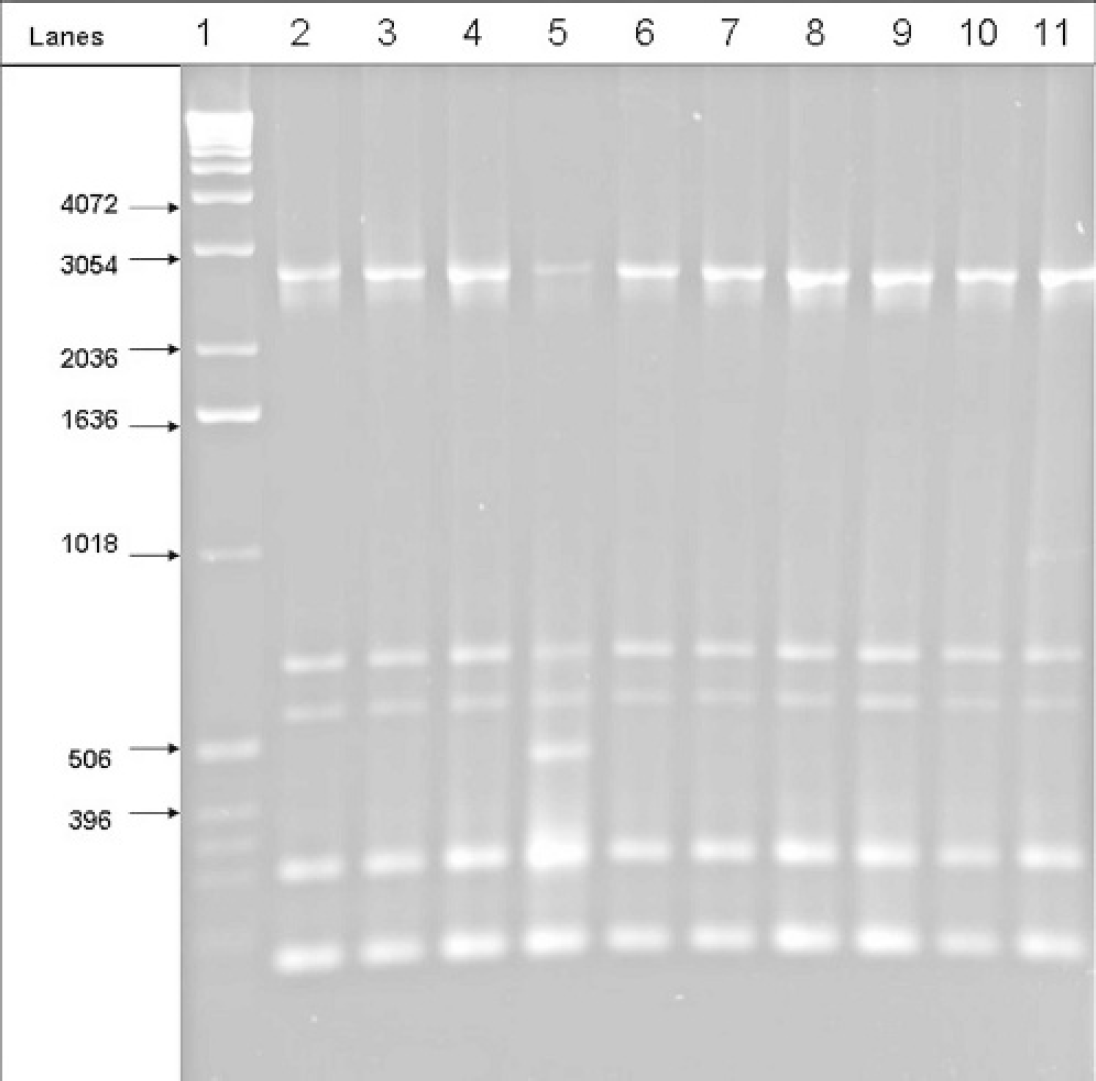

M13 fingerprints of ORT serotype A isolates. Lane 1 is a 1-Kb DNA ladder. Isolates in lane 5 and lane 11 yielded different fingerprint patterns.
In the present study, no cross-reactions with any of the ORT isolates tested were found in the AGP test. The cross-reactions in the AGP test could possibly be due to the presence of some shared antigens among ORT serotypes. The presence of more than 1 precipitating antigen and cross-reactions in the AGP test among ORT serotypes A, B, D, and E have been previously reported, indicating that the AGP test might not be able to serotype some field isolates of ORT. But no cross-reactions were reported between ORT A, I, and C serotypes in the AGP test. 13 Previous findings (Nagaraja, unpublished data) from more than 200 US field isolates of ORT tested by AGP did show a cross-reaction between some of the ORT A serotype strains with ORT I type-specific sera and also a few isolates with ORT C type-specific sera. Indeed, most of the fingerprints of ORT serotype C US isolates were found to be different from the reference ORT serotype C pattern and had a pattern indistinguishable from ORT serotype A. This suggests that the ORT isolates circulating in turkeys in the United States may be less heterogeneous and might have originated from a common ancestral lineage irrespective of the serotypes.
More than 1 distinct fingerprint pattern was obtained in this study within the ORT serotypes A, C, and I. Although M13 was successful in distinguishing most of the reference serotypes, both ERIC 1R and M13 were helpful in generating different fingerprint patterns within the serotypes. This suggests the possibility of further characterizing strains or isolates based on their fingerprints within the ORT serotypes for outbreak investigations 5 as well as for selection of autogenous vaccine candidates. Because the current method of serotyping of ORT by the AGP test can be subjective and because cross-reactions among serotypes are common, DNA fingerprinting techniques offer a possible means of characterizing ORT isolates. This can be achieved by M13 and ERIC 1R fingerprinting alone or in combination with the AGP test to characterize ORT isolates within the serotypes. The present results suggest that PCR subtyping methods offer promise in furthering the understanding of the epidemiology of ORT outbreaks in United States. In the near future, it might be possible to group and identify ORT isolates based on their fingerprint patterns alone; such a development would improve autogenous vaccine development.
Number of US isolates of ORT tested in each serotype and the type of fingerprinting pattern obtained
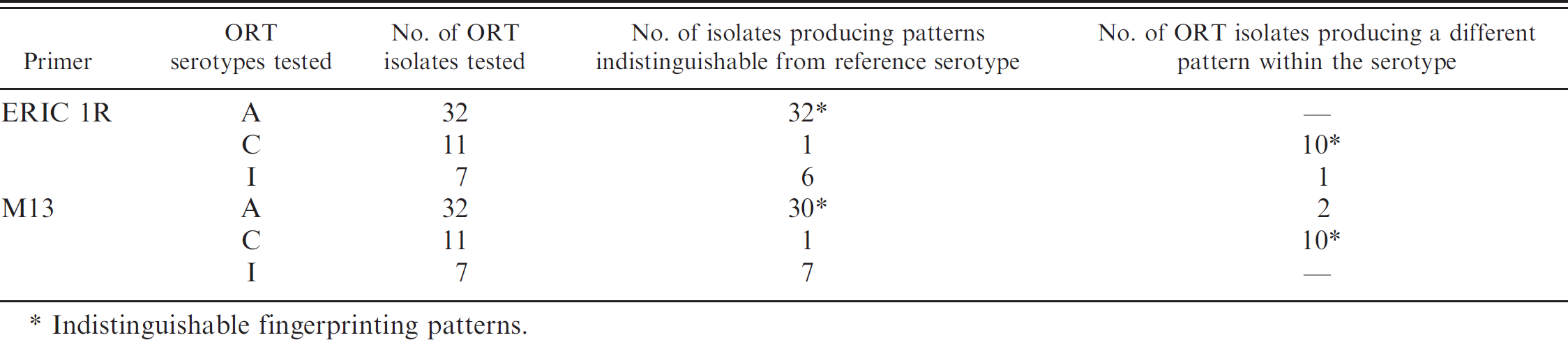

Indistinguishable fingerprinting patterns.
Acknowledgement. The authors would like to thank Dr. Paul Van Empel for providing the ORT reference serotype strains.
Footnotes
a.
Reference isolates were kindly provided by Dr. P. Van Empel, Intervet International, the Netherlands.
b.
bioMerieuX, Marcy l'Etoile, France.
c.
Applied Biosystems, Foster City, CA.
d.
Bio-Rad, Hercules, CA.
e.
Biomedical Genomics Center, University of Minnesota, St. Paul, MN.
f.
PE Thermal Cycler 480, Norwalk, CT.
g.
UVP Biochemi, Upland, CA.
