Abstract
Neospora caninum is a protozoan parasite that causes an important reproductive disease in cattle. Neospora caninum surface antigen 1 (Nc-SAG1) is an immunodominant candidate for the development of a diagnostic reagent for neosporosis. The current study describes the development and evaluation of an antigen print immunoassay (APIA) with recombinant Nc-SAG1 for the detection of specific antibodies to N. caninum in cattle. The concordance between APIA and a commercial enzyme-linked immunosorbent assay (ELISA) was evaluated with 232 serum samples from experimentally and naturally infected cattle. Sixty-one (26.7%) samples were positive for antibodies to N. caninum by ELISA and 58 (25.4%) by APIA. The new assay had a sensitivity of 85% and a specificity of 96%. These results, along with the potential of APIA to evolve into a multiple antigen detection format, suggest that this method would be a reliable diagnostic test for detection of antibodies to N. caninum in cattle.
Neospora caninum is a protozoan parasite first recognized in dogs in Norway. 4 The parasite was isolated into tissue culture and identified as a new genus in 1988. 12 Disease was first associated with congenital encephalomyelitis in puppies. Subsequently, fetal and neonatal infections were described in cattle, sheep, goats, deer, and horses.6,13,21 In the last decade (2000–2010) neosporosis has emerged as an important reproductive disease in cattle. Most of the studies of N. caninum infections have examined their impact mainly on dairy cattle. 10 However, some authors have established that Neospora infections are important in beef herds as well.27,28
A report from 1995 indicated that neosporosis was present in Argentinean beef and dairy herds. The infection status was corroborated by serological evidence using indirect fluorescent antibody test (IFAT) and was suspected as a cause of abortion in these herds. 29 In 1998, lesions of nonsuppurative encephalitis, hepatitis, and myocarditis were found in histological examination of aborted fetuses. 7 One year later, a diagnosis of N. caninum infection was confirmed in these aborted fetuses based on histopathology and serology results. 30
Neospora caninum infection can be diagnosed by the use of histological, immunohistochemical, and polymerase chain reaction (PCR) techniques on the brain or other tissues of aborted fetuses. Serological tests are also a diagnostic tool currently available. Such tests include immunoblotting, 1 IFAT, 15 agglutination test, 26 and several enzyme-linked immunosorbent assays (ELISAs).3,5,25
Serological evaluation of neosporosis in cattle herds requires methods that enable rapid processing of large numbers of samples. The use of recombinant antigens in serological tests has additional benefits: they are easily produced in large quantities and can be readily standardized. In addition, the use of highly specific antigens may minimize the risk of cross-reactivity with other parasite species. In this sense, the N. caninum surface antigen 1 (Nc-SAG1) was shown to be an effective candidate to diagnose N. caninum infection in cattle when it was used in ELISA 8 and rapid immunochromatographic test. 20
The current study described the development and evaluation of an antigen print immunoassay (APIA) with recombinant Nc-SAG1 for detection of specific antibodies to N. caninum in cattle. The APIA or line immunoassay is an antibody detection method that employs the direct application of proteins sprayed in lines onto nitrocellulose membranes followed by the performance of classical antibody detection methods. A similar method called MAPIA (multiple antigen printing immunoassay) was previously developed using a multi-antigen cocktail of Mycobacterium tuberculosis with greater serological reactivity than that observed with ELISA. 22 Recently, a MAPIA test was also developed for cysticercosis and taeniasis. 17 In the current work, the performance of the APIA was evaluated for the serological detection of N. caninum and compared with the standard diagnostic methods for bovine neosporosis.
Neospora caninum isolates used in the current study were strains Nc-1,a,14 Nc-6,a,2 and Nc-7. b Tachyzoites were grown in monolayers of Vero (African green monkey kidney epithelial) cells and purified as previously described. 23 For the cloning of the truncated Nc-SAG1 gene, the genomic DNA of purified N. caninum tachyzoites Nc-1 strain (1 × 108) was extracted using a commercial kit. c The DNA pellets were dissolved in a TE buffer (10 mM Tris–HCl [pH 8.0], 1 mM ethylenediamine tetra-acetic acid) and used as a template DNA for PCR. The truncated Nc-SAG1 gene does not include the hydrophobic signal peptide and the C-terminus as previously described. 8 The Nc-SAG1 gene was amplified by PCR using the oligonucleotide primers Nc-SAG1F (5’-GGCATCAGAAAAATCACC-3’) and Nc- SAG1R (5’- GACCAACATTTTCAGCCG-3’) at an annealing temperature of 54°C. The PCR product was ligated into a cloning vector, d and this fragment was further subcloned into the EcoRI site of the bacterial expression vector pRSET. e The correct orientation and fusion of the open reading frame of Nc-SAG1 with the vector’s histidine tag was confirmed by sequencing at the sequencing facility of the Biotechnology Institute–National Institute of Agricultural Technology with a genetic analyzer. f The Nc-SAG1 gene was expressed in Escherichia coli (BL21 pLys strain). Escherichia coli transformed with pRSET–Nc-SAG1t was cultured at 37°C until the OD600 nm (optical density at 600 nm) level reached 0.5. Synthesis of Nc-SAG1 was induced with 1 mM isopropyl beta-D-thiogalactoside at 37°C for 4 hr. The resulting E. coli were harvested by centrifugation (10,000 × g for 15 min at 4°C) and treated by sonication in a native lysis buffer (50 mM K2HPO4, 400 mM NaCl, 100 mM KCl, 10% glycerol, 0.5% Triton X-100, 10 mM imidazole [pH 7.8]) plus 1 mg/ml lysozyme and 2 mM phenylmethylsulfonyl fluoride. The suspension was centrifuged again (10,000 × g for 15 min at 4°C), and the pellet containing the inclusion bodies was further resuspended in 2 ml of 8 M urea and incubated at 37ºC for 1 hr. The suspension was sonicated and centrifuged again as before, and the supernatant was harvested. The Nc-SAG1 recombinant protein present in this supernatant was purified with a nickel resin g with denaturing conditions according to the manufacturer’s instructions.
Sera used in the current study were as follows: 1) A “gold standard” panel was composed of 16 samples from cattle experimentally infected with either Nc-1 USA strain (n = 2 intravenously inoculated with 2 × 108 parasites and n = 4 inoculated with 2.4 × 108 by conjunctival route), Nc-6 Argentina strain (n = 4 intravenously inoculated with 5 × 107 parasites), or Nc-7 Spain strain (n = 6 intravenously inoculated with 2 × 108 parasites). Blood was extracted when seroconversion occurred (as evaluated by IFAT), and sera were stored at −20°C. Pre-immune sera from the cattle were obtained before inoculation and were negative by IFAT at a titer ≤1:25 so they were further used as negative controls. 2) Field bovine sera (n = 212) were collected from cattle pastured on different farms in central Argentina (Buenos Aires, La Pampa, and San Luis provinces) in which isolated cases of abortions were reported. All serum samples diluted 100-fold were analyzed in parallel with a commercial ELISA. h In this test, the presence or absence of antibodies to N. caninum was determined by calculating the sample-to-positive (S/P) ratio according to the following equation: S/P = sample (OD) – negative control(OD)/positive control(OD) – negative control(OD). Serum samples with S/P ratios <0.50 were classified as negative, and samples with S/P ratios ≥0.50 were classified as positive. According to the manufacturer, the specificity and sensitivity of this ELISA are 99.2% and 100%, respectively. Indirect FAT was also performed with sera from experimentally infected animals using a protocol described elsewhere. 11 The specificity and sensibility of IFAT are between 82.4–97% and 85.7–90%, respectively, as reported previously. 1 Slides were examined with an epifluorescence microscope. i A serological titer >1:25 was considered an evidence of N. caninum infection. 11
For the APIA, NcS-AG1 was immobilized on nitrocellulose membranes j as a narrow band by using a semiautomatic airbrush device k as previously described. 22 An image of a real APIA membrane after color development is shown in Figure 1A. The protein was printed at a concentration of 173 µg/ml in Tris buffer solution (TBS; 150 mM NaCl, 10 mM Tris [pH 7.5]). This concentration of Nc-SAG1 was used according to previously published data. 22 Higher concentrations of antigen did not show significant differences in the signal intensity. Twenty µl of antigen solution was sufficient to print a band that was 15 cm long. After antigen printing, the membrane was stored at room temperature until used. For the APIA, nitrocellulose membranes were blocked for 1 hr with 5% nonfat skim milk in TBS (blocking buffer). Afterwards, the membrane was placed on a support cushion into a miniblotter device l such that the slots were perpendicular to the line pattern of the applied antigen. The device was adjusted tightly to avoid leakage of sera among slots. Each membrane was incubated for 1 hr with samples diluted 1:50 in blocking buffer in a final volume of 180 µl. Optimal serum concentration was chosen based on previous serial titrations between 1:50 and 1:200 dilutions. In each experiment, 6 positive sera from the experimentally infected cattle and 4 negative sera were included in the membrane along with 28 test serum samples (Fig. 1A). To avoid cross contamination, each sera was individually aspirated before the washing procedure. After incubation, the membrane was removed from the miniblotter, washed with TBS, incubated for 1 hr with protein G–horseradish peroxidase conjugate m diluted 1:250 in TBS and washed again with TBS for 5 min. Enzyme activity was visualized by incubating the membrane for 5 min with a chromogen consisting of 0.5 mg/ml 4Cl-naphtol in TBS (pH 7.5) with the addition of 1% H2O2. Membranes were extensively rinsed in distilled water to stop the reaction. The entire procedure was performed at room temperature. APIA results were evaluated visually, with the presence of a spot of any intensity being considered a positive result. All sera were evaluated twice. As a measure of reproducibility, the results were read by 2 independent readers who were blinded to the origins of the sera; discrepant results were resolved by a third reader who was also blinded to the status of the sera tested. Densitometry analysis of the APIA membrane was also performed. n Spot intensity was normalized against one of the negative sera spots. Concordance between ELISA and APIA was calculated using the kappa coefficient 9 (κ) according to the following formula: κ = Pr(a) – Pr(e)/1 – Pr(e), where Pr(a) is the relative observed agreement among raters, and Pr(e) is the hypothetical probability of chance agreement. If there is complete agreement then κ = 1. Sensitivity and specificity were calculated by the following equations: sensitivity = (number of animals disease positive/number of animals test negative + animals disease positive) and specificity = (number of animals disease negative/number of animals test positive + animals disease negative)
The APIA developed herein used recombinant Nc-SAG1 expressed in E. coli. This antigen was completely expressed as an insoluble form and did not appear in the supernatant before urea treatment (Fig. 2A, lane 3) although the removal of the hydrophobic signal peptide and C-terminus had been reported to improve its hydrophilicity and solubility. 8 Results of the current study are in accordance with a previous report 16 in which the authors also purified recombinant Nc-SAG1 using denaturing conditions. The molecular weight of recombinant Nc-SAG1 was estimated at approximately 35 kDa by sodium dodecyl sulfate–polyacrylamide gel electrophoresis analysis (Fig. 2A). The identity of the purified protein was confirmed by Western blot with a positive sera from a N. caninum– inoculated cow (lane 3) using the pre-immune serum as control (lane 2; Fig. 2B). The recombinant Nc-SAG1 did not cross-react with a bovine serum positive to Toxoplasma gondii (Fig. 2, lane 1) as previously reported. 8
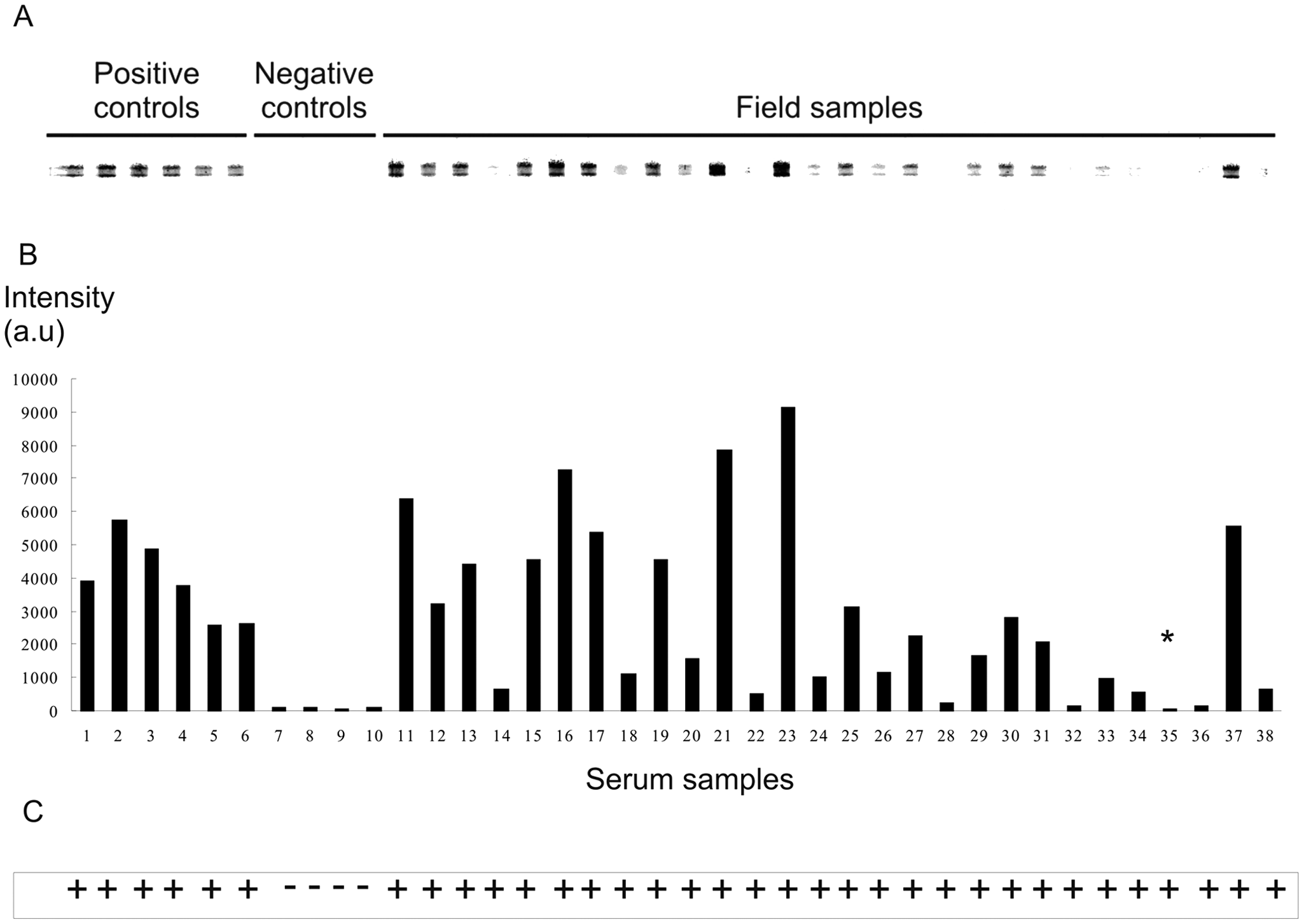
Antigen print immunoassay (APIA).
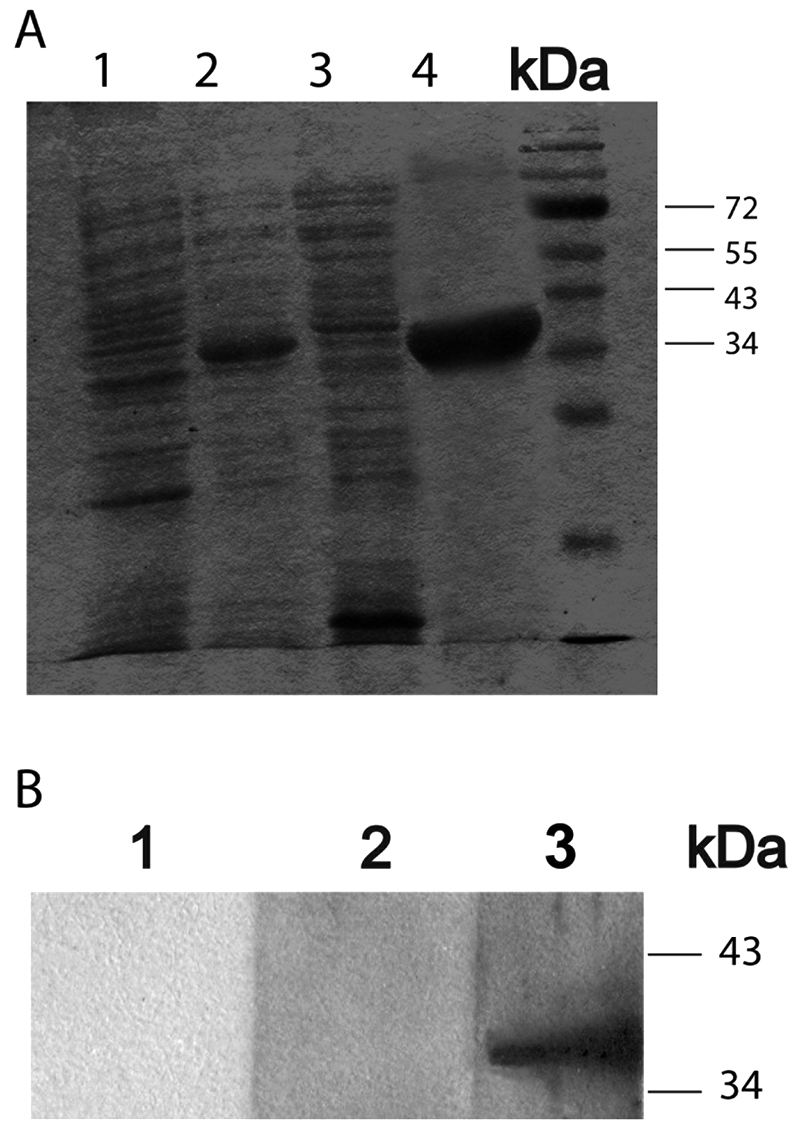
Purification of Neospora caninum surface antigen 1 (Nc-SAG1) and Western blot analysis of recombinant Nc-SAG1.
In order to analyze the performance of the APIA, the membrane was tested with sera of experimentally and naturally infected cattle. From a total of 228 sera analyzed, 16 corresponded to a “gold standard” panel of samples of cattle inoculated with 3 strains of N. caninum from different geographical origin (Table 1). The recombinant Nc-SAG1 was recognized by antibodies from cattle inoculated with strains from the United States, Argentina, and Spain. This result confirms previous findings that the Nc-SAG1 protein is antigenically conserved among geographically distant isolates. 18 The results showed that APIA detected 15 positive samples out of the 16 sera. In these samples, when APIA results were compared with ELISA, the latter showed to be less sensitive and detected antibodies in only 13 out of the 16 sera (Table 1). In view of the findings, the 16 sera were also analyzed by IFAT, which detected all samples as positive (Table 1). According to the data, IFAT performed better than ELISA, which is in accordance with a previous report on N. caninum 26 in which the authors compared the performance of a new agglutination test, ELISA, and IFAT. According to the authors, the latter performed perfectly for the experimentally infected animals but gave either false-positive or -negative results for the naturally infected animals. However, IFAT is a laborious and time-consuming technique that also requires the production of live tachyzoites. In contrast, ELISA can simultaneously process a large number of sera and has higher sensitivity and specificity values compared to IFAT. For these reasons, ELISA was selected for the further analysis of a larger and more diverse serum panel of 212 field samples from central Argentina. The samples came from herds with isolated cases of abortion. From the 212 total samples, 40 were positive and 160 were negative by both techniques, respectively. Eight samples were false negatives and 4 were false positives by APIA. A seroprevalence of 26% in this region was estimated from the results. This value is in agreement with a previous report 24 in which a prevalence of 25% was found for herds with history of reproductive losses in the same area.
Summary of name, geographical origin, and serological status of the 16 experimentally infected cattle included in the current study.*
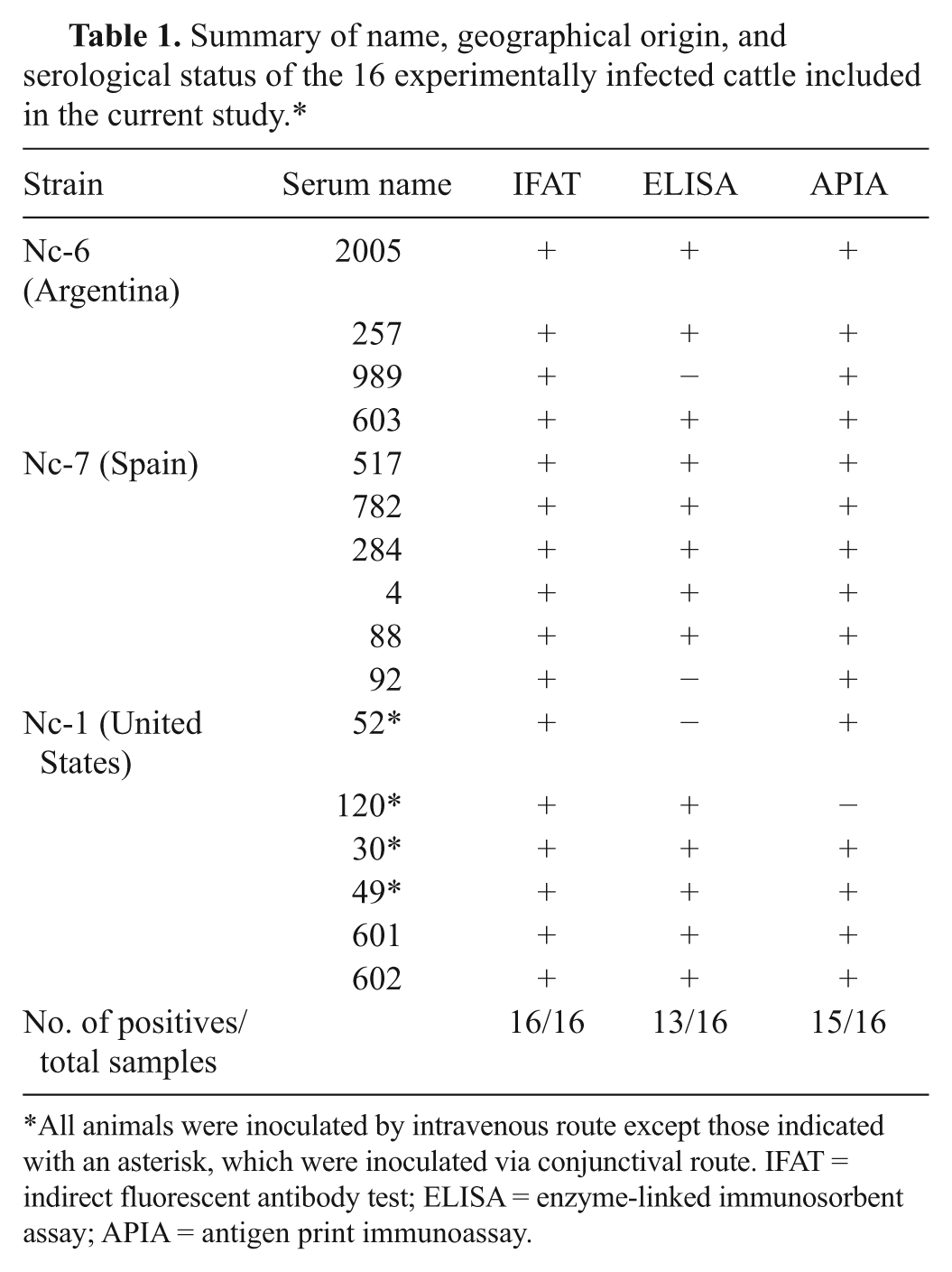
All animals were inoculated by intravenous route except those indicated with an asterisk, which were inoculated via conjunctival route. IFAT = indirect fluorescent antibody test; ELISA = enzyme-linked immunosorbent assay; APIA = antigen print immunoassay.
Comparison between antigen print immunoassay (APIA) and enzyme-linked immunosorbent assay (ELISA) analysis for the diagnosis of neosporosis in cattle.*
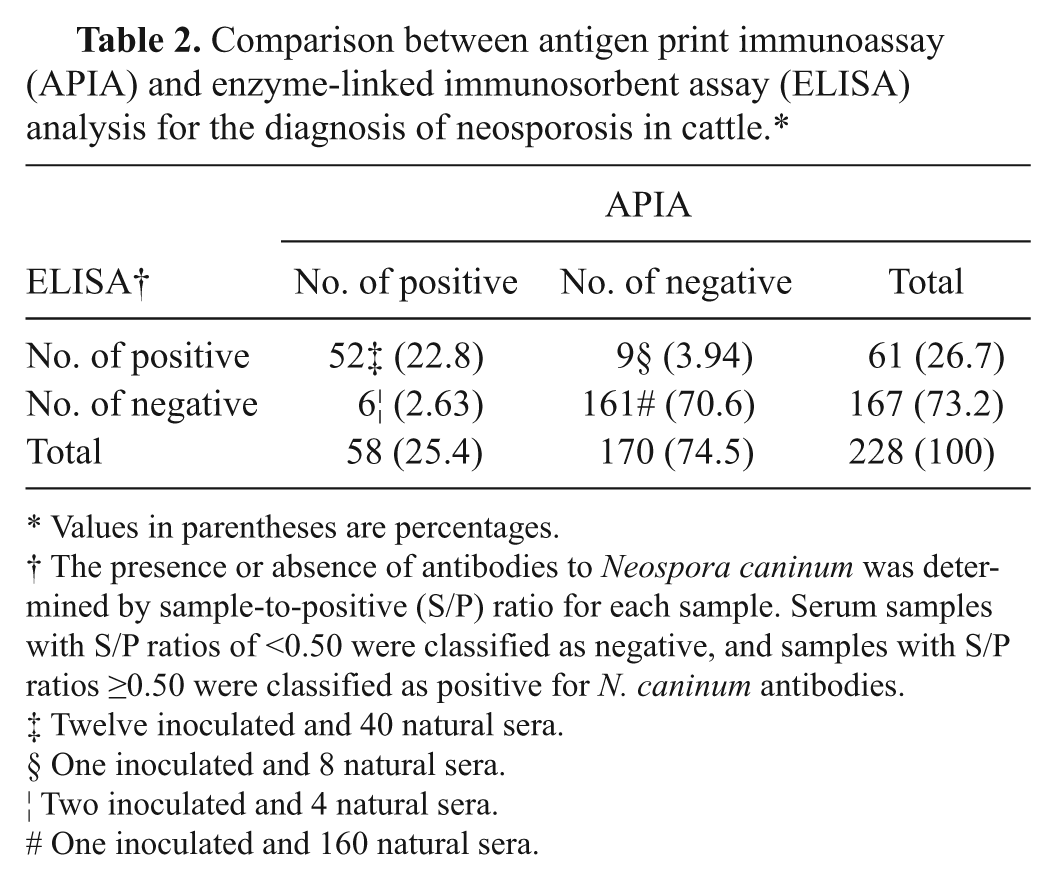
Values in parentheses are percentages.
The presence or absence of antibodies to Neospora caninum was determined by sample-to-positive (S/P) ratio for each sample. Serum samples with S/P ratios of <0.50 were classified as negative, and samples with S/P ratios ≥0.50 were classified as positive for N. caninum antibodies.
Twelve inoculated and 40 natural sera.
One inoculated and 8 natural sera.
Two inoculated and 4 natural sera.
One inoculated and 160 natural sera.
For concordance, sensitivity, and specificity calculation purposes, the 212 samples were analyzed along with the 16 samples of the experimentally infected heifers. The results shown in Table 2 indicate that APIA has a relative sensitivity of 85% (95% confidence interval [CI]: 73–92) and a specificity of 96% (95% CI: 91–98). The degree of agreement between APIA and ELISA was estimated by calculating the kappa value (http://faculty.vassar.edu/lowry/VassarStats.html). The kappa value of 0.83 (95% CI: 0.74–0.91) indicates a very good agreement between APIA and ELISA. Reproducibility of APIA was 100% since the 2 trained readers that examined each membrane were in total agreement. Although a large number of sera will be required to validate APIA performance, the results suggest that APIA with recombinant Nc-SAG1 would be a reliable tool for a first screening of antibodies to N. caninum in cattle.
The choice of nitrocellulose membrane as the solid phase offers several advantages. First, nitrocellulose membranes have a higher protein-binding capacity than polystyrene microtiter plates. 19 Moreover, nitrocellulose membrane is amenable to simultaneous testing of many antigens, 22 and in the current test, a maximum of 40 sera. Visual evaluation of APIA results is simple and rapid. Although the interpretation of results could be considered subjective, the results of 2 independent readers were in almost complete accordance with the ELISA results and the densitometric analysis of the membranes as it is shown in Figure 1. This “yes-or-no” result is suitable for efficient sera screening and is highly desirable for use in low-income, low-technology countries because the membrane can be manufactured in a core facility and easily shipped at room temperature to distant laboratories in which only the miniblotter, reagents, and a minimum of immunological expertise is required. The cost of APIA was estimated at US$1 per sample, which is acceptable for veterinary testing. Although APIA is not intended to be a quantitative diagnostic test, this membrane-based assay can easily evolve into a low-cost multi-antigen test that includes several antigens of N. caninum or other bovine pathogens coexisting in the same area that can be analyzed simultaneously as a first screening step.
Footnotes
Acknowledgements
Silvina E. Wilkowsky, Dadin Prando Moore, and María I. Romano are researchers of the National Research Council of Argentina (CONICET). The authors thank Drs. M. C. Venturini and G. Moré from Universidad de La Plata, Argentina, for kindly providing Toxoplasma gondii–positive sera.
a.
Strains Nc-1 and Nc-6 kindly provided by Dr. M. C. Venturini, Veterinary College, La Plata National University, Argentina.
b.
Strain Nc-7 kindly provided by Dr. L. M. Ortega-Mora, Complutense University of Madrid, Spain.
c.
Puregene® DNA Purification Kit, Qiagen Inc., Valencia, CA.
d.
pCR-TOPO 4, Invitrogen Corp., Carlsbad, CA.
e.
pRSET C, Invitrogen Corp., Carlsbad, CA.
f.
BigDye® Terminator v3.1 Cycle Sequencing Kit, ABI 3130XL; Applied Biosystems, Carlsbad, CA.
g.
ProBond Nickel-Quelatin Resin, Invitrogen Corp., Carlsbad, CA.
h.
IDEXX Neospora X2 Ab Test, IDEXX Laboratories Inc., Westbrook, ME.
i.
Nikon Fluophot 40 1.3, Nikon Instruments Inc., Melville, NY.
j.
Bio-Rad Laboratories, Hercules, CA.
k.
Linomat IV, CAMAG, Muttenz, Switzerland.
l.
Isogen Biosolutions, De Meern, The Netherlands.
m.
Hybond-ECL, GE Healthcare, Pittsburgh, PA.
n.
Image Quant TL Array Version 7.0 Software, GE Healthcare, Pittsburgh, PA.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
This work was financially supported by a grant from Biotech MERCOSUR UE 1271117.
