Abstract
A listeriosis outbreak, in dairy cattle, with a high case mortality and acute death after onset of symptoms was investigated using gross pathology and bacteriologic approaches, including molecular characterization of a clinical Listeria monocytogenes isolate. In a herd of 315 animals, 9 animals showed clinical symptoms consistent with listeriosis, including 3 animals that died within 2–4 days after acute onset of clinical signs, 4 animals that were euthanized, and 2 that survived. Initial EcoRI ribotyping and serotyping indicated that this outbreak was caused by an unusual L. monocytogenes serotype 4b strain, which was classified into lineage III. Further characterization of this isolate by DNA sequencing–based subtyping methods indicated that the strain responsible for this outbreak represented a unique genotype as supported by its classification into a new sigB allelic type, which has not been identified previously among >290 isolates, and by compelling phylogenetic evidence. While lineage III isolates are generally rare, they seem to be more common among L. monocytogenes isolates from animals with clinical signs of listeriosis. This is the first report of a particularly severe clinical course of disease associated with infection by a lineage III strain. The high prevalence of Listeria spp., including L. monocytogenes, in the farm environments may favor emergence and evolution of novel, and possibly more virulent, L. monocytogenes strains. Continued monitoring of animal listeriosis cases and outbreaks may not only improve animal health but also aid in the early discovery of newly emerging L. monocytogenes strains.
Listeria monocytogenes is a Gram-positive bacterial pathogen of both human beings and animals commonly isolated from farm environments. 15 The bacterium can be classified into 4 genetic lineages (I–IV) 22 and 13 serotypes 18 ; the vast majority of human listeriosis cases (i.e., >90%) are caused by lineage I serotype 1/2b and 4b and lineage II serotype 1/2a strains. 9 While lineage III and IV strains are generally rare, they have been involved in a number of animal listeriosis cases and even in some outbreaks. 17,27 Most listeriosis outbreaks in livestock have been linked to contaminated silage. 5,16,25–27 While plant materials can contain L. monocytogenes, silage contamination can also originate from contamination of the raw plant material with soil, wildlife or bird feces, or manure. In addition to initial contamination, growth of L. monocytogenes in silage is typically needed to reach pathogen levels likely to cause disease. In improperly fermented silage with a pH >5, growth typically can occur 15 ; L. monocytogenes numbers in poorly fermented silage can be as high as 108 colony-forming units (CFU)/g wet weight of silage. 27 Ingestion of contaminated silage can result in listeriosis with invasion occurring in the intestinal tract or possibly in the oral cavity. It has been suggested that oral lesions in young animals that are changing teeth may be a point of entry for L. monocytogenes, which is then thought to migrate from the oral cavity along cranial nerves to the brainstem, resulting in listerial encephalitis. 20,26
While our understanding of the transmission of Listeria in ruminants is still limited, 23 several studies on smaller outbreaks in cattle and small ruminants have contributed to the general understanding of listeriosis in ruminants. 5,15,16,21,25,26 During an outbreak, disease prevalence in cattle is usually comparably low, in the range of 8%–10%, although it can reach 15%, 23 and case mortality rates can range from 20% to 100%. 14 Subclinical infections are also known to exist, and multiple studies have shown that up to 50% of feces collected from animals without clinical symptoms contain L. monocytogenes. 11,23 While some studies on ruminant listeriosis have also used molecular tools to characterize the specific L. monocytogenes strains involved in a given outbreak, 6,8,21,27 additional studies are needed to improve our understanding of animal listeriosis and to better characterize L. monocytogenes strains that cause animal disease.
In February 2009, a suspected listeriosis outbreak was identified on a dairy farm with approximately 180 milking cows, 15 dry cows, and 120 calves and heifers. Over a period of 12 days, 9 animals (5 milking cows, 2 dry cows, and 2 heifers) displayed symptoms typical of encephalitic listeriosis (i.e., facial paralysis, drooling, circling, and recumbency); 3 animals died and 4 were euthanized due to severity of the clinical signs (Table 1). The time between initial detection of listeriosis symptoms and death for the fatal cases ranged from 2 to 4 days. Treatment consisted of 3 × 106 IU per animal of penicillin twice a day or 12 g per animal of florfenicol a every other day (Table 1). The affected animals were from 2 groups kept in separate areas but fed the same forages, including barley silage from an upright silo. The pH of a silage sample was 7.0, indicating improper fermentation and suggesting silage as a possible source of L. monocytogenes in this outbreak.
Description of animals with clinical symptoms associated with the outbreak in the current study. *
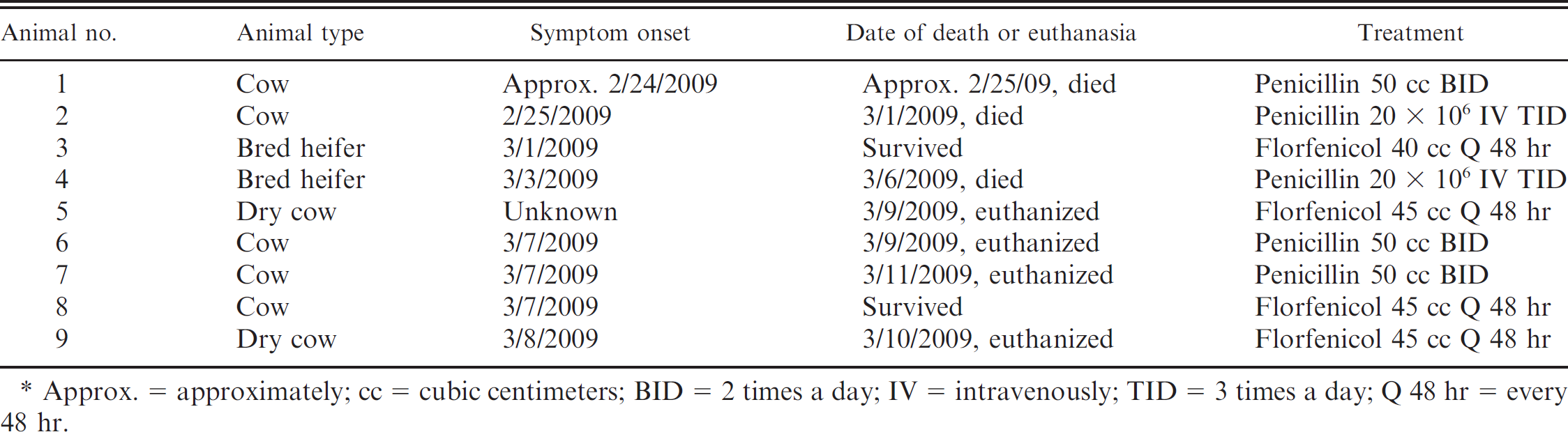
Approx. = approximately; cc = cubic centimeters; BID = 2 times a day; IV = intravenously; TID = 3 times a day; Q 48 hr = every 48 hr.
The clinical diagnosis of encephalitic listeriosis was confirmed by bacteriologic and histopathologic testing of fresh and fixed brain tissues from a single deceased animal submitted to Lynchburg Regional Laboratory (Lynchburg, Virginia). Histopathological examination of fixed brain tissue samples stained with hematoxylin and eosin showed brainstem lesions characteristic of listerial infection, 20 although the micro-abscesses, commonly associated with encephalitic listeriosis, were not recognized microscopically. Fresh brain tissue was streaked onto Columbia blood agar (CBA) b and Levine eosin methylene blue agar (EMB) b plates and incubated at 37°C for up to 48 hr; the CBA plate was incubated in 7% CO2 and the EMB plate was incubated without added CO2. Both the CBA and EMB plates had bacterial colonies consistent with L. monocytogenes colony morphology after 24 hr of incubation. A Gram stain showed short Gram-positive rods, and a catalase test and CAMP (Christie, Atkins, Munch-Petersen) test were consistent with L. monocytogenes. A corynebacteria and coryne-like organism identification kit c also identified the isolate as L. monocytogenes. A rabies fluorescent antibody test 10 was conducted by the Division of Consolidated Laboratory Services (Richmond Virginia). The result on fresh brain tissue was negative, excluding rabies as the cause of the central nervous system signs.
The bacterial isolate from 1 animal was sent to the Cornell Listeria Strain collection and deposited under the accession number FSL F6-706. It was characterized by traditional serotyping as well as molecular methods, including molecular serotyping, ribotyping, allelic typing of sigB, and multilocus sequence analysis (MLSA). Molecular serotyping, a polymerase chain reaction (PCR) assay for sigB allelic typing, and MLSA were performed using a crude lysate, prepared as previously described, as template. 7 Multiplex PCR-based molecular serotyping, which involves PCR-based amplification of genes that are unique to different serotypes as previously described, 3 yielded a single band, which did not allow for serotype assignment, suggesting that the isolate may represent an unusual L. monocytogenes strain. Serotyping with a commercial Listeria antisera kit, d performed at the U.S. Centers for Disease Control and Prevention (Atlanta, Georgia), classified this isolate as serotype 4b. Automated EcoRI ribotyping, a subtyping method that generates banding patterns after EcoRI restriction digestion of chromosomal DNA followed by Southern blot hybridization with a ribosomal RNA gene–based probe, 1 was performed using a commercial system e as previously described. 1 This method classified the isolates as ribotype DUP-1061A, indicating the isolate belonged to lineage III. Listeria monocytogenes lineage III (including IIIA and IIIC) and IV (previously designated as lineage IIIB and involved in an outbreak among goats described in an earlier study 27 ) have previously been reported among animal isolates. 17 These lineages are rare among L. monocytogenes overall but have been found to show, on average, high cytopathogenicity (i.e., cell-to-cell spread) in tissue culture assays, 24 even though some virulence-attenuated lineage III isolates have also been reported. 4 Overall, serotyping and ribotyping results are consistent, as serotype 4b is 1 of the 3 serotypes associated with lineage III. 17 While lineage I serotype 4b isolates are common, and isolates with this lineage-serotype combination are frequently associated with human listeriosis cases and outbreaks, 9 lineage III serotype 4b isolates (such as the one reported herein) are unusual and much less common.
To further characterize the unusual strain responsible for this listeriosis outbreak, 2 subtyping methods were performed: 1) sigB allelic typing, 13 which is based on DNA sequencing of a 660–base pair (bp) fragment of the sigB gene; and 2) subtyping with a 10-locus MLSA scheme, which uses partial sequences of sigB, addB, ldh, pbpA, lmo0490, lmo2763, polC, prs, rarA, and lmo1555 as described elsewhere. 2 Sequence data for these genes were obtained through PCR amplification of the target genes followed by dideoxynucleotide f sequencing at the Cornell University Life Sciences Core Laboratories Center (Ithaca, New York). Sequence data were edited with Sequencher version 4.8 g and analyzed using PAUP* version 4.010b 19 ; midpoint rooted, neighbor-joining (NJ) phylogenies were created for the sigB locus data and for the concatenated sequence data of all 10 loci. The sigB locus has been successfully used as a typing method for Listeria species, and in combination with other subtyping methods gives a high level of discrimination. 13 Both sigB allelic typing as well as the 10-locus MLSA scheme showed that this isolate represented a new subtype (i.e., a new multilocus sequence typing type as well as a new sigB allelic type), which was not previously found among 292 sigB sequences from L. monocytogenes isolates in the Cornell Listeria Strain collection (www.pathogentracker.net). Phylogenetic analyses, including a bootstrapped NJ tree based on the 10-gene MLSA data, confirmed that this isolate belongs to the lineage III cluster and placed it on its own distinct branch, further supporting that the isolate has a unique genotype (Fig. 1).
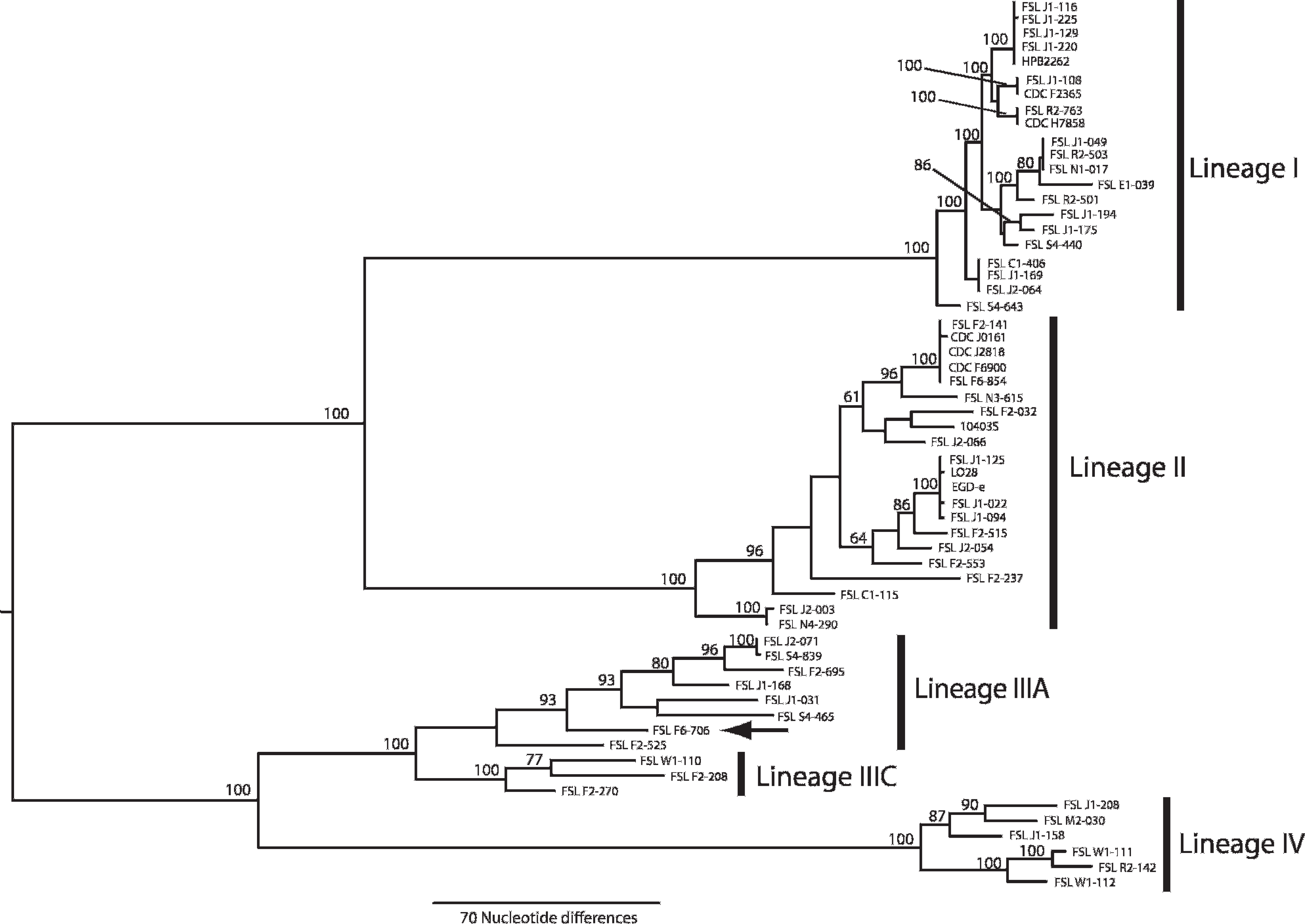
Multilocus sequence analysis phylogeny of Listeria monocytogenes. Midpoint-rooted, neighbor-joining tree based on concatenated partial sequences of sigB, ldh, prs, pbpA, rarA, polC, lmo0490, lmo1555, lmo2763, and addB sequences (6798 nucleotide positions total). Bootstrap values are based on 1,000 bootstrap replicates. The black arrow indicates the phylogenetic placement of the outbreak-associated isolate FSL F6-706 in the phylogeny of L. monocytogenes.
In summary, the current report is of an atypical outbreak of listeriosis in dairy cattle, which included unique clinical features (i.e., a high case mortality rate and a short time between the onset of clinical signs and death) and appears to have been caused by a unique L. monocytogenes serotype 4b lineage III strain. Unfortunately, an L. monocytogenes isolate for characterization was only available from 1 of the affected animals; this is not unusual, as the expense of bacteriologic diagnostics often prevents submission of specimens from multiple animals. While the epidemiologic features of this outbreak (i.e., a considerable number of cases over a short time) make it likely that this outbreak was caused by a single source and a single strain, it cannot be excluded that some of the affected animals may have been infected by another strain. The clinical features of this outbreak suggest a high virulence of the strain isolated in this outbreak, which is consistent with previous reports indicating increased virulence-associated phenotypes of some lineage III strains (i.e., relatively high cell-to-cell spread in tissue culture). 24 One cannot exclude, however, that additional factors (e.g., host and environmental factors) may have been responsible for, or contributed to, the unique clinical features of this outbreak. Future phenotypic and genetic studies (e.g., tissue culture assays, genome sequencing) of this strain, as well as other lineage III and IV strains involved in animal listeriosis cases and outbreaks, 26 will be critical to explore mechanisms that may be responsible for unique virulence characteristics hypothesized for these strains and lineages. The distribution of cases among animals of different developmental stages (i.e., heifers, dry cows, and milking cows) and physical locations suggests that the outbreak in the current study originated in the feed. Although L. monocytogenes was not isolated from the silage, the high pH of the barley silage makes this a likely source. Barley silage can be difficult to ferment correctly, particularly if a mature cutting is used, as the water content is often too low for effective fermentation. 12 It is tempting to speculate that drier silage may predispose to the development of oral lesions, thus, potentially providing entry points for L. monocytogenes in the feed.
Identification of a unique lineage III L. monocytogenes strain from a severe outbreak of listeriosis in dairy cattle also highlights the potential of animal-associated agricultural environments to represent a source of unique pathogen strains. Previous studies have reported a high prevalence of L. monocytogenes in ruminant farms, 15 which may facilitate emergence of new strains.
Acknowledgements. The current work was partially supported by U.S. Department of Agriculture (USDA) Special Research Grants (2006-34459-16952; 2008-34459-19043) and USDA-CSREES Hatch Funds (NYC-143445). Lewis Graves (U.S. Centers for Disease Control and Prevention) is acknowledged for performing serotyping of L. monocytogenes.
Footnotes
a.
Intervet/Schering-Plough Animal Health, Boxmeer, The Netherlands.
b.
Remel Inc., Lenexa, KS.
c.
API®, bioMérieux Inc., Durham, NC.
d.
Denka Seiken USA Inc., Campbell, CA.
e.
RiboPrinter® Microbial Characterization System, DuPont Qualicon, Wilmington, DE.
f.
BigDye Terminator®, Applied Biosystems, Foster City, CA.
g.
Gene Codes Corp., Ann Arbor, MI.
