Abstract
PCR-based approach was used to examine the rate of Chlamydia positivity in raptors from wild bird rehabilitation centers in Oregon. Three of 82 birds were identified as positive for Chlamydia with this PCR. Sequence analysis of 16S ribosomal DNA from 2 of these birds confirmed the presence of DNA from phylum Chlamydiae. One bird was positive for Chlamydia psittaci in both choanal and cloacal swabs. The second bird, a louse-infested red-tailed hawk, had evidence of choanal colonization by “Candidatus Rhabdochlamydia” spp. Our study describes evidence of this Chlamydia-like organism in the United States. This survey also suggests that the carriage rate of C. psittaci is low in raptors in Oregon wild bird rehabilitation centers, and that care must be taken in the design of PCR primers for phylum Chlamydiae such that colonization by insect endosymbionts is not mistaken for an infection by known chlamydial pathogens.
Keywords
Phylum Chlamydiae is a group of obligate intracellular pathogens that infect many animals and humans. Different Chlamydiae also infect invertebrates, including mollusks, insects, and protozoans. 1 Chlamydia psittaci is a serious zoonotic pathogen that colonizes a variety of avian species. 6 Birds of prey are a potential host for C. psittaci, 6 and a previous study conducted on captive raptors in California indicated a high percentage of C. psittaci–infected birds. 4 This study, as well as the zoonotic potential of wildlife rehabilitation birds, motivated our inquiry into the prevalence of C. psittaci in Oregon raptors.
Samples from 82 raptors, collected in 9 different Oregon wildlife centers, were used in the prevalence study. A total of 12 different species were sampled. The Oregon State University Animal Care and Use Committee approved all animal-handling procedures. On admission, polyester swabs a were used to sample choanal and cloacal tissues by trained staff at the rehabilitation center. The swabs were then kept at −18°C prior to laboratory analysis. Total DNA was extracted from the dry swabs using a commercial kit b with some modifications that led to the most efficient harvesting of chlamydial DNA. The most significant modification involved the addition of 5 mM dithiothreitol (final concentration) to the lysis buffer, and following the steps for purification of DNA from gram-positive bacteria. Body lice were also collected from 14 raptors at one rehabilitation center in Oregon, and DNA was purified using the same commercial kit and protocol. The extracted DNA was quantified by spectrophotometry c and stored at −18°C prior to use.
A PCR assay targeting the 5′-end of the Chlamydia spp. 16S ribosomal DNA (rDNA) sequence was used to evaluate the presence of Chlamydiaceae in the birds and lice. General Chlamydiaceae primers 10 were initially used, and a set of similar primers 3 were included to confirm and expand the results (Table 1). BLAST analysis confirmed that amplification of samples with these oligonucleotide pairs would yield an amplicon from the 16S rDNA of all known C. trachomatis, C. psittaci, C. avium, C. pecorum, and C. muridarum. The PCR reaction mixtures consisted of 10 µL of commercial master mix, d 6 µL of nuclease-free water, 2 µL of template, and 1 µL each of the forward and reverse primers (final primer concentration 5 pM). The PCR reactions for amplifying louse 18S rDNA were conducted using identical methods, with the 18S rDNA–specific primers shown in Table 1. The extracted DNA was used as a positive control for the PCR targeting louse rDNA in the bird choanal samples.
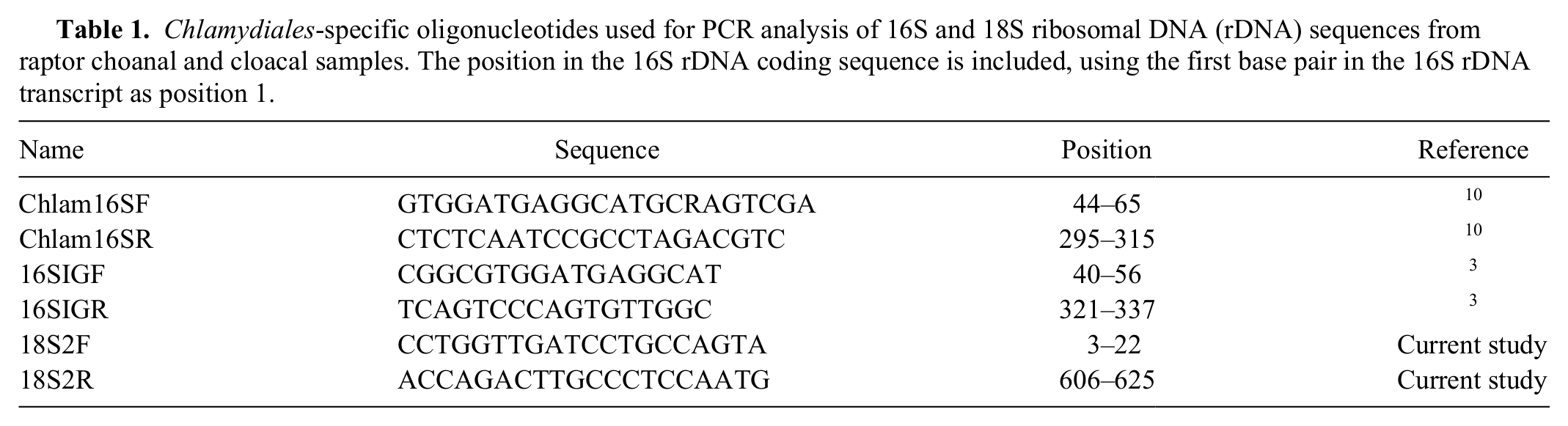
Chlamydiales-specific oligonucleotides used for PCR analysis of 16S and 18S ribosomal DNA (rDNA) sequences from raptor choanal and cloacal samples. The position in the 16S rDNA coding sequence is included, using the first base pair in the 16S rDNA transcript as position 1.

After amplification, each reaction mixture was electrophoresed on a 1% agarose gel. For each assay, sterile and nuclease-free water was used as a negative control, and Chlamydia caviae DNA was used as a positive control. The DNA sequence was determined for PCR-amplified products in positive samples using Sanger sequencing at the Oregon State University Center for Genome Research and Biocomputing (Corvallis, Oregon).
The DNA collected from 3 different birds were PCR positive for chlamydial 16S DNA in the initial screening of choanal samples, prompting analysis of the cloacal samples as well. Cloacal samples from the other birds were not analyzed. The first of these PCR-positive birds, an osprey nestling (Pandion haliaetus) that was presented with multiple fractures from a fall, was positive in both cloacal and choanal swabs. Sequence analysis showed that this bird was infected at both sites with C. psittaci. A second PCR-positive bird was an emaciated great horned owl (Bubo virginianus). Sequence analysis of PCR products from this bird was unsuccessful, and thus the diagnosis was incomplete. The third positive bird, a dehydrated and emaciated young red-tailed hawk (Buteo jamaicensis) with a heavy louse infestation, had a PCR-positive choanal sample but a PCR-negative cloacal sample. Nucleotide sequence analysis of PCR products from this bird, using 2 different primer pairs, demonstrated that the positive PCR did not result from C. psittaci infection. Clustal-based analysis of the fragment derived from the choanal sample showed clearly that this sequence was more closely related to Rhabdochlamydia spp. than to C. psittaci, or any other member of the genus Chlamydia (Fig. 1A, and data not shown). Phylogenetic positioning of this sequence was then analyzed using a neighbor-joining approach, conducted using a commercial package. e These results indicated strong support for the relationship between the PCR product from our clinical sample and members of the genus Rhabdochlamydia (Fig. 1B). The results also confirm that sequences from Chlamydia-like organisms can be detected using generalized chlamydia primers, and that subsequent sequence analysis is required to confirm the actual nature of a 16S rDNA–based PCR-positive sample.
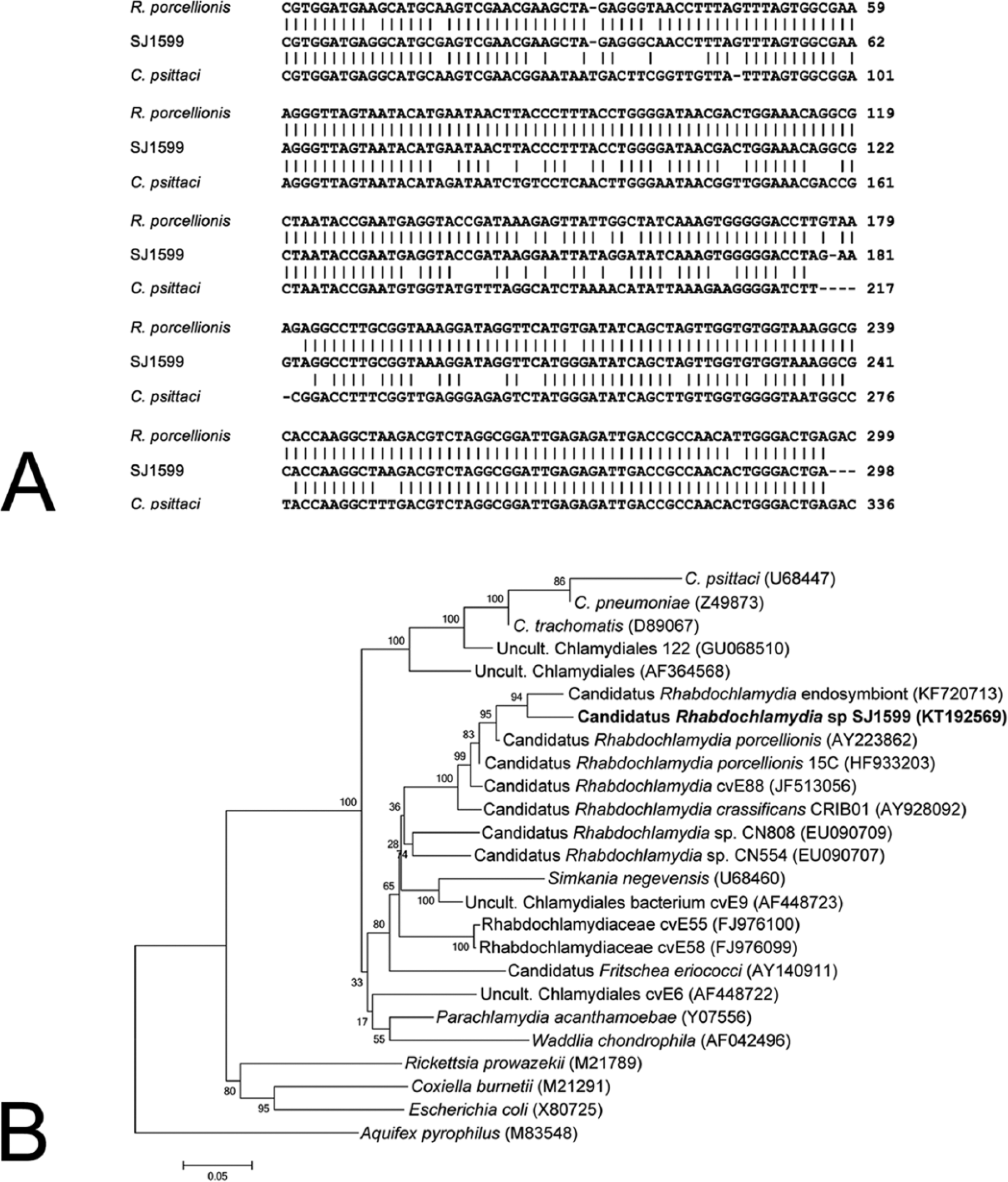
Taxonomic placement of the 16S ribosomal DNA (rDNA) sequence obtained from a captive red-tailed hawk.
The bird that was colonized by Rhabdochlamydia spp. was heavily infested with lice. This observation led to the hypothesis that the bird was positive because it had been preening lice off its body, and the choana therefore became contaminated with an arthropod endosymbiont. This was not supported by our analyses, as louse-specific 18S rDNA was not amplified from the choanal swab sample, although raptor 18S DNA was amplified from the swab.
None of the 14 louse DNA preparations were positive for Rhabdochlamydia. These results suggest that neither birds nor lice within the geographic region serviced by these raptor centers (northwest Oregon) are routinely positive for Rhabdochlamydia spp. Also, because we were only able to demonstrate C. psittaci in 1 of 82 sampled birds, it appears that C. psittaci prevalence is not high in these bird populations. These data are in contrast to the enzyme-linked immunosorbent assay (ELISA)-based testing of California raptors published in 1990, in which more than 40% of animals were considered positive. 4 As our techniques were quite different to those used in the California study (PCR vs. ELISA), and because of the lengthy time interval between the 2 studies, it is challenging to assess the meaning of the relatively high prevalence indicated in the previous study to the very low prevalence indicated in our work. A culture-based survey of wild birds in UK rehabilitation centers also reported a low percentage of positive raptors (1/9), although overall prevalence in all avian species sampled was 26%. 7
Rhabdochlamydia is a genus currently represented by endosymbionts of arthropods in Europe,2,8,11 and by rDNA sequences in environmental samples obtained from a variety of sources. 9 The presence of “Candidatus Rhabdochlamydia” spp. has also been documented in mammals, including cattle in Great Britain, 12 and in human respiratory specimens collected in Germany. 5 Our study describes evidence of Rhabdochlamydia spp. in the United States. It is likely, however, that this is a result of the absence of directed testing and not because the organism is necessarily rare in this region of the world.
Footnotes
Acknowledgements
We thank the contributing raptor centers for their great help and cooperation.
Authors’ contributions
SJ Jouffroy contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. AH Schlueter contributed to acquisition, analysis, and interpretation of data and critically revised the manuscript. RJ Bildfell and DD Rockey contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; critically revised the manuscript; and gave final approval. All authors agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
PurFlock Ultra, Puritan Medical Products, Guilford, ME.
b.
Qiagen GmBH, Hilden, Germany.
c.
NanoDrop Technology, Thermo Fisher Scientific Inc., Waltham, MA.
d.
Phusion master mix, New England Biolabs Inc., Ipswich, MA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Oregon State University College of Veterinary Medicine Summer Student Research Program (VBS330 VBSR). We also thank the National Veterinary College of Toulouse Alumni Fund for their financial support.
