Abstract
The aim of the current study was to evaluate the efficacy of gastric washing for detecting the presence of Helicobacter spp. in feline gastric mucus. Gastric fluids were collected from 11 cats undergoing routine surgical procedures. The fluids were centrifuged and the pellets were subjected to cytological examination by May-Grünwald/Giemsa (MGG) staining and testing by polymerase chain reaction (PCR) for the presence of Helicobacter after DNA extraction. Helicobacter spp. were detected in 8 of 11 samples by MGG staining and in 10 of 11 samples by PCR. Gastric washing proved to be a useful sampling technique and a valuable alternative to the gold-standard method of gastric biopsy, which has the risk of missing colonized spots in the case of patchy Helicobacter colonization. Gastric washing, a noninvasive way of sampling, could be applied to cats undergoing general anesthesia for different causes. Detection of Helicobacter in cats could allow epidemiological investigations, and sequencing of samples could assist in assessing the distribution of various Helicobacter species.
Keywords
Gastric bacteria categorized in the genus Helicobacter are Gram negative and spiral shaped; at least 24 species have been reported, and most Helicobacter spp. are suspected or proved gastric or hepatic pathogens in humans. 7 Helicobacter pylori, colonizing the stomach of half of the world's human population, is a proven primary cause of gastritis and peptic ulceration 4 and is considered a major risk factor in the development of mucosa-associated lymphoid tissue lymphoma 22 and gastric adenocarcinoma. 19
Studies of the relationship between gastric disease and Helicobacter spp. infection in animals have resulted in the discoveries of Helicobacter mustelae in ferrets with gastritis and peptic ulcers, Helicobacter acinonychis in cheetahs with severe gastritis, and Helicobacter heilmannii in pigs with gastric ulcers. 5,8,21 The role of Helicobacter spp. in gastrointestinal disease in dogs and cats is uncertain. It is known that virtually all healthy adult cats harbor spiral helicobacter in their gastric mucosa, but the clinical significance and ecology of this microorganism within the stomach of domestic carnivores remain largely unknown. 12 The marked morphological resemblance and the genetic relatedness between the spiral organisms identified in both animal and human gastric tissues led to the hypothesis that spiral gastric bacteria are zoonotic agents. The high prevalence of these gastric Helicobacter spp. in animals, 2,12 in contrast with their low prevalence in humans, 1,11 strongly indicates that animals may act as natural reservoirs for transmission to humans. 3 The frequent detection of gastric Helicobacter—like organisms (GHLOs) in the cat led to the hypothesis that feline Helicobacter spp. may be transmitted to humans. 10,11,17
Diagnostic tests for Helicobacter spp. can be noninvasive, as is the urea breath test, or invasive, as is biopsy. Breath samples can be obtained either through the endotracheal tube in anesthetized cats or, in conscious, cooperating animals, through a face mask. 15 Gastroscopy requires general anesthesia and should always be followed by biopsy because the mucosa may appear macroscopically normal but may nevertheless be inflamed, 24 and a tissue sample is essential for histopathological examinations and strain identification. However, in cases of patchy Helicobacter colonization, biopsy sampling has the risk of missing colonized spots. Some authors reported that the prevalence of Helicobacter spp. detected in gastric biopsies using polymerase chain reaction (PCR) technique was low, because the results could be affected by factors such as sampling location, specimen preparation, and bacterial density. 20 The aim of the current study was to evaluate a novel noninvasive sampling technique, gastric washing followed by May-Grünwald/Giemsa (MGG) staining or PCR, to detect the presence of Helicobacter spp. in feline gastric mucus.
Eleven privately owned cats (4 males and 7 females), ranging between 2 and 16 years of age, undergoing routine surgical procedures were included in the study. Owner consent was obtained for all cats. After a 12-hr fasting, all cats were intubated and anesthetized with isoflurane in oxygen. Before the scheduled surgery, the animals were placed in left lateral recumbency, and gastric washing was performed using a gastric tube (4.0 mm in diameter). Ten milliliters of sterile saline solution was slowly injected into the stomach and almost immediately retrieved through a syringe. The fluid was centrifuged (5 min, 3,000 x g), the supernatant removed, and the pellet resuspended in 500 μl of phosphate buffered saline. The resuspension was split into 2 pieces, and one part was subjected to cytocentrifugation (100 μl, 5 min, 22.4 x g in a Shandon Cytospin 2 centrifuge) for cytological examination following MGG staining. 6 The other part was used for DNA extraction for PCR. DNA extraction was carried out by a commercial tissue extraction kit. a The extracted DNA was subjected to PCR using specific primers that amplify a fragment of the 16S ribosomal RNA (rRNA) gene of Helicobacter spp.
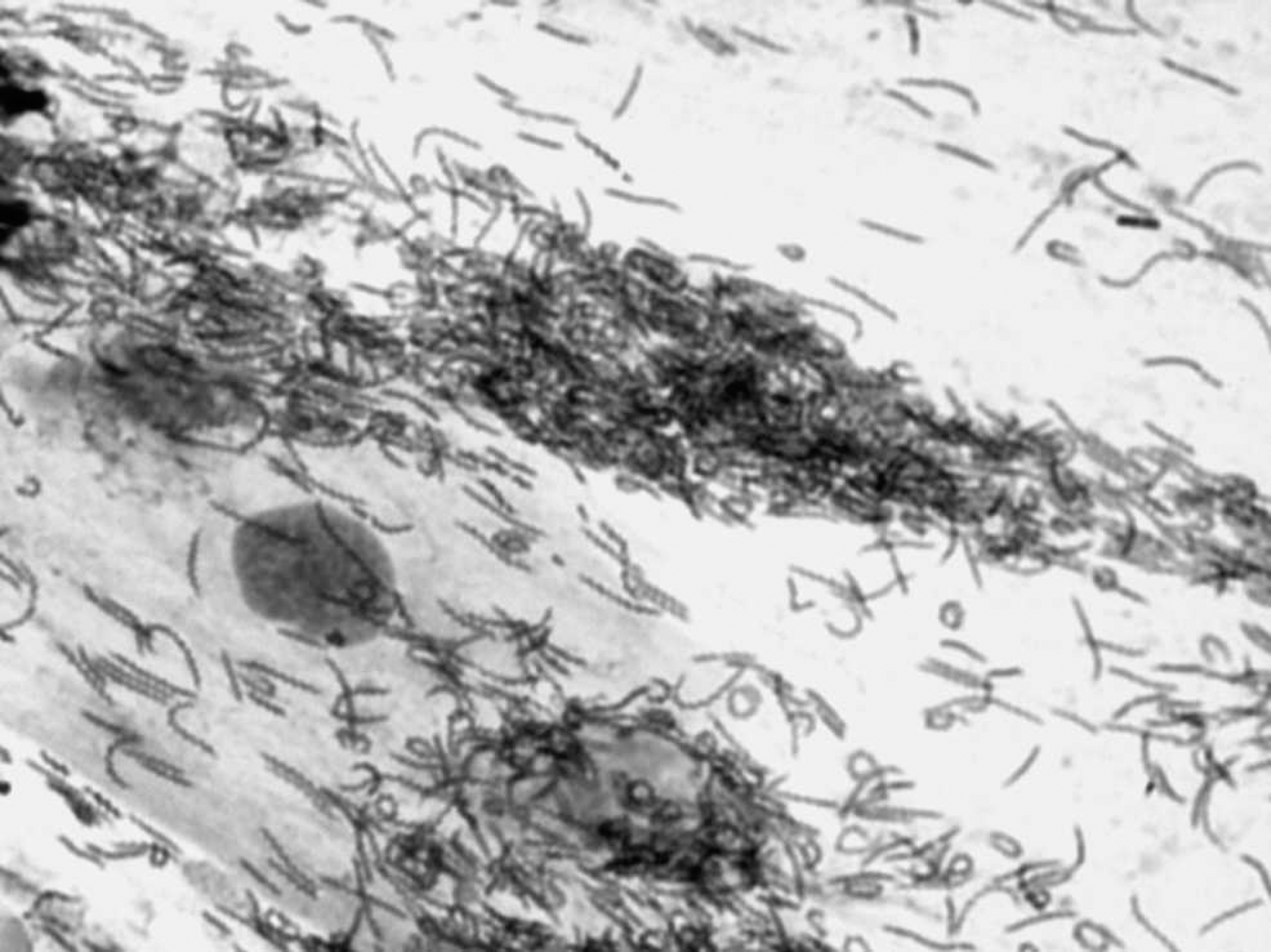
Massive presence of Helicobacter in feline gastric washing. May-Grünwald/Giemsa stain. 1,000X.
Polymerase chain reaction was performed using 0.5 U of Taq DNA polymerase, 10 mmol of Tris HCl, 50 mmol of KCl, 1.5 mmol of MgCl2, 200 μmol deoxyribonucleotide triphosphate, and 25 μmol of each primer (sense primer: innS [GAACCTCTAGGCTTGACATTG] and antisense primer: innR [GGTGAGTACAAGACCCGGGAA]). The samples were subjected to 30 amplifying cycles at the following temperatures: denaturation at 94°C for 30 sec, annealing at 47°C for 30 sec, and extension at 72°C for 30 sec. Products from the PCR were evaluated by means of 7% polyacrylamide gel electrophoresis, comparing the obtained amplicons to a DNA molecular weight marker. b Five of the 11 amplified products from the samples were further purified from agarose gel by means of a commercial extraction kit c and were sequenced. d
Helicobacter was detected in 8 of 11 samples by MGG staining and in 10 of 11 samples by PCR. May-Grünwald/Giemsa—stained cytological samples showed dark-purple spiral microorganisms, presumptive of Helicobacter spp. (Fig. 1). Polymerase chain reaction—positive samples presented a typical 420—base pair fragment of the 16S rRNA gene. Nucleotide sequencing reflected sequences with a homology range of 99–100%, with a partial sequence from 16S rRNA gene of H. heilmannii and an uncultured Helicobacter sp. from a cheetah. 16,23
Gastric washing proved to be a useful noninvasive technique to detect the presence of Helicobacter spp. in cat stomachs. In the current study, gastric washing showed a prevalence of 90.9% of Helicobacter spp., which is in accordance with results of previous investigations, reporting values ranging from 57% to 100% in gastric biopsies. 9 A previous study 15 showed that the urea breath test can be a reliable noninvasive diagnostic method as well, although it is not useful for species identification. At present, the gold standard of diagnostic tests in cats is considered to be gastric biopsy followed by morphological observations and PCR for species differentiation. 15 Results from the present study indicate that MGG staining represents a useful and uncomplicated way to detect the presence of the microorganism, 15 although PCR may be a more sensitive method of detection when few bacteria are present.
Results from the current study also indicate that cats are frequently colonized by H. heilmannii without showing any gastrointestinal symptoms. 15,18 Previous studies that differentiated Helicobacter spp. (e.g., PCR or electron microscopy) showed that H. heilmannii-like species were the most commonly found GHLO in the stomach of the cat, Helicobacter felis could be isolated less frequently, and the presence of H. pylori remains anecdotal. 15 Also, infection with more than 1 species may occur. 13,14 The role of the cat as a reservoir of zoonotic agents should be investigated further, not only because a small proportion (approximately 0.25–1.7%) of human patients suffering from gastritis, peptic ulcers, and gastric cancer are diagnosed with gastric Helicobacter-like bacteria other than H. pylori 11 but because a previous study 10 that detected H. pylori in cats indicated that the cat might represent a reservoir for this bacterium, whose natural host is human.
The great majority of GHLOs of carnivorous pets are confined to the mucus layer (Lecoindre P, Chevallier M, Berger I, Peyrol S: 1997, Contribution to the study of gastric helicobacters. Ultrastructural aspects of the cat gastric mucosa. In: European Society of Veterinary Internal Medicine Congress Proceedings. Ecole Nationale Veterinaire de Lyon, Lyon, France, p. 166), and gastric washing could, therefore, be a valuable technique to sample the entire gastric mucosa, avoiding the need to select the biopsy area and the risk of missing colonized spots. Gastric washing, an uncomplicated and noninvasive method of sampling, could be applied to cats submitted to general anesthesia for different causes, thus allowing epidemiological investigations on transmission of these bacteria. Furthermore, sequencing the samples would allow the assessment of species distribution in the stomach and the evaluation of whether H. pylori and H. heilmannii are potential zoonotic agents. The relationship between DNA sequence of Helicobacter spp. found in cats and in their owners could determine if these bacteria are zoonotic agents that may have important public health implications.
The proposed technique could also be of clinical relevance, since it might increase the rate of Helicobacter detection during patchy mucosal colonization. The method could also be used in association with endoscopy and biopsy for symptomatic cats to assess the distribution of inflammation and the location of the bacteria in tissues. Because the role of Helicobacter in cat gastritis has not been fully elucidated, further investigation of feline gastric pathology is needed to understand strain-dependent virulence.
Acknowledgements. The authors wish to thank Drs. Federico Stoppani and Elio Bossi, small animal practitioners in Ospedaletti (IM) and Sanremo (IM), Italy, for sample collection.
Footnotes
a.
NucleoSpin® Tissue Extraction Kit, Macherey-Nagel, Düren, Germany.
b.
DNA Molecular Weight Marker V, Roche Diagnostics, Basel, Switzerland.
c.
NucleoSpin® Extract Kit, Macherey-Nagel, Düren, Germany.
d.
BMR Genomics Srl, Padova, Italy.
