Abstract
Respiratory diseases in pigs are mostly polymicrobial, and the involved pathogens can vary from farm to farm. The impact of Pneumocystis carinii (P. c.) f. sp. suis on respiratory disorders has not been comprehensively appraised because tests were limited to lung tissue samples and, for this reason, it was not possible to detect the fungus in living animals. In the present study, bronchoalveolar lavage fluid (BALF) from 12 pigs and oral fluid samples from 9 pigs were tested for the presence of Pneumocystis by quantitative real-time PCR. The results from these 2 clinical specimens were compared with Pneumocystis quantities in lung tissue samples. Pneumocystis quantities in BALF correlated significantly to those in lung tissue. BALF has proved to be an adequate specimen for detection of various respiratory pathogens in pigs, and the collection procedure directly on farms is also well established. In contrast to the BALF results, all oral fluid samples were negative. Thus, specimens from the lower respiratory tract should generally be preferred for the detection of Pneumocystis. Additionally, under farm conditions, oral fluid is mainly collected in the form of collective samples per pen. In the present study, oral swab sampling of individual pigs was intended but failed in 3 of 12 pigs because they did not salivate sufficiently. As a conclusion, only BALF can be recommended as a useful tool for Pneumocystis herd monitoring in vivo.
Introduction
Pneumocystis carinii (P. c.) f. sp. suis is an opportunistic fungus that is mainly associated with milder lung lesions, such as interstitial pneumonia. 1 The fungus is found in pigs with a relatively high prevalence (32%), and coinfections with other respiratory pathogens are frequently observed. 1 Respiratory diseases in pigs have been described to be polymicrobial and multifactorial, and the involved pathogens can vary markedly from farm to farm. 8 To date, the impact of P. c. f. sp. suis on complex respiratory disorders in pigs has not been investigated intensively. 10 One of the reasons for this lack of data is that most analyses have been of lung tissue samples collected at the slaughterhouse. 3 Also, not all age classes susceptible to respiratory pathogens have been covered. The ultrasound-guided collection of biopsy samples from living animals can only be carried out under the risk of anesthesia complications, development of hemorrhage caused by the procedure, and a large technical and time-consuming effort per pig. 9 In human medicine, the sample of choice for Pneumocystis detection is bronchoalveolar lavage fluid (BALF).5,14 BALF has already proved to be an adequate specimen for the identification of various respiratory pathogens in pigs, and the collection procedure directly on farms is well established. 15 The second specimen that has gained increasing importance in porcine veterinary testing is oral fluid. Collection of sputum using cotton ropes is a commonly used procedure for detection of Porcine reproductive and respiratory syndrome virus (PRRSV) and other infectious agents. 16 The establishment of a quantitative real-time (q)PCR to detect porcine Pneumocystis in BALF and oral fluid provides a promising opportunity to obtain information about the presence of the fungus in pig production units and was the central aim of the present study.
Materials and methods
A total of 12 lung tissue, 12 BALF, and 9 oral fluid samples of pigs from 2 farms were included in the present study. The pigs were sent to the University Clinic for Swine (Vienna, Austria) and forwarded to the Institute of Pathology and Forensic Veterinary Medicine of the University of Veterinary Medicine (Vienna, Austria) for routine testing. Pigs were anesthetized with a combination of 2 mg of azaperone a per kg body weight and 10 mg of ketamine b per kg body weight, both administered intramuscularly, and euthanized with 0.3 mL of embutramid c per kg body weight administered intracardially. This method is routinely applied by the University Clinic for Swine and is consistent with the recommendations of the AVMA Guidelines for the Euthanasia of Animals. The owners’ consents for using samples were obtained on admission of the pigs, and no further ethics permission was required. Compliance with the Austrian federal law for animal experiments (Tierversuchsgesetz) and the University for Veterinary Medicine Vienna ethics of good scientific practice was ensured.
Selection criteria were age of the pigs and the presence of mild respiratory, gastrointestinal, or multisystemic signs (Table 1). A portion of lung tissue samples was fixed in 10% formalin, embedded in paraffin, and subjected to histologic investigation (hematoxylin and eosin staining) and in situ hybridization (ISH). For every pig, sterile collection equipment was used, and the samples were handled cautiously and processed under sterile conditions preventing sample-to-sample cross-contamination. BALF samples were collected as described previously 7 and centrifuged at 2,000 × g for 10 min. The supernatant was discarded, and the pellet was resuspended in phosphate buffered saline (PBS). Oral fluid samples could only be obtained from 9 of the 12 pigs and were collected by centrifugation of oral swabs that had been chewed by the pigs for several minutes. BALF pellets, oral fluid, and native lung tissue samples were stored at −20°C until analysis.
Semiquantitative Pneumocystis results of in situ hybridization (ISH) and quantitative Pneumocystis results of quantitative real-time (q)PCR for native lung tissue, bronchoalveolar lavage fluid (BALF), and oral fluid.*
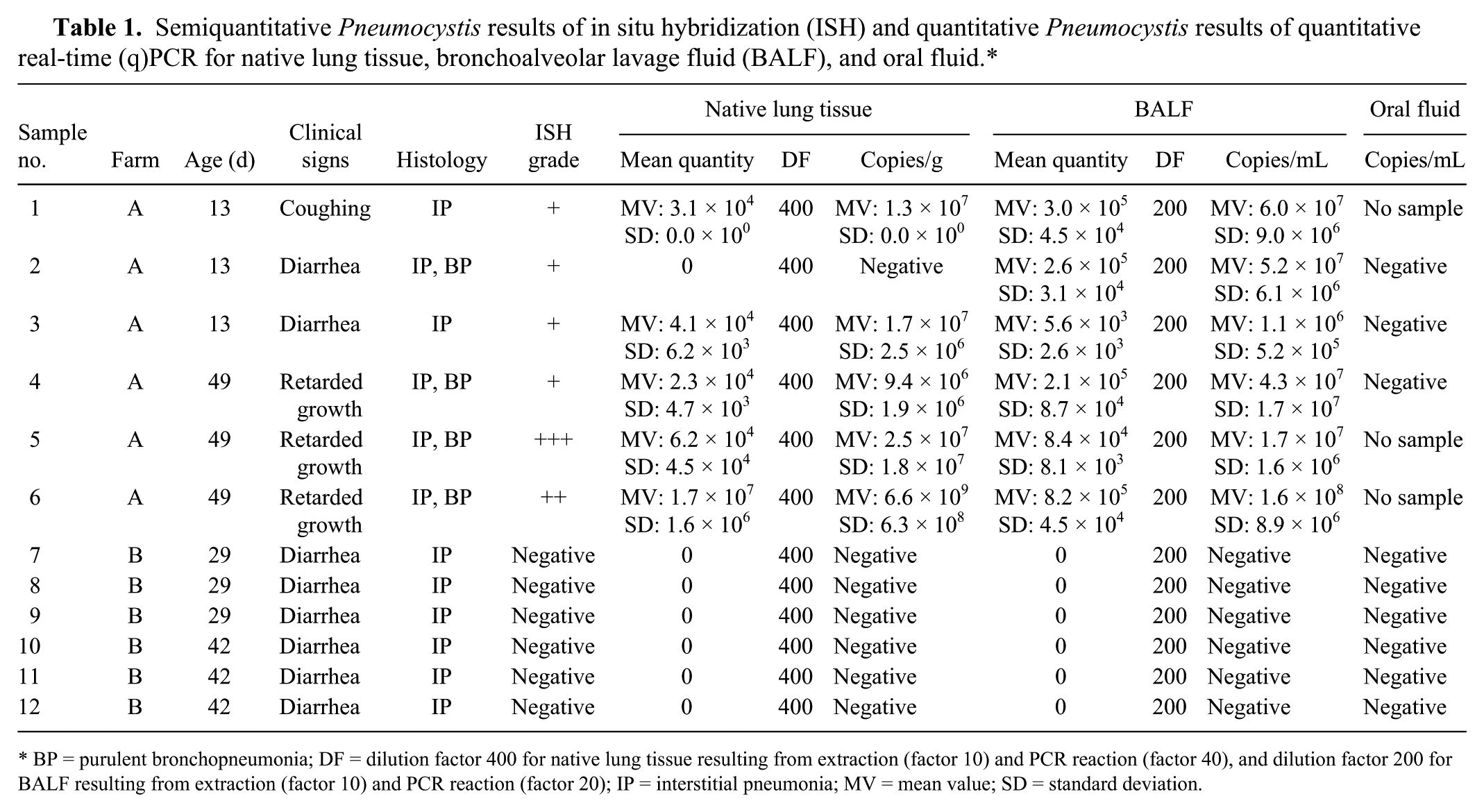
BP = purulent bronchopneumonia; DF = dilution factor 400 for native lung tissue resulting from extraction (factor 10) and PCR reaction (factor 40), and dilution factor 200 for BALF resulting from extraction (factor 10) and PCR reaction (factor 20); IP = interstitial pneumonia; MV = mean value; SD = standard deviation.
In situ hybridization
In situ hybridization was applied as described previously. 1 The oligonucleotide probe sequence was designed within the 18S ribosomal (r)RNA gene (5′-GGAACCCGAAGACTTTGATTTCTCATAAGATGCCGAGCGA-3′). The semiquantitative analysis was assessed using the score +++ for multiple, ++ for moderate, and + for few organisms (Fig. 1).
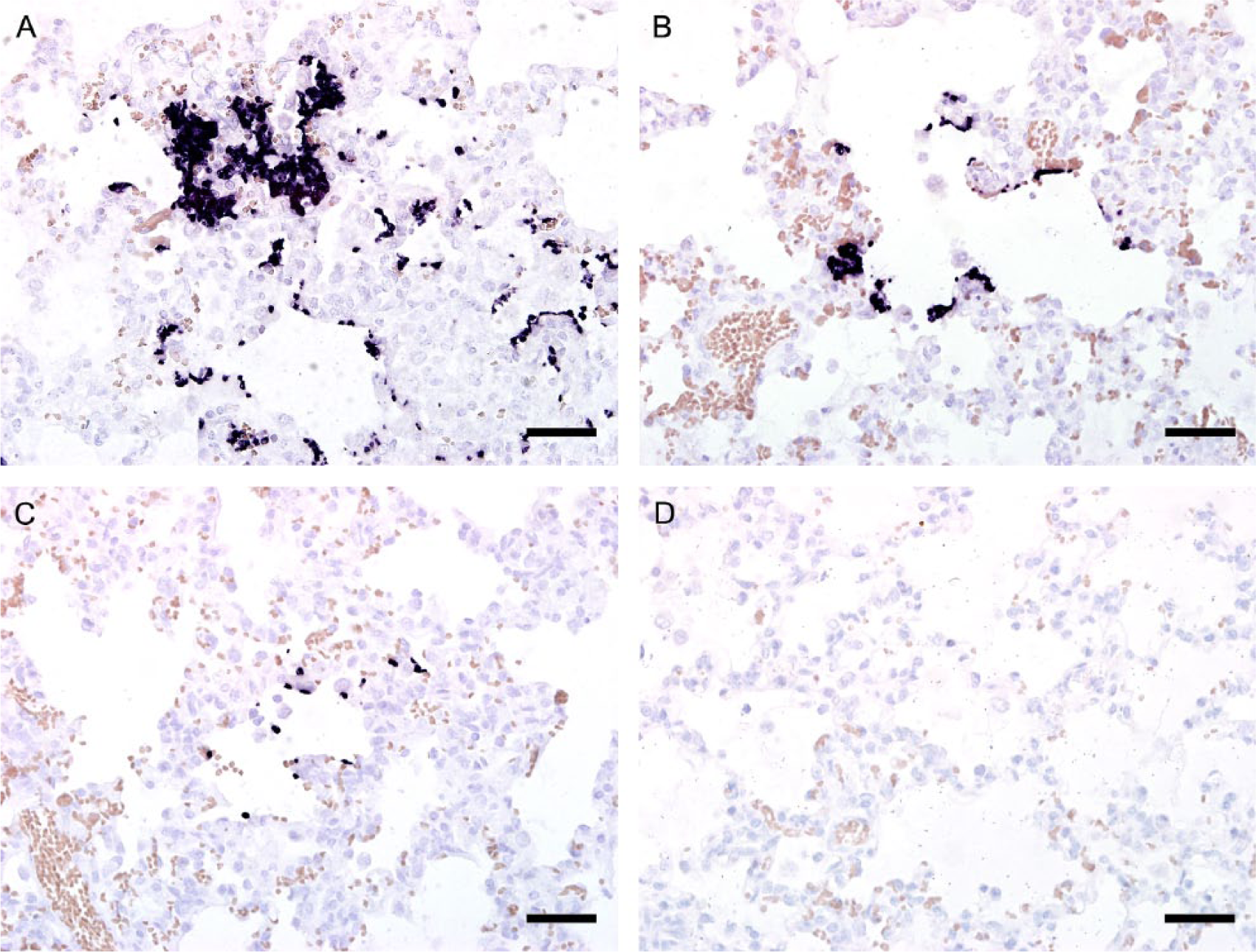
Pig lung, in situ hybridization. The semiquantitative analysis was assessed using the score +++ for multiple (
Quantitative real-time PCR
Primers and probe with the potential to detect all representatives of P. c. f. sp. suis were designed d according to published guidelines 2 after extensive homology studies on all available GenBank sequences of the mtLSU rRNA gene. The sequences were submitted to BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to search against GenBank sequences and to exclude unintentional cross-reactivity with other organisms. Additionally, specificity was tested in vitro. Samples positive for Aspergillus spp., Candida spp., and Penicillium spp. remained negative.
Prior to qPCR, DNA was extracted from ~0.5 mg of native lung tissue e and from 100 µL of the resuspended BALF pellet or oral fluid f according to the manufacturer’s instructions. DNA from native lung tissue was eluted in a volume of 100 µL of buffer, and DNA from BALF and oral fluid in 50 µL of buffer. The qPCR reaction master mixture consisted of 10 µL of master mix, g 0.5 µM of each primer (forward: 5′-TAGCTGGTTTTCTGCGAAAT-3′; reverse: 5′-TTCTGGGCTGTTTCCCTTTA-3′), 3.125 µM of the probe (5′-ATCTCAGAATGGCTAATAAGTTAGG-3′), 2.5 µL template DNA, and distilled water to a total volume of 20 µL. The PCR reactions were started with a first heat denaturation step at 95°C for 20 s, followed by 40 cycles at 95°C for 3 s and 57°C for 2 min.
For the production of standards, an amplification product of a positive control, which had been confirmed by sequencing, h was cloned.i,j Ten separate colonies were selected from the lysogeny broth agar k and incubated overnight at 37°C in lysogeny broth. k Plasmid DNA was extracted, l and the concentration was determined with a spectrophotometer. m A range of dilutions of the targeted gene was produced for establishment of the standard curve.
Statistical analysis
The correlations between Pneumocystis quantities in native lung tissue and BALF, as well as the semiquantitative ISH score results, were calculated using Pearson coefficient of correlation r. n
Results
Histologically, all pigs had interstitial pneumonia (Table 1). Additionally, 4 pigs from farm A had purulent bronchopneumonia. In ISH, all lung tissue samples of farm A were Pneumocystis positive. In qPCR, 5 of 6 lung tissue and all BALF samples from pigs from farm A were positive. Samples from farm B as well as oral fluid samples of both farms were negative. The semiquantitative ISH results as well as the quantitative PCR results of the positive samples are presented in Table 1. The ISH signals were restricted to the lung alveoli and were either directly attached to the type I pneumocytes lining the alveolar spaces or, in high-grade cases, filling alveoli (Fig. 1). Colonization with Pneumocystis was shown to be mild in 4 samples, moderate in 1, and severe in 1.
The limit of detection of the qPCR was 4 × 101 copies/µL. For the establishment of the standard curve, plasmid DNA in the concentrations 1.0 × 109 to 1 copy/2.5µL template DNA were used in 1:10 serial dilution steps. The standard curve showed an average slope of −3.44, a regression r2 of 0.988, and an efficiency of 95.7% (Fig. 2). For the investigation of the samples, plasmid DNA in the concentrations 1.0 × 108, 1.0 × 106, and 1.0 × 104 was used. All standards were run in triplicate.
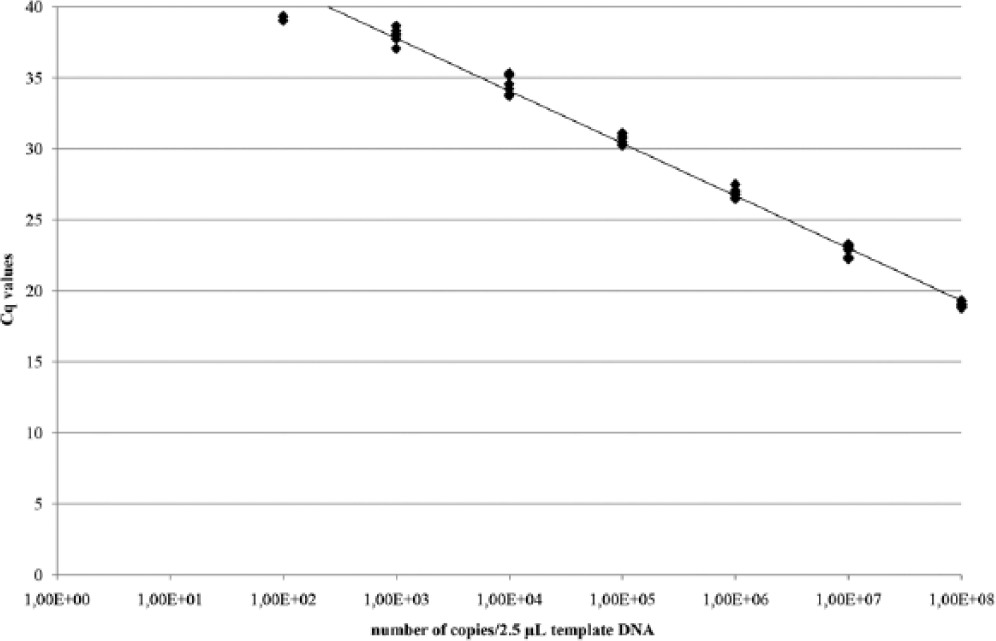
Standard curve of the plasmid DNA in the concentrations 1.0 × 109 to 1 copy/2.5 µL template DNA in 1:10 serial dilution steps. Cq = quantification cycle.
The qPCR results of native lung tissue ranged from 9.4 × 106 to 6.6 × 109 copies/g tissue. The BALF results ranged from 1.1 × 106 to 1.6 × 108 copies/mL lavage fluid. One lung sample was negative but the corresponding BALF sample was positive. Nevertheless, qPCR results of native lung tissue and BALF correlated significantly (Fig. 3A; r = 0.883, p < 0.001), whereas ISH and qPCR results did not correlate (ISH/native lung tissue (Fig. 3B): r = 0.411, p = 0.184; ISH/BALF (Fig. 3C): r = 0.547, p = 0.066). The ISH(+++)-positive case had 107 copies/g in native lung tissue and 107copies/mL in BALF in the qPCR, whereas the (++)-positive case had 108 copies/mL in BALF and 109 copies/g in native lung tissue. In the ISH(+)-positive cases, there were Pneumocystis quantities between 106 copies/g in native lung tissue and 107 copies/mL in BALF.
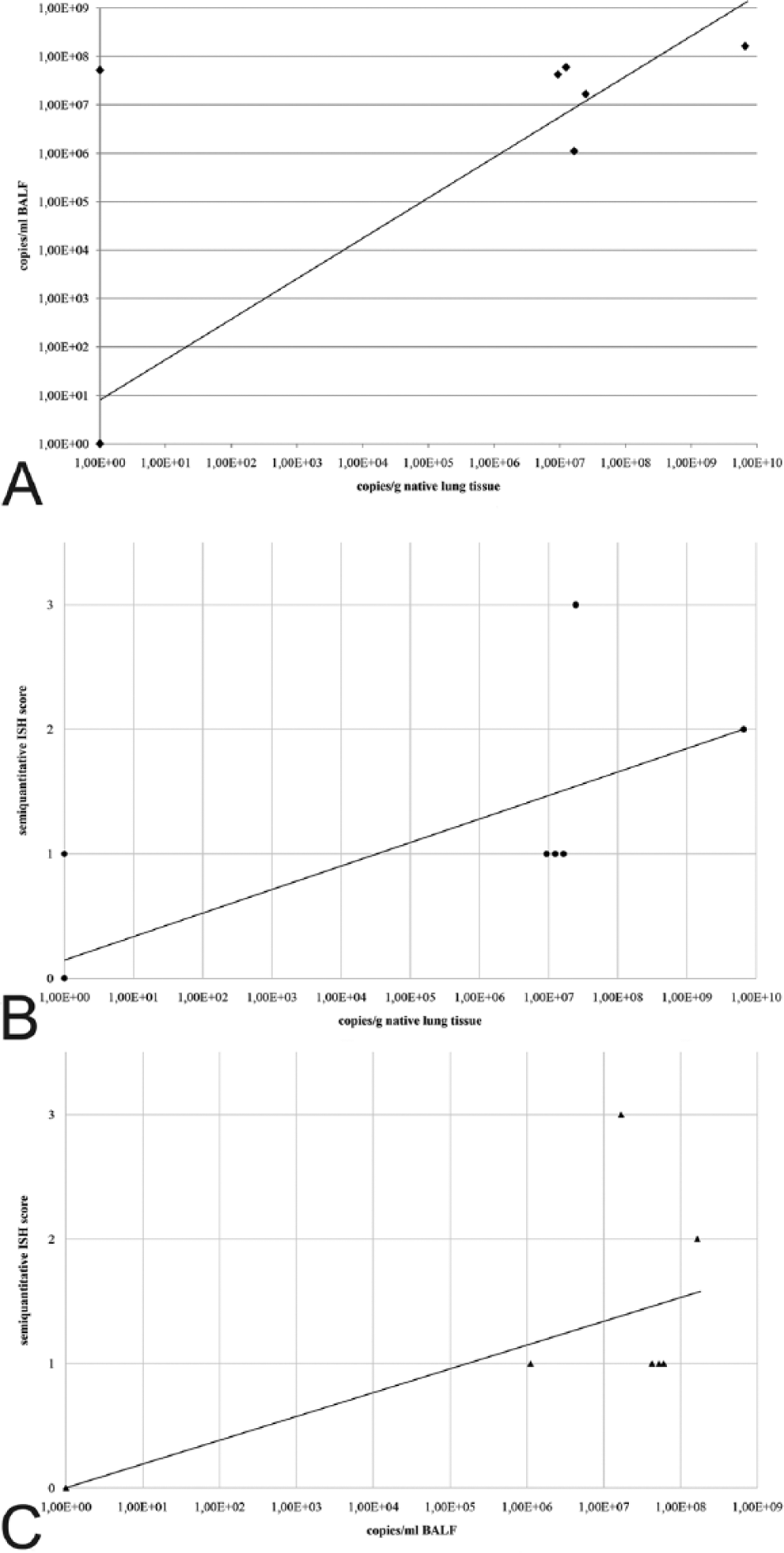
Scatterplot diagrams of the correlations.
Discussion
Pneumocystis pneumonia is a major cause of mortality in immunocompromised humans. 4 To date, the impact of Pneumocystis on the respiratory disease complex of pigs has not been investigated in depth. Interactions and synergies between various viral and bacterial pathogens of the respiratory tract have been described, but the role of fungi in general has not been elucidated adequately. 13 Analytical methods that have been established for specimens collected from dead animals are often not suitable for herd screening. Biopsy techniques have been refined, but most animal species must be anesthetized for this procedure. Species such as pigs are easily excited by simple immobilization and can develop stress-induced circulatory collapse. Furthermore, a targeted biopsy is nearly impossible if the animal is not adequately immobilized. Dependent on the species-related differences of the site(s) to be sampled, the collection of biopsy samples is often only possible under ultrasound guidance, which can have a direct impact on the sampling time and duration of anesthesia. As a consequence, tissue-based methods, such as ISH, are unsuitable for herd screening purposes.
The detection of respiratory pathogens does not need to be carried out on lung tissue samples. The investigation of BALF 15 or oral fluid 16 mainly by PCR has been well established for pig herd monitoring. In the present study, lung tissue samples were analyzed by ISH, 1 which is a classical molecular pathology method and a recommended tool for rapid tissue localization of infectious agents. ISH allows semiquantitative evaluation of slides and therefore provides a perfect basis for establishment of a quantitative method, such as qPCR, because the amount of ISH signal should correspond directly to the quantity in the qPCR. In the present study, ISH and qPCR results from native lung tissue were not concordantly positive or negative; in addition, the semiquantitative/quantitative evaluation did not correlate perfectly.
Pneumocystis is a fungus that attaches to the alveolar epithelium and proliferates from this starting point. As a consequence, the distribution of the fungus is not uniform. In contrast, not only areas with dense colonization but also ones with only a few ISH signals, can be seen next to each other (Fig. 4). This nonhomogeneous distribution pattern could be the reason for the variation of the results. Whether Pneumocystis is restricted to certain lung lobes or whether special locales are preferred by the fungus needs to be investigated. If an exact determination of the quantity is not of prior diagnostic value, a slightly reduced accuracy of results is not significant. The trophic stages of Pneumocystis adhere tightly to type I pneumocytes using filopodial structures.11,12 Attachment is supported by interactions between the fungi and adhesive glycoproteins (e. g., fibrinogen) present in increased concentration in the lower respiratory tract during Pneumocystis pneumonia. 18 As the strength of this filopodial attachment was unclear, we were uncertain whether a sufficient amount of fungus could be obtained by lung lavage. Only 5 native lung samples were positive, whereas 6 BALF samples were positive. The irregular distribution of the fungus in tissue might be compensated for by flushing the lung whereby different lung parts can be reached equally. The collection of BALF is a well-established procedure and is routinely used for investigation of respiratory pathogens in pigs. 15 During lung lavage, the pigs also have to be anesthetized, but a skilled veterinarian is able to carry out the sampling in a few minutes and, as a consequence, stress factors and cardiovascular risk are negligible.
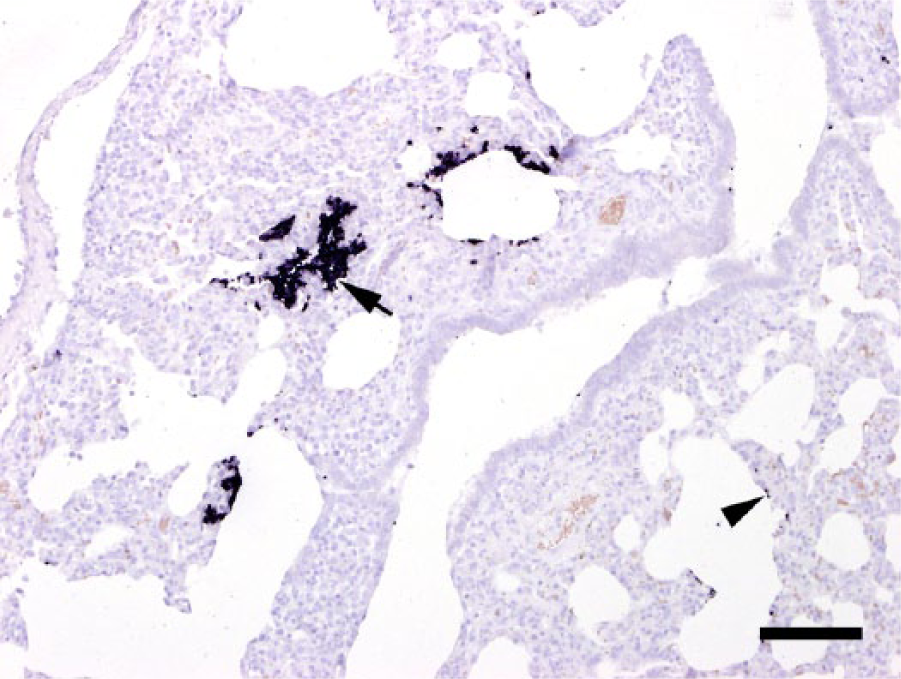
Pig lung. In situ hybridization labels large colonies of numerous Pneumocystis developmental stages in areas with dense colonization within the alveolar lumen (arrow), but also singular stages along the alveolar wall (arrowhead). Bar = 100 µm.
The second specimen of interest was oral fluid, which is mainly used for PRRSV testing. Cotton ropes are hung in the pens, and the pigs, motivated by curiosity and play instinct, start chewing them. Oral fluid is collected by mechanical compression of the ropes. 16 The disadvantage of this method is that only collective samples per pen are obtained. In our study, oral cotton swabs were used for sputum collection to obtain individual samples. Only 9 pigs salivated and produced a sufficient amount of sputum. To minimize stress, the duration of anesthesia was kept as short as possible. During a longer anesthesia period, the pigs might have salivated more. Considering these facts, BALF collection is more promising. In the present study, all oral fluid samples were negative. Only 3 oral fluid samples of Pneumocystis-positive pigs were available, and these 3 cases were categorized as mild based on BALF and lung tissue results. Induced sputum has been described as a useful technique for the detection of Pneumocystis jirovecii in HIV patients.6,17 Pneumocystis infections of immunocompromised humans can be severe. 19 The fungus concentration in the lungs is likely extremely high and, as a consequence, Pneumocystis is easily expectorated with the sputum by coughing. The oral fluid samples in the present study came from mildly infected pigs and, in those cases, sputum seems to be inadequate for accurate detection. Oral fluid samples of the pigs with moderate to severe Pneumocystis infections could not be obtained. For this reason, it remains unknown whether these samples would have been positive or whether specimen of the lower respiratory tract should generally be preferred. P. c. f. sp. suis can be detected in BALF successfully. This method renders Pneumocystis monitoring on a farm possible and facilitates the assessment of the fungus’ impact on respiratory diseases in pigs.
Footnotes
Acknowledgements
We thank Klaus Bittermann and Karin Fragner for their excellent technical support. The positive controls for excluding cross reactivity with other fungi were kindly provided by Dr. Joachim Spergser, Institute of Bacteriology, Mycology and Hygiene, University of Veterinary Medicine Vienna, Austria.
Authors’ contributions
C Weissenbacher-Lang and H Weissenböck contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and drafted the manuscript. N Nedorost contributed to conception of the study, and contributed to analysis and interpretation of data. C Knecht and I Hennig-Pauka contributed to conception of the study, and contributed to acquisition and interpretation of data. All authors critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Stresnil, Janssen-Cilag AG, Baar, Switzerland.
b.
Narketan, Vetoquinol GmbH, Ravensburg, Germany.
c.
T61, MSD Animal Health Austria, Vienna, Austria.
d.
Sci Ed Central for Windows 95, Scientific & Educational Software, Cary, NC.
e.
Nexttec clean column kit, Nexttec Biotechnologie GmbH, Hilgertshausen, Germany.
f.
QIAamp DNA micro kit, Qiagen GmbH, Hilden, Germany.
g.
TaqMan fast universal PCR master mix (2×) No AmpErase UNG, Life Technologies Austria Zweigniederlassung, Vienna, Austria.
h.
Microsynth AG, Wolfurt-Bahnhof, Austria.
i.
StrataClone PCR cloning kit, Agilent Technologies, Santa Clara, CA.
j.
StrataClone Solopack competent cells, Agilent Technologies, Santa Clara, CA.
k.
Life Technologies Austria Zweigniederlassung, Vienna, Austria.
l.
PureLink quick plasmid miniprep kit, Life Technologies Austria Zweigniederlassung, Vienna, Austria.
m.
NanoVue plus spectrophotometer, GE Healthcare, Vienna, Austria.
n.
PASW 17, SPSS, Quarry Bay, Hong Kong SAR.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the profile lines, University of Veterinary Medicine Vienna, Austria, and by the Verein der Freunde und Förderer der Schweinemedizin, University of Veterinary Medicine Vienna, Austria.
