Abstract
The
Various methods have been previously described to detect REV in avian blood and tissue samples, including virus isolation, immunohistochemistry, in situ hybridization, enzyme-linked immunosorbent assay, and fluorescent antibody test, but they can be time consuming (e.g., several weeks for virus isolation). 12 In comparison, molecular techniques such as polymerase chain reaction (PCR) have been used in the diagnosis of various infectious diseases and are considered one of the best alternatives to conventional assays because of their sensitivity, specificity, and reproducibility.15,17 Conventional gel-based PCR tests,5,6,10 as well as a nested PCR assay, 2 have been developed for routine REV diagnostics; however, these methods require manipulations post-PCR amplification including gel electrophoresis, gel staining steps, or transfer of amplified products to secondary tubes for further amplification. These methods can be time consuming and are prone to carryover contamination.
The objective of the present study was to develop a high-throughput process for the rapid diagnosis of REV with high sensitivity and specificity. To accomplish this, a duplex real-time PCR assay was designed to detect both the envelope protein gene (
For the real-time PCR assay described herein, primers and probes were designed using commercially available software
a
based on published sequences of the REV
Two DNA extraction methods were compared in the current study: 1) phenol-chloroform and 2) a microsphere paramagnetic bead kit
d
and processor.
e
The phenol-chloroform extraction was performed according to standard methods for the isolation of DNA. Briefly, 10 µl of whole blood was added to 500 µl of 1× STET buffer
f
(100 mM NaCl, 10 mM Tris–HCl [pH 8.0], 1 mM ethylenediamine tetra-acetic acid [pH 8.0], and 5% Triton X-100) and 20 µl of proteinase K
g
(20 mg/ml) and incubated for 15 min at 56°C. Then, 500 µl of buffered phenol-chloroform–isoamyl alcohol 25:24:1
f
was added, and the sample was vortex mixed and then centrifuged for 5 min at 12,000 ×
The duplex real-time PCR reactions were performed using a commercially available kit,
h
with 10 pmol of each
Conventional gel-based PCR for detection of REV LTR was performed using a previously described method
6
with minor modifications. Briefly, the PCR reaction mixture consisted of 1.5 mM MgCl2, 200 µM of each deoxyribonucleotide triphosphate (dNTP), 1.25 U of
The specimens included in the present study consisted of 176 avian blood samples (168 Attwater’s prairie chickens, 6 chickens, 1 duck, and 1 pheasant) submitted to the Texas Veterinary Medical Diagnostic Laboratory (College Station, Texas) for various reasons, such as health checks or disease diagnostics, 27 of which had been confirmed REV positive and 149 negative by conventional gel-based PCR. Whole blood samples were stored at −20°C following initial screening. For the current study, blood samples were thawed and DNA extracted using both the phenol-chloroform and magnetic bead methods and tested by the newly developed duplex real-time PCR. Additionally, to test the specificity of the assay, DNA extracted from 9 Marek’s disease virus (MDV)-positive and 16
The limit of detection of the duplex real-time PCR was determined using a plasmid containing the full-length REV proviral DNA diluted 10-fold serial dilutions (10−1–10−9). In addition, the detection limit of the duplex real-time PCR was also compared to the conventional gel-based PCR using DNA extracted from serial 10-fold dilutions (10−1–10−8) of an REV-positive whole blood sample. Plasmid containing REV proviral DNA was purified from bacterial culture using a plasmid mini prep kit,
g
and the DNA concentration was determined.
o
Results show that the limit of detection for the duplex assay was 46 and 9 copies for
Comparison of the
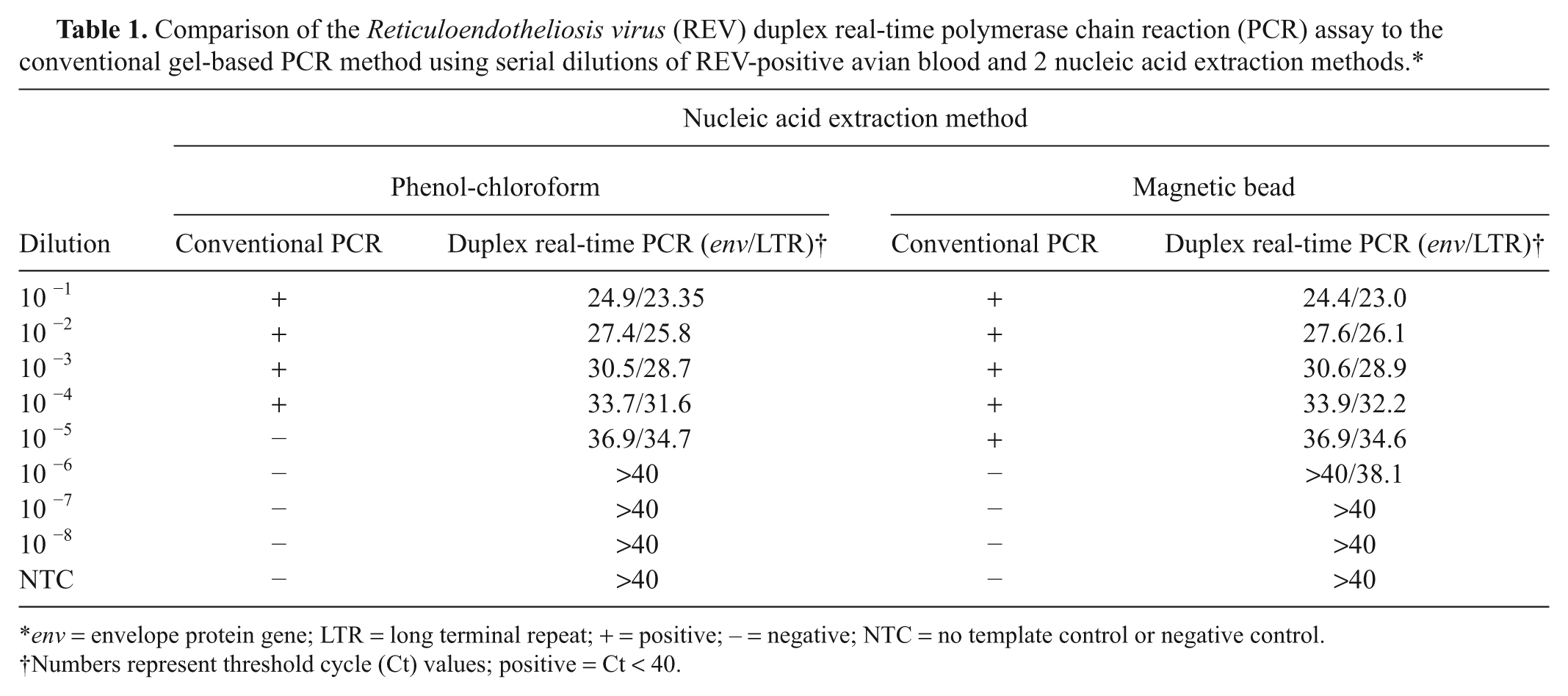
Numbers represent threshold cycle (Ct) values; positive = Ct < 40.
The diagnostic sensitivity and specificity of the duplex real-time PCR assay was determined by comparison of DNA extracted from 176 avian blood samples previously screened for the presence of REV using the standard gel-based assay (27 REV positive, 149 REV negative).
6
Results show all 27 previously identified positive samples were positive for both
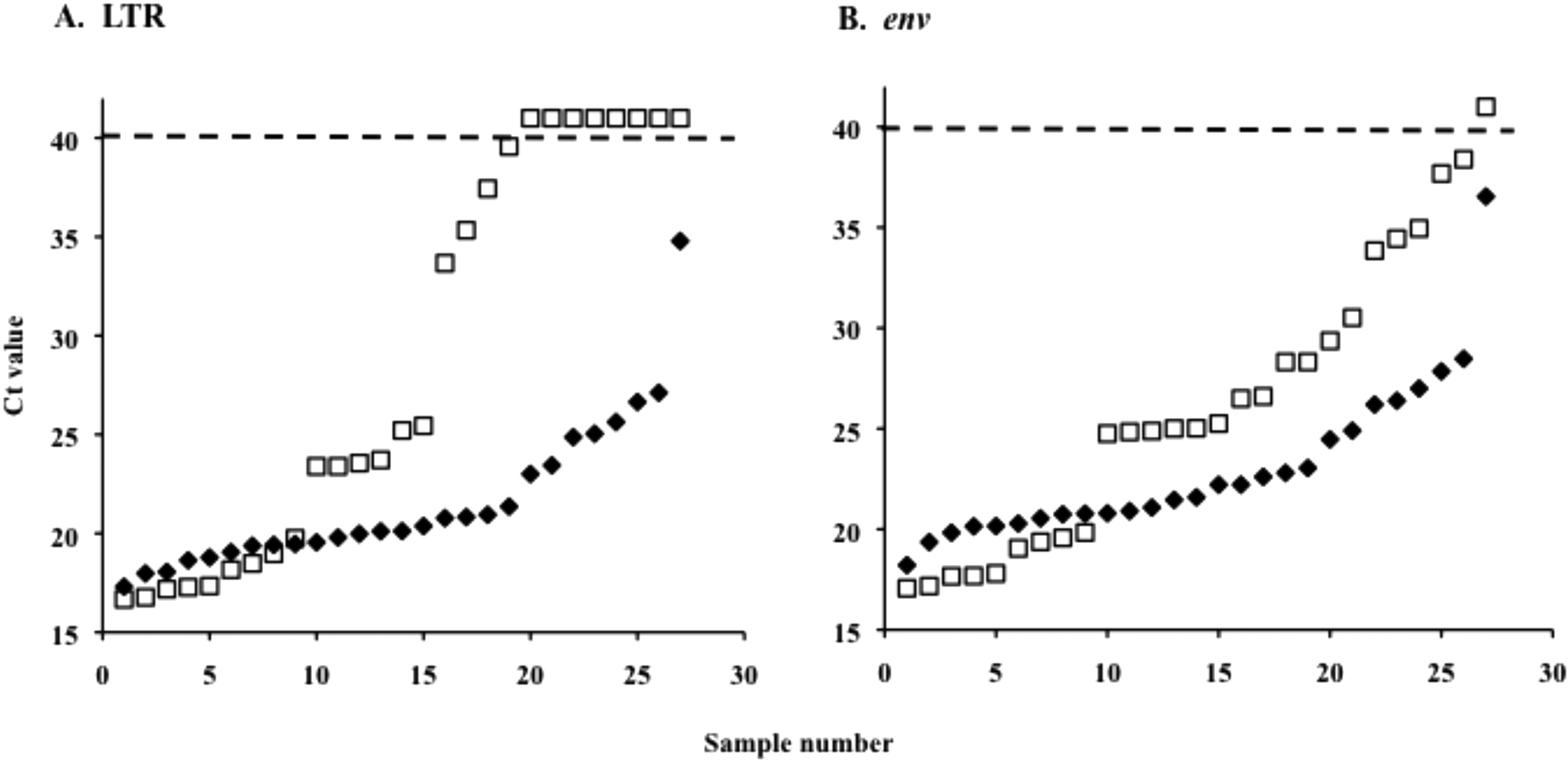
Comparison of 27
Decision makers rely on diagnostic tests to aid in management decisions regarding disease status.
Since REV is not a widespread problem in poultry and, when detected, terminal methods of control are implemented, a limitation in available REV-positive test specimens existed. The unique situation with the Attwater’s prairie chicken allows for the availability of REV-positive field samples from the captive flock. Although the majority (95.5%) of samples tested and all REV-positive samples were from Attwater’s prairie chickens, the assay could be applied to other species and specimens. Genomic sequences among REV isolates have been shown to have little variability, emphasizing the applicability of PCR-based tests for the detection of REV to various avian species.6,9,18 Additionally, the currently used assay for the detection of REV in clinical specimens was designed for use in poultry and is routinely used for other avian species, including the Attwater’s prairie chicken.
In conclusion, the use of real-time PCR increased the diagnostic sensitivity of the assay 10-fold compared to the traditional gel-based PCR assay, regardless of extraction method. Combining the magnetic bead extraction method and the real-time assay increased the diagnostic sensitivity 100-fold compared to the traditional phenol-chloroform extraction with the gel-based PCR assay. The ability of the duplex assay to detect fewer copies of LTR compared to
Although not shown in the present study, the duplex real-time PCR assay described herein potentially has broader application since the REV proviral DNA is known to integrate in part (LTR only) or in whole into the genome of the infected host and large DNA viruses such as FPV and MDV.4,10,14,19,20,24 Thus far, the genomes of commercial vaccine strains of FPV and MDV field strains have been shown to contain complete or truncated REV-LTR, but not REV-
The combination of nucleic acid isolation with magnetic beads and real-time PCR has been shown to greatly increase testing capacity of other avian pathogens, such as
Footnotes
Acknowledgements
The authors wish to thank Jennifer Meier for her expert technical assistance. This work was a collaborative effort between the Texas Veterinary Medical Diagnostic Laboratory (TVMDL), the Texas A&M University College of Veterinary Medicine (TAMU-CVM), the U.S. Fish and Wildlife Service Attwater Prairie Chicken National Wildlife Refuge, and the Houston Zoo, Inc. The findings and conclusions in this article are those of the author(s) and do not necessarily represent the views of the U.S. Fish and Wildlife Service.
a.
Primer Express™ 3.0, Applied Biosystems, Foster City, CA.
b.
Sigma Genosys, Sigma-Aldrich, St. Louis, MO.
c.
TaqMan®, Applied Biosystems, Foster City, CA.
d.
MagMAX 96 Viral RNA Isolation Kit, AM1836; Applied Biosystems, Foster City, CA.
e.
KingFisher 96, Thermo Scientific, Waltham, MA.
f.
Fisher Scientific, Pittsburg, PA.
g.
Qiagen Inc., Valencia, CA.
h.
FastStart Universal Probe Master (Rox), Roche Applied Science, Indianapolis, IN.
i.
ABI Prism 7000 and 7500 Fast Real-Time PCR detector, Applied Biosystems, Foster City, CA.
j.
TOPO-TA cloning® kit, Invitrogen Corp., Carlsbad, CA.
k.
BigDye Terminator v3.1 Cycle Sequencing Kit, ABI 3100 Genetic Analyzer; Applied Biosystems, Foster City, CA.
l.
Promega Corp., Madison, WI.
m.
GeneAmp PCR System 9700, Perkin-Elmer, Waltham, MA.
n.
GelRed™, Phenix Research Products, Candler, NC.
o.
NanoDrop 8000, Thermo Scientific, Waltham, MA.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) received no financial support for the research, authorship, and/or publication of this article.
