Abstract
To investigate the prevalence and genetic characterization of
Introduction
The first
A 2009 report describes PiCV infections in the pigeon population in eastern China. 28 However, the genetic characterization of the PiCV strains in the region is not known. The objective of the present study was to characterize the prevalence and the genetic features of the PiCV strain that was infecting the pigeon population in eastern China.
Materials and methods
Sample sources
A total of 104 infected pigeons displaying clinical symptoms (such as respiratory signs, weight loss, and diarrhea) and 40 healthy pigeons were collected from 12 different regions in 6 provinces (Shandong, Anhui, Zhejiang, Shanghai, Fujian, and Jiangsu) in eastern China between 2009 and 2010 (Table 1). The 12 regions were chosen because they had the highest pigeon population density in China. The tissue samples were collected from the pigeons and immediately frozen at 80°C.
Molecular characterization and epidemiological investigation of
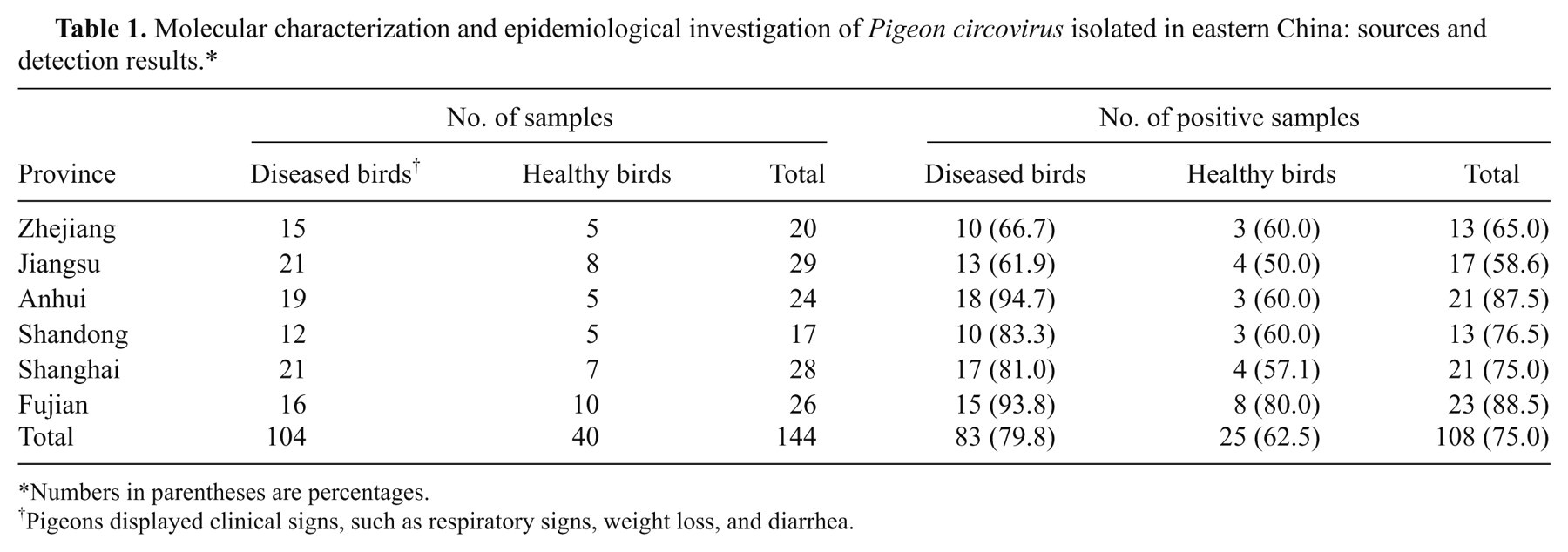
Numbers in parentheses are percentages.
Pigeons displayed clinical signs, such as respiratory signs, weight loss, and diarrhea.
Nucleic acid isolation
The genomic DNA was extracted from 200 µl of tissue homogenate (10%, w/v) or from approximately 25 mg of tissue samples using a commercial kit. a The extracted DNA was eluted twice with 25 µl of sterile distilled water to give a final extract volume of 50 µl.
The detection of PiCV by polymerase chain reaction
To detect the presence of PiCV sequence, the extracted genomic DNA was PCR amplified using PiCV-specific primers (forward: 5’-TTGAAAGGTTTTCAGCCTGGC-3’; reverse: 5’-AGGAGACGAAGGACACGCCTC-3’) as previously described. 4 The expected PCR product was 325 base pairs (bp) in size. The PCR mixture (50 µl) contained 40.5 µl of water, 5 µl of 10× buffer, 1 µl of each deoxyribonucleotide triphosphate (2.5 mM each), b 2 µl primers (10 µM each), 1 µl of template DNA (extracted as described above) and 0.5 µl of Taq DNA polymerase (5 U/µl). b The PCR reaction was conducted in a thermal cycler c using the following conditions: denature for 5 min at 94°C, followed by 40 cycles of 94°C for 30 sec, 60°C for 30 sec, and 72°C for 30 sec, and elongation for 5 min at 72°C. The amplicons were resolved by electrophoresis in a 2% agarose gel, which was stained with ethidium bromide (0.5 g/ml) in Tris–borate–ethylenediamine tetra-acetic acid buffer (89 mM Tris; 89 mM boric acid; 2 mM ethylenediamine tetra-acetic acid, pH 8.4) and visualized under ultraviolet light.
Inverse primer polymerase chain reaction on the PiCV genome
A positive DNA sample extracted above from each region were amplified with inverse primer polymerase chain reaction (IP-PCR) using the inverse primer pair 5’-CCCCACTGAAGAAGAGATTAAGAGCCTGGAAA CGTG-3’ (forward) and 5’-CTTAATCTCTTCTTCAGTGGGGTT GTTCAA-3’ (reverse). Six full-length clones were obtained in the current study, and each PCR product was approximately 2,000 bp in size. The IP-PCR reagents were the same as described above, and the amplification conditions were set as following: 30 cycles of 94°C for 45 sec, 53°C for 1 min, and 72°C for 2 min with an initial denaturation of the template DNA at 94°C for 5 min. The PCR products with the expected sizes were purified and sequenced after subcloning into the pMD18-T vector with a Taq-amplified cloning kit according to the manufacturer’s protocol. b
The phylogeny of the PiCV strains
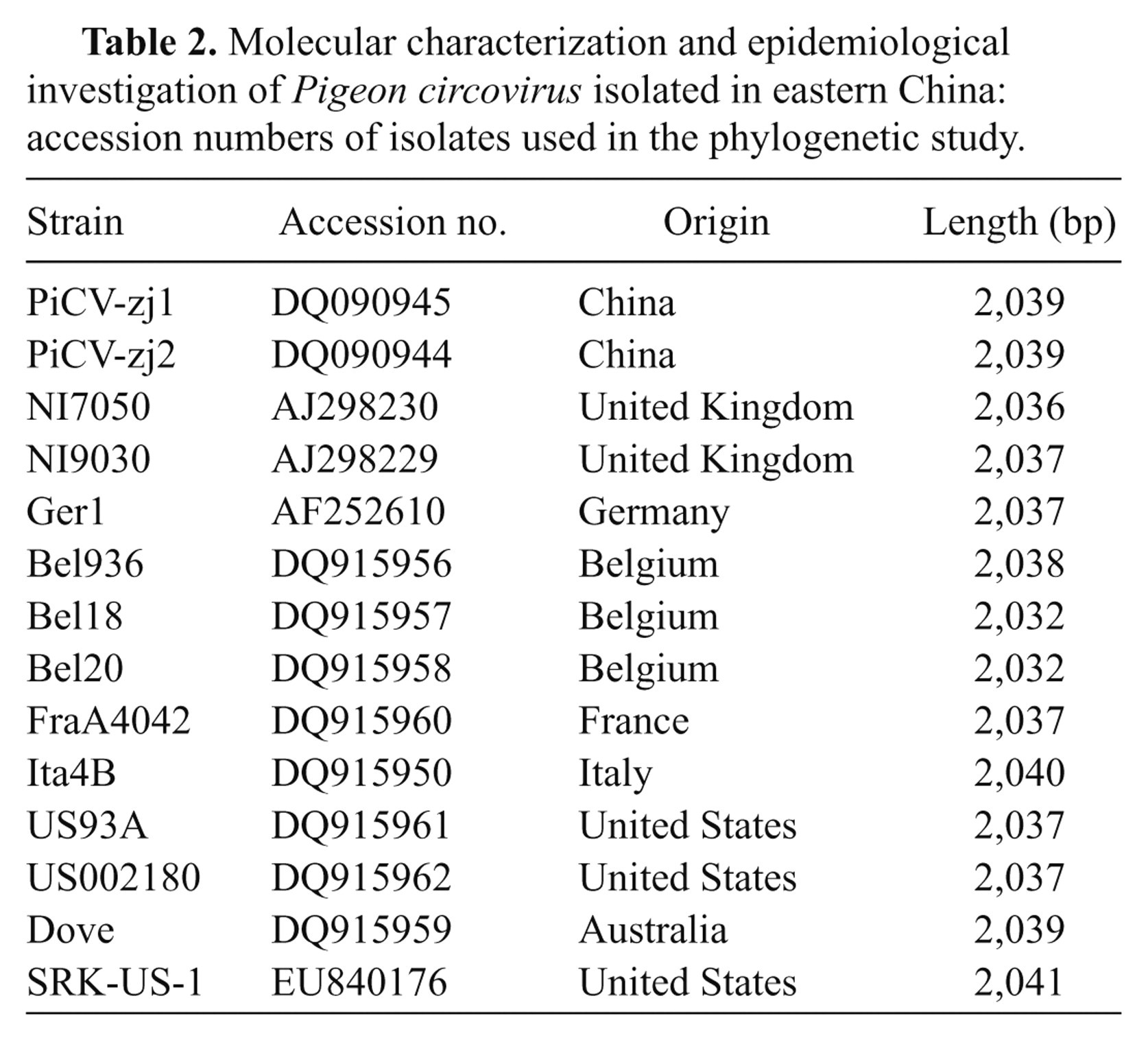
The DNA sequences of the 6 PiCV strains obtained in the current study were compared with 14 published full-length PiCV sequences from the GenBank database (Table 2) using the sequence analysis software DNASTAR d and ClustalW. 6,20 The phylogenetic tree was generated using the distance-based, neighbor-joining program MegAlign. 6,10 The bootstrap values were calculated on 1,000 replicates of the alignment.
Molecular characterization and epidemiological investigation of
The amino acid sequence analysis of the replication association and capsid proteins
The amino acidic sequences of the replication-associated (
Results
Polymerase chain reaction testing of the clinical samples
A specific DNA fragment was PCR-amplified to produce the amplicons with expected size. The amplicons were then purified, sequenced, and further confirmed by BLAST (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi). A total of 108 samples out of 144 tissue field samples were positive for PiCV (Fig. 1). Among the PiCV-positive samples, 83 were detected in 104 diseased pigeon samples (79.8%), and 25 were detected in 40 healthy pigeon samples (62.5%).
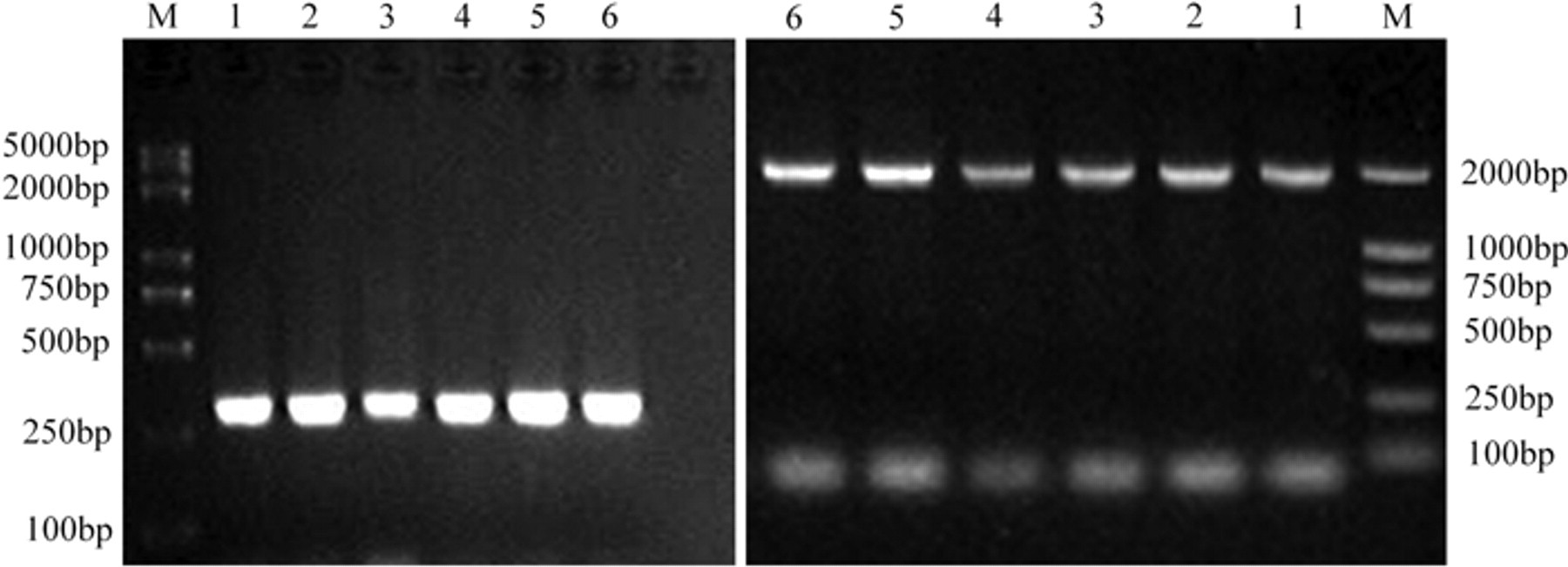
The result of
Inverse amplification of the PiCV genome
The IP-PCR technique was performed using PiCV-positive samples to produce 6 genomic DNA fragments (Fig. 1), which were gel-purified and sequenced. The sequences were confirmed by BLAST and designated as SHWH-AB4 (2,031 bp), NJPK-21 (2,035 bp), HBLF-E2 (2,031 bp), JSJN (2,039 bp), SDDZ (2,037 bp), and AHBZ (2,035 bp).
The phylogeny of the PiCV
A phylogenic tree was constructed using the MEGALIGN software program d for the 6 PiCV genomic sequences identified in the present study and the 14 additional PiCV sequences published in the GenBank database (Fig. 2). The PiCV phylogenic tree was divided into 2 branches. All 6 strains detected in the current study belonged to the same branch. The Dove strains that were isolated from Australia solely constituted the second branch. In the dendrogram, the sequences of the 3 isolates (DQ090945 PiCV-zj1 China, JSNJ China Seq, and DQ090944 PiCV-zj2 China) formed a cluster and collectively differed from the remaining 5 isolates.
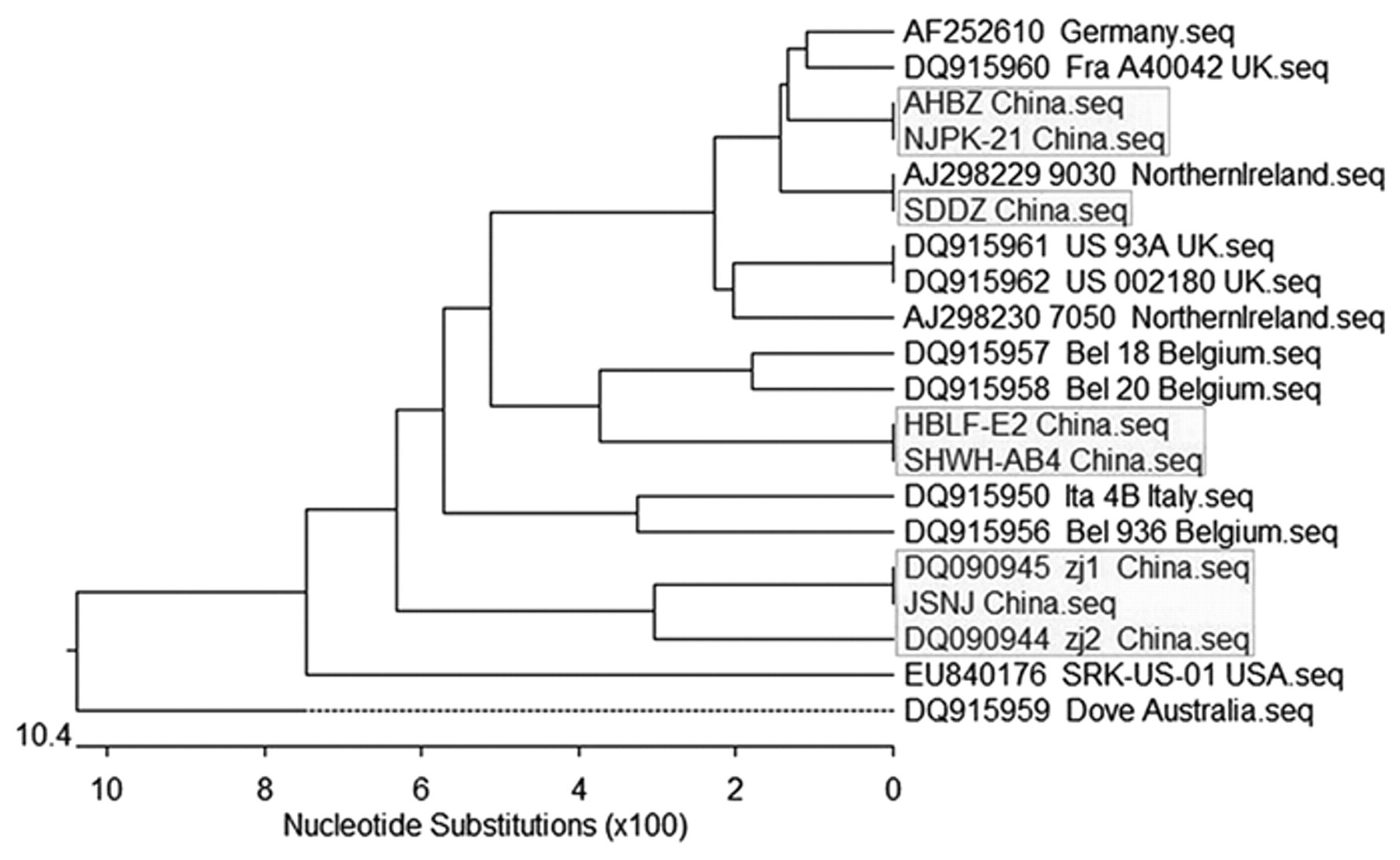
Phylogenetic tree based on the comparison of entire
The amino acid sequence analysis of the Rep and Cap proteins
The amino acid sequences were translated from the sequenced PiCV genome using the ClustalW program. The
Pairwise comparison of twenty
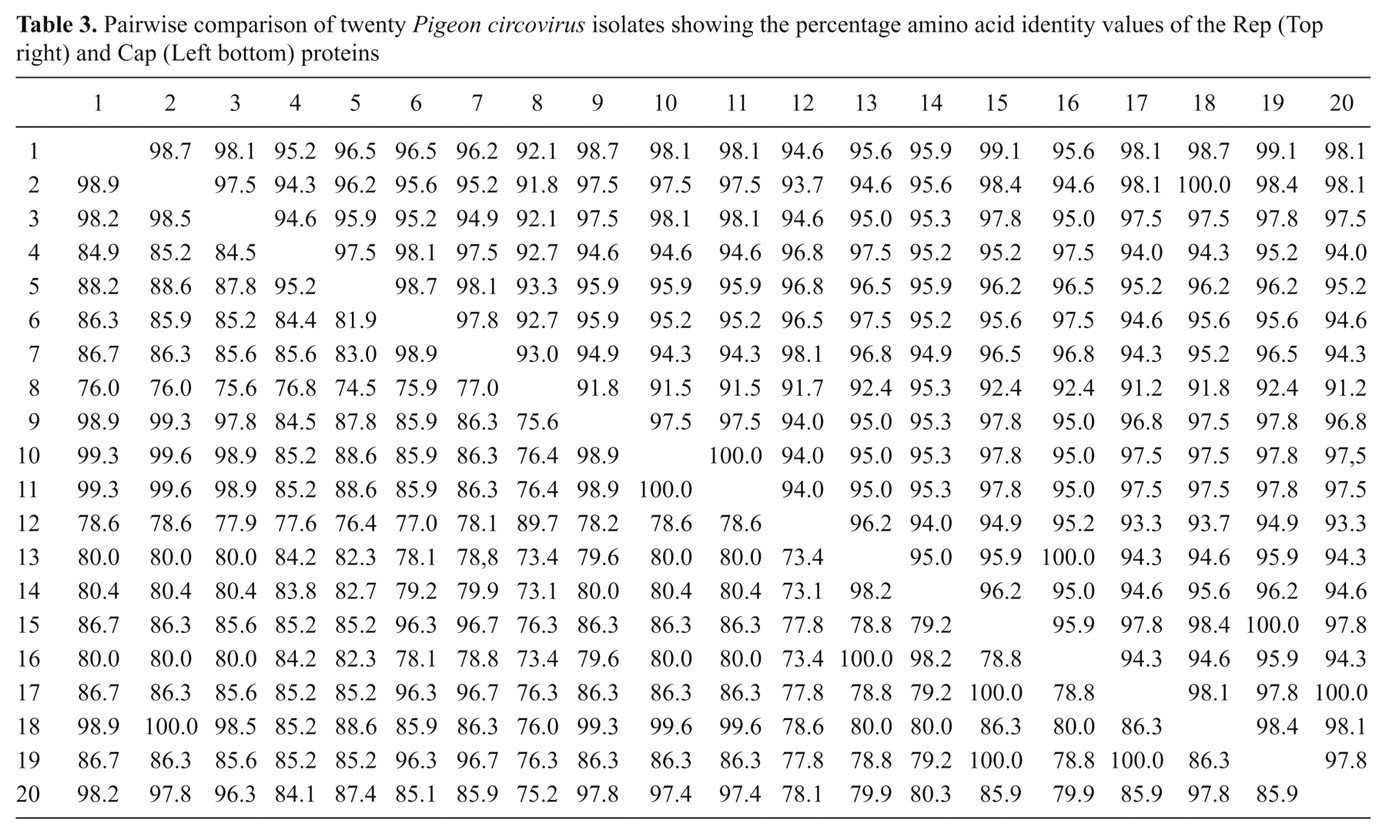
The
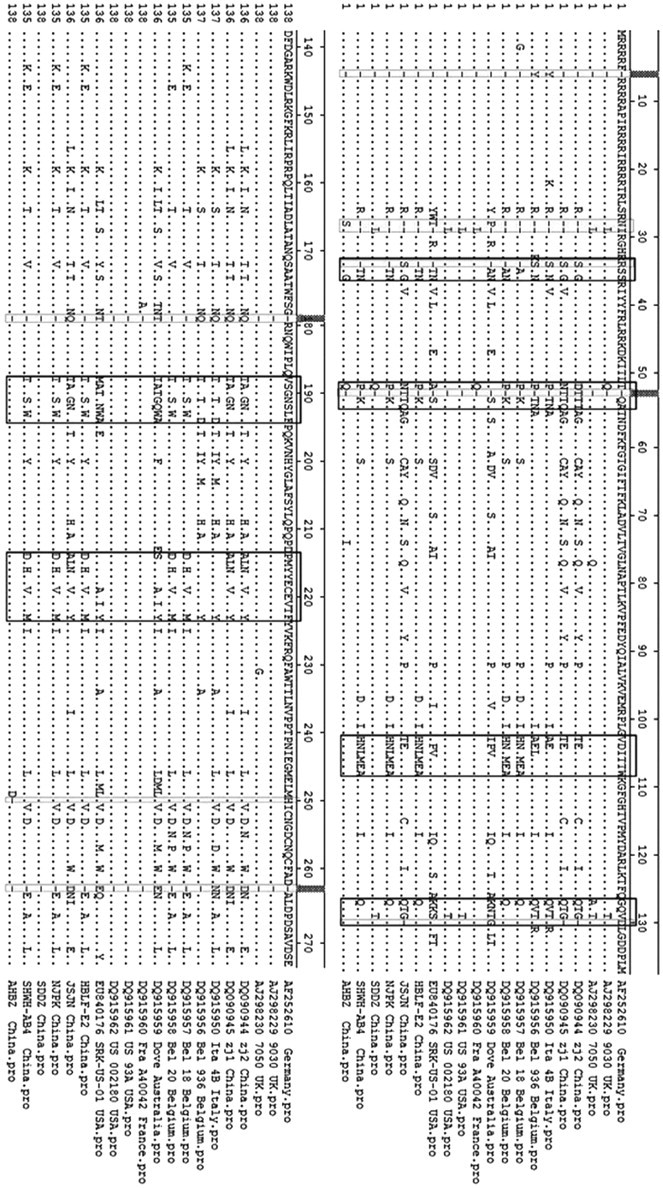

Alignment of the full-length amino acid sequences of
The size of the

Alignment of the full-length amino acid sequences of
Discussion
The genetic diversity of the virus may lead to variations in their pathogenicity.
The major advantage of IP-PCR is the ability to amplify an unknown flanking region using 2 specific primers, which could avoid sequence assembly and minimize interference from other circovirus-like agents. A 2001 study found that the positive rate of PCV-2 detection using PCR was enhanced by the amplification of PCV-2 ORF2 rather than the amplification of PCV-2 ORF1 or the complete PCV-2 genome. 24 In the same study, the authors suspected that contaminants having similar sequences to the PCV-2 ORF2 were present in porcine sera and, therefore, may have led to PCR interference. 25
Pairwise comparisons showed that the genome sequence of PiCV detected from samples isolated from pigeons in eastern China appeared to be closely related. The sequence identity was found to be 98.9%–100%, and the genome size ranged from 2,031 nt (SHWH-AB4 and HBLF-E2) to 2,039 nt (JSJN, PiCV-zj1, and PiCV-zj2). The PiCV-zj1, PiCV-zj2, and NJPK strains were isolated from the Zhejiang Province, while the JSNJ strains were isolated from Jiangsu Province. However, the PiCV-zj1, PiCV-zj2 strains (2005) were found closely related to the JSNJ strains (2009), but not from the NJPK strains (2009). Results indicated that geographical and time factors had almost no effects on the genome diversity of PiCV.
The phylogenic tree and aa sequence comparison found that the epidemic PiCV strains in eastern China had low genomic separation and diversity. The variation was mainly due to the variability within the
Footnotes
a.
QIAamp® DNA Mini Kit, Qiagen Ltd., West Sussex, United Kingdom.
b.
Takara Bio Inc., Otsu, Shiga, Japan.
c.
Eppendorf Canada, Mississauga, Ontario, Canada.
d.
DNASTAR Inc., Madison, WI.
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The current study was supported by the Program for New Century Excellent Talents (NCET) in the University (no. NCET-08-0794) of China, the Qing Lan Project, China, Natural Science Foundation of Jiangsu Province (BK2008059), Research Foundation of Jinling Institute of Technology (no. Jit-n-2009-011), Natural Science Foundation of the Higher Education Institutions of Jiangsu Province, China (no. 10kjd230003), and Graduate Research and Innovation Project of Jiangsu Province (CX09B_ 246Z).
