Abstract
Centronuclear myopathy (CNM) is an autosomal recessive hereditary disease affecting Labrador Retriever dogs. The disease is characterized by muscle lesions, typically encompassing reduction in the number and atrophy of type II fibers, and is caused by a short interspersed repeat element insertion in exon 2 of the protein tyrosine phosphatase-like member A. The actual allele frequency is unknown; a study was undertaken to ascertain it using a convenience-sample population composed of 217 Labrador Retrievers. In addition to 3 subjects already diagnosed with CNM, used as positive controls for polymerase chain reaction, only 2 unrelated dogs were heterozygous wild-type/mutation (wild-type/mut). Thus, the frequency of the CNM allele observed in the present study was 1.8% and 0.47% when including and excluding the 3 mut/ mut homozygous cases, respectively. Based on the Hardy–Weinberg exact test (P = 1.00), the genotype frequency without the CNM-affected dogs was in agreement with the Hardy–Weinberg equilibrium. Assuming the Hardy–Weinberg equilibrium law, the expected frequency of the homozygous mutated genotype was calculated to be approximately 0.00005, which corresponds to 1 case of CNM out of 20,000 dogs. In conclusion, the present study indicates that the CNM allele is present but rare in a convenience sample of Labrador Retrievers in Italy.
Keywords
Centronuclear myopathy (CNM) was first described 3 in 1976 in Labrador Retriever dogs as a form of hereditary myopathy. The disease showed an autosomal recessive pattern of inheritance and was characterized by typical histopathological muscle lesions involving type II fibers. 1–5 Histopathological findings of muscle biopsies show a characteristic centralization of myonuclei, often located in areas devoid of myofibrils with mitochondrial aggregation; selective type II muscle fibers atrophy is observed. 6 The term centronuclear myopathy was subsequently introduced in 2002, since the typical histopathological muscle lesions were similar to those of CNM in human beings. 9 The disease has recently been characterized at the molecular level, and the mutation responsible was identified as a major insertion of a short interspersed repeat element (SINE) element in exon 2 of the protein tyrosine phosphatase-like member A (PTPLA). 7
Centronuclear myopathy affects both male and female dogs, regardless of coat color. The age of onset ranges from 6 weeks to 7 months, and the severity of the clinical signs is quite variable, progressive, and characterized by muscle weakness, exercise intolerance, and abnormalities of gait and posture with ventroflexion of the neck. As the condition progresses, generalized atrophy of the skeletal muscles develops. A definitive diagnosis is based on molecular testing. Most notably, a CNM DNA test also permits the identification of asymptomatic heterozygous carrier dogs. This information is particularly relevant for breeders, enabling them to produce healthy litters. From a clinical standpoint, knowledge of the frequency of the mutated allele is valuable for establishing the relative importance of the disease among similar syndromes. 8 Since, to the authors' knowledge, these data are not available, a study was undertaken to ascertain the frequency of the allele responsible for CNM as well as to estimate the prevalence of the disease using a convenience-sample population of Labrador Retrievers examined for unrelated complaints at the Veterinary Clinical Department of the University of Bologna (Ozzano dell'Emilia, Bologna, Italy) or at a commercial laboratory over a period of 8 years (2001–2009).
The present study analyzed DNA purified from 217 canine samples of different individual Labrador Retrievers from a convenience-sample population. The samples had been obtained from Labrador Retrievers who underwent routine blood testing either at the Veterinary Clinical Pathology Service of the University of Bologna or at a referral laboratory between January 1, 2001, and July 1, 2009. The samples consisted of 155 ethylenediamine tetraacetic acid–anticoagulated blood samples and 62 serum samples kept frozen at −20°C. The convenience sample included 3 subjects belonging to 2 different litters known to be affected with CNM. Nucleic acids were purified using a commercial extraction kit, a following the manufacturer's instructions with minimal modifications. Briefly, the thawed blood and serum samples were lysed using a lysis buffer with the addition of 1% Triton X-100 b and were subsequently extracted. a All polymerase chain reaction (PCR) reactions were carried out in 25 μl of final volume. The PCR mixture was composed of 5 μl of 5 × green GoTaq buffer, a 2 μl of MgCl2 (25 mM), 2.5 μl of deoxyribonucleotide triphosphates (250 μM each), 2 μl of forward (x2F2, 5′-GGAAAAAGGAACACACAAAGG-3′) and reverse (Ex3R1, 5′-ACCAATTAAACAGTGGAC TAT-3′) primers, 1 U GoTaq polymerase, a and 2.5 μl of DNA template brought up to 25 μl with molecular biology–grade water. c Each PCR reaction was carried out in duplicate. Positive and negative PCR controls were included in each PCR batch. Positive controls were represented by genomic DNA (gDNA) purified from dogs diagnosed with CNM; as a negative control, molecular biology–grade water was added, instead of gDNA, into the PCR mixture. Polymerase chain reaction was carried out using a thermal cycler, c as previously described. 7 The PCR products were evaluated after electrophoresis on a 1.5% agarose gel stained with ethidium bromide. According to a previous study, 7 the wild-type allele was represented by a 282–base pair (bp) band, while the mutated allele with the SINE insertion was represented by a 518-bp band. Once the allelic variants in the affected dogs were confirmed with direct sequencing, no uncertainties occurred in interpreting the PCR assay in the study (Fig. 1).
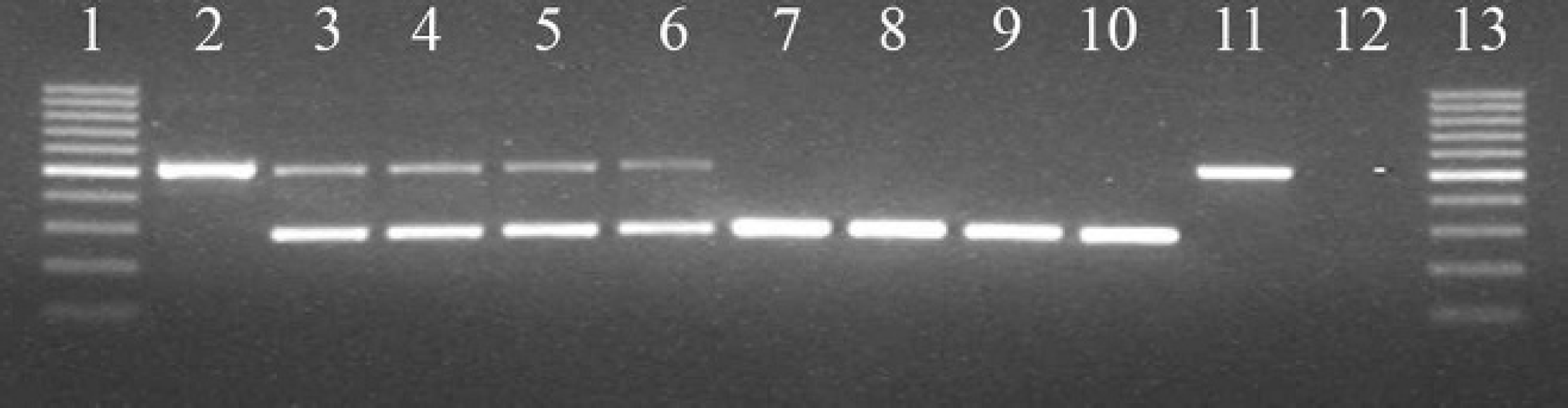
Representative picture of a 1.5% agarose gel for visualization of the polymerase chain reaction products of the centronuclear myopathy assay. Lane 1: 100–base pair (bp) ladder; lane 2: positive control; lanes 3–6: duplicates of 2 different heterozygous dogs; lanes 7–10: duplicates of 2 different wild-type homozygous dogs; lane 11: positive control; lane 12: no DNA control; lane 13: 100-bp ladder.
In addition to the 3 subjects affected by CNM, which were homozygous for mutated allele (mut/mut), only 2 additional unrelated dogs were heterozygous wild-type/mut. The frequency of the CNM allele observed in the current study was 1.8% and 0.47%, including and excluding the 3 mut/mut homozygous cases, respectively.
Based on both the chi-square test (P = 0.95) and the Hardy–Weinberg (HW) exact test d (P = 1.00), the genotype frequency without the CNM-affected dogs was in agreement with the HW equilibrium (HWE). Assuming the HWE law and using the equation p2 + 2pq + q2 = 1, the frequency of the homozygous mutated genotype was calculated to be approximately 0.00005, which corresponds to 1 case of CNM out of 20,000 dogs. The inclusion of the CNM-affected dogs, which were referred to the Veterinary Clinical Department of the University of Bologna, eventually strongly biased the allele frequency assessment; in fact, according to the HWE law, approximately 50 heterozygous dogs were expected.
Although violations of the underlying assumptions, such as the inclusion of multiple generations of dogs, are possible, the associated P value of the chi-square and HW exact tests nevertheless argues in favor of the fact that those assumptions were respected and that the predicted frequency of the homozygous-mutated genotype is quite accurate. Convenience samples usually represent a feasible way of sampling pet animals since truly random samples are impractical or even impossible. In fact, the true frequency of deleterious traits is likely overestimated by a genetic screening laboratory (as a result of the tendency to test individuals related to affected individuals or known heterozygous individuals) and underreported in a genetic registry where only “clear” animals are registered. Indeed, the convenience sample has comprised dogs that had some ancillary clinicopathological exams different from genetic testing of CNM. No evident bias in the convenience-sample population could be ascertained, although convenience samples in general have some drawbacks. Convenience samples are not truly random samples; they may not reflect the general Labrador Retriever population, and, therefore, the actual frequencies may differ from those reported herein. In conclusion, the present study indicates that the CNM allele is present but rare in an Italian convenience sample of Labrador Retrievers.
