Abstract
Pasteurellaceae are bacteria with an important role as primary or opportunistic, mainly respiratory, pathogens in domestic and wild animals. Some species of Pasteurellaceae cause severe diseases with high economic losses in commercial animal husbandry and are of great diagnostic concern. Because of new data on the phylogeny of Pasteurellaceae, their taxonomy has recently been revised profoundly, thus requiring an improved phenotypic differentiation procedure to identify the individual species of this family. A new and simplified procedure to identify species of Actinobacillus, Avibacterium, Gallibacterium, Haemophilus, Mannheimia, Nicoletella, and Pasteurella, which are most commonly isolated from clinical samples of diseased animals in veterinary diagnostic laboratories, is presented in the current study. The identification procedure was evaluated with 40 type and reference strains and with 267 strains from routine diagnostic analysis of various animal species, including 28 different bacterial species. Type, reference, and field strains were analyzed by 16S ribosomal RNA (rrs) and rpoB gene sequencing for unambiguous species determination as a basis to evaluate the phenotypic differentiation schema. Primary phenotypic differentiation is based on β-nicotinamide adenine dinucleotide (β-NAD) dependence and hemolysis, which are readily determined on the isolation medium. The procedure divides the 28 species into 4 groups for which particular biochemical reactions were chosen to identify the bacterial species. The phenotypic identification procedure allowed researchers to determine the species of 240 out of 267 field strains. The procedure is an easy and cost-effective system for the rapid identification of species of the Pasteurellaceae family isolated from clinical specimens of animals.
Introduction
Bacterial species belonging to the family Pasteurellaceae are Gram negative, aerobic, coccoid- or rod-shaped, non-spore-forming, and nonmotile. In vitro, these species are routinely grown under aerobic conditions supplemented with 5% carbon dioxide (CO2) on growth medium containing the growth factors hemin and β-nicotinamide adenine dinucleotide (β-NAD), the latter being required for certain species in this family. Many of the species of the Pasteurellaceae family inhabit the mucosal membranes of the alimentary, respiratory, and genital tract of mammals, birds, and reptiles. Several Pasteurellaceae are known as opportunistic secondary invaders, able to cause infections under predisposing circumstances, whereas some species are primary pathogens that cause severe diseases in certain animals. 20 Pasteurellaceae show ecological preferences for specific surfaces and hosts. However, very little is known about the factors that govern this host specificity. 3 The taxonomic classification of these fastidious bacteria has recently been revised as a consequence of increasing phylogenetic data, which have redefined the relationships among the various species of the family based on the 16S rRNA gene (rrs), rpoB, and infB sequence analyses. 1,2,4,7,12,18,19 To date, the following 13 different genera have been described: Actinobacillus, Aggregatibacter, Avibacterium, Bibersteinia, Gallibacterium, Haemophilus, Histophilus, Lonepinella, Mannheimia, Nicoletella, Pasteurella, Phocoenobacter, and Volucribacter.
The purpose of the current study was to establish an easy and accurate procedure to identify the most commonly encountered species of the Pasteurellaceae family during routine bacteriological diagnostics of animal specimens. The procedure is based on the most recent taxonomy from the International Committee on Systematics of Prokaryotes (ICSP) Subcommittee on the Taxonomy of Pasteurellaceae, the List of Prokaryotic Names With Standing in Nomenclature (LPSN), the description of the biochemical characteristics of the strains from the Dictionnaire de Bactériologie Vétérinaire, 9 Bergey's Manual of Systematic Bacteriology 20 and the revised definition of Actinobacillus sensu stricto. 6 The spectrum of species contains 23 validated bacterial species, 6 subspecies, 4 biovars, and 1 candidate species (‘Actinobacillus porcitonsillarum’), all of which are currently encountered in routine veterinary bacteriological examinations. Identification is mainly based on β-NAD dependence, hemolysis pattern, tryptophanase activity (indole test), and the animal origin of the strains, followed by further phenotypic characterizations.
Materials and methods
Bacterial strains
A total of 307 strains of the Pasteurellaceae family were investigated in the current study from the following sources (Table 1): 40 type and reference strains originating from bacterial culture collections (for Gallibacterium anatis, biovar haemolytica, an internal reference strain was used, which was originally isolated from a chicken and confirmed by rrs and rpoB sequence analysis and multilocus sequence analysis [MLSA] 12 , 16 ), 267 field strains isolated from necropsy or clinical material of various animals, and 9 bacterial species that do not belong to the Pasteurellaceae family but that share phenotypic properties with it, including Acinetobacter baumannii, Acinetobacter radio-resistens, Aeromonas hydrophila, Bordetella avium, Bordetella bronchiseptica, Escherichia coli, Pseudomonas aeruginosa, and Yersinia pseudotuberculosis. These 9 species were used as controls. The field strains for the current study were differentiated phenotypically and genotypically.
Genetic typing and identification
Type, reference, and field strains were analyzed by rrs and rpoB sequence analysis for identification as described. 12 Threshold values for species designation were set at a conservative criterion of 99% and 98% for rrs and rpoB, respectively. Sequence comparisons were done using the SmartGene IDNS software package. a , 12 , 21 Actinobacillus pleuropneumoniae biovars I and II were identified by the Apx toxin gene (apx) detection. 10 For the identification of A. pleuropneumoniae serotype 2 biovars I and II, a cps2-based polymerase chain reaction (PCR) was used. 11 Actinobacillus porcitonsillarum was identified by characterization of the RTX toxin ApxII genes apxIICABD and its particular flanking genes in this species. 17
Culture conditions and phenotypic and biochemical identification
The isolates were cultivated on 5% sheep blood trypticase soy agar (TSA) b with a β-NAD-producing “nurse” strain (Staphylococcus sciuri strain JF700, strain collection of the Institute of Veterinary Bacteriology, Bern, Switzerland). The isolates were additionally cultivated on chocolate agar medium supplemented with PolyViteX. c The plates were incubated for 24 hr at 37°C in an atmosphere of 5% CO2. Afterward, cultures were evaluated for hemolysis. All strains were tested for tryptophanase activity, which leads to indole production from tryptophan, using a dimethyl cinnamaldehyde (DMCA) indole test, d for catalase activity (measuring gas production from 3% hydrogen peroxide [H2O2]) using a catalase test, d and for oxidase activity using oxidase reagent. c For strains of Actinobacillus equuli subsp. haemolyticus, use of a cover slip was necessary to assess catalase reaction to trap the oxygen bubbles. Fermentation of carbohydrates was assessed by culture medium supplemented with mannitol, mannitol plus β-NAD, saccharose, trehalose, sorbitol, glucose, and dulcitol. The following test kits were used according to the manufacturers' instructions: urease test, b ornithine decarboxylase test, b and motility test. d Inoculation of test tubes was carried out with a sterile loop with 10–20 colonies incubated for 24 hr at 37°C under aerobic conditions. In certain cases, a further incubation time of 24 hr was necessary to obtain clear, interpretable results.
Results
Identification procedure
The identification procedures presented in Figure 1 and corresponding Tables 2–5 was elaborated on the basic phenotypic characteristics of Pasteurellaceae as described previously, 9 , 20 taking into account the amended taxonomy. As an additional decisive criterion for identification, the animal species from which the isolate originated (see Tables 2–5) was used because that is a valid parameter for routine diagnosis. 6 Growth dependence on β-NAD was used as the first factor for differentiation (Fig. 1). Subsequently, hemolysis, indole, oxidase, and catalase activity determinations allowed the separation of different species and subspecies into 6 different groups (Fig. 1). Based on these criteria, the corresponding biochemical tests were then chosen for further identification. Strains that showed β-NAD dependence were analyzed for mannitol catabolism and urease activity (Table 2). If the strain was β-NAD independent and showed β-hemolysis, 9 supplementary phenotypic tests were performed, including motility, urease activity, ornithine decarboxylase, and catabolism of saccharose, trehalose, and sorbitol (Table 3). The third group was β-NAD-independent, nonhemolytic, and indole-negative strains, which were differentiated by oxidase and catalase activity and by glucose, saccharose, trehalose, and mannitol catabolism; urease activity; nonmotility; and gas production on a medium containing glucose as a carbon source (Table 4). The fourth group included the β-NAD-independent, nonhemolytic, and indole-positive strains, which were further identified by oxidase and catalase activity and nonmotility (as decisive criteria for Pasteurellaceae); mannitol, sorbitol, dulcitol, and trehalose catabolism; and urease and ornithine decarboxylase activity (Table 5). Basically, all field strains tested displayed the same phenotypes as their respective type or reference strains unless stated specifically.
Bacterial strains used in this study.
Continued.
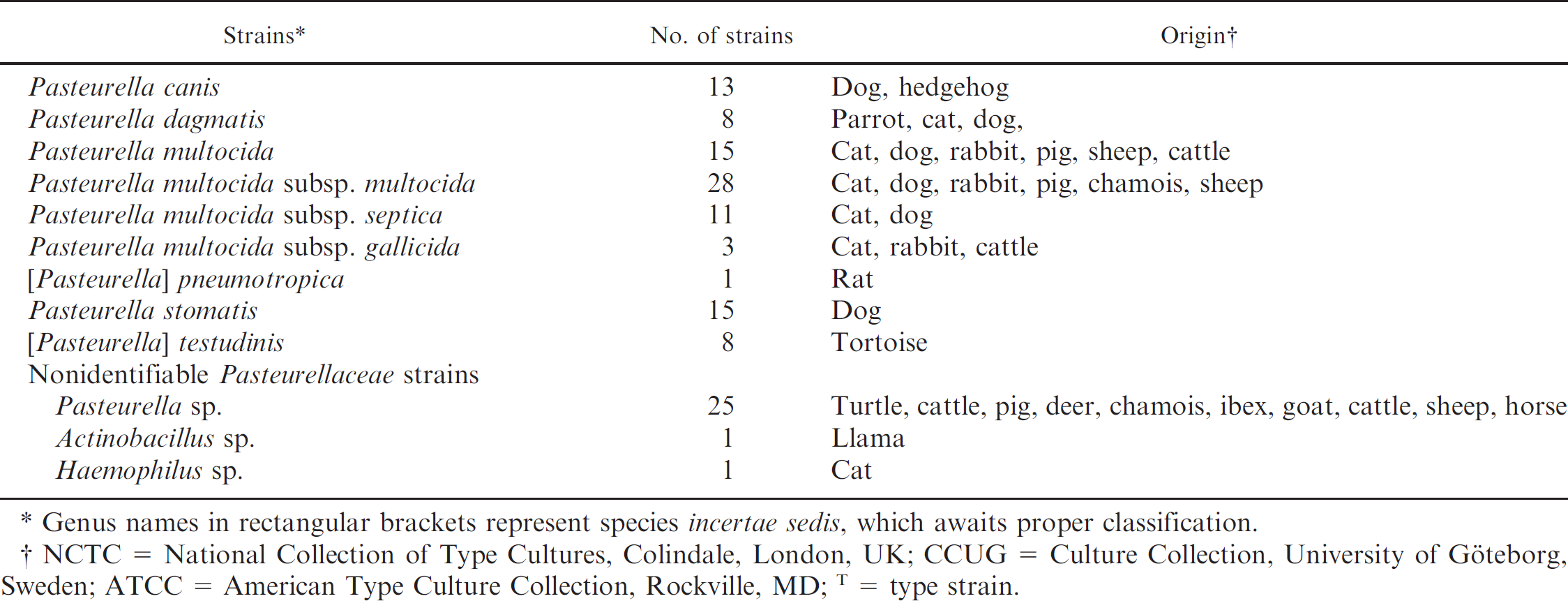
Genus names in rectangular brackets represent species incertae sedis, which awaits proper classification.
NCTC = National Collection of Type Cultures, Colindale, London, UK; CCUG = Culture Collection, University of Göteborg, Sweden; ATCC = American Type Culture Collection, Rockville, MD;T = type strain.
Identification of β-NAD-dependent strains
In the group of β-NAD-dependent Pasteurellaceae, 9 species could be differentiated using the phenotypes given in Table 2. The 14 field strains of Actinobacillus pleuropneumoniae biovar I and the 8 field strains of ‘Actinobacillus porcitonsillarum’ showed concordant results with the corresponding type and reference strains tested. The strains had been previously identified by analysis of the apx gene patterns. [Actinobacillus] porcinus, Avibacterium avium, Avibacterium volantium, and Avibacterium paragallinarum were tested using the corresponding type strain only because no field strains were available. Between the 2 [Haemophilus] felis field strains analyzed, one showed β-hemolysis after 72 hr and the other isolate after 48 hr. Neither isolates grew in the mannitol medium supplemented with β-NAD. The isolates were confirmed as [Haemophilus] felis by rrs and rpoB sequence analysis. In total, 25 [Haemophilus] parasuis field strains have been examined and isolated during the past 40 years. All 25 strains, plus the type strain representatives CCUG3712T and NCTC7440T, showed clear positive oxidase reaction. This is in contrast to the identification characteristics given in Bergey's Manual of Systematic Bacteriology. 20 However, all [Haemophilus] parasuis strains that were confirmed by rrs and rpoB sequence analysis were clearly oxidase positive.
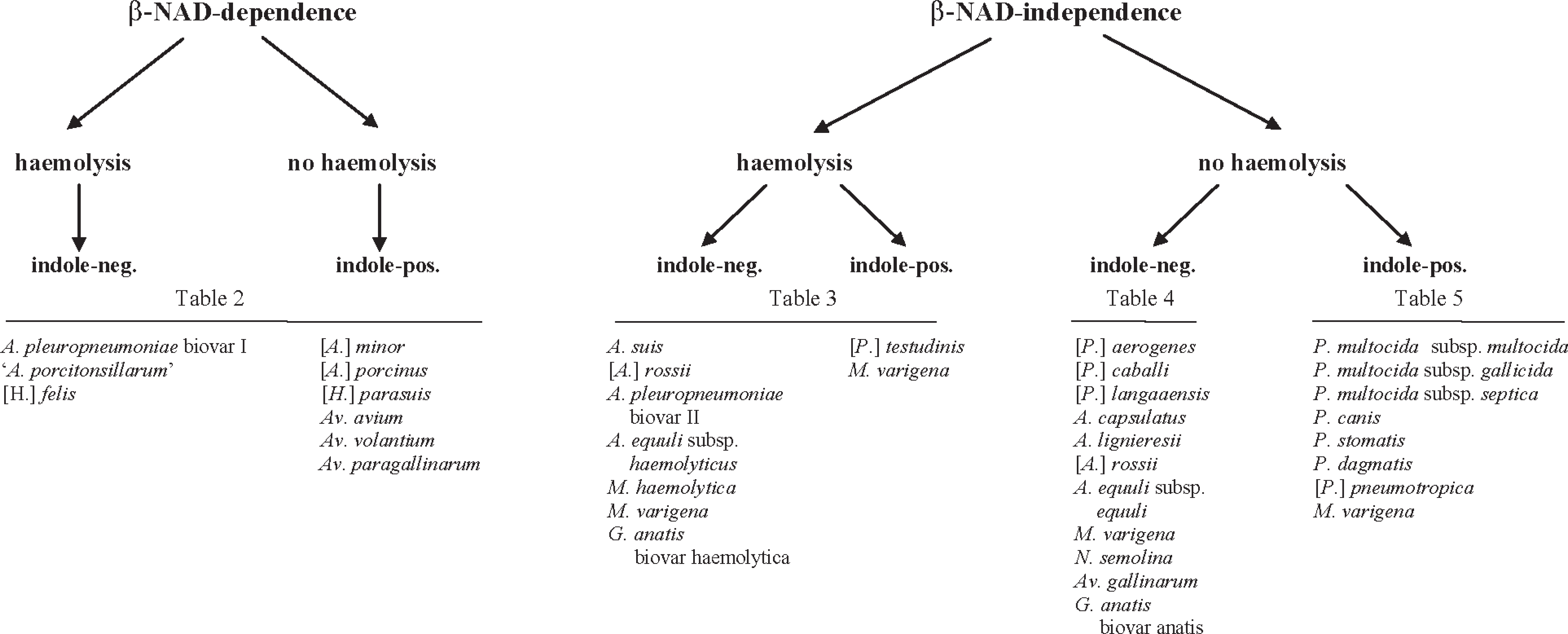
Flow diagram of the phenotypic identification process for Actinobacillus sp., Avibacterium sp., Gallibacterium sp., Haemophilus sp., Mannheimia sp., Nicoletella sp., and Pasteurella sp. that are typically isolated in veterinary bacterial diagnostic laboratories. Table numbers refer to tables in this article.
Identification of β-nicotinamide adenine dinucleotide (β-NAD)-dependent strains in the current study.*
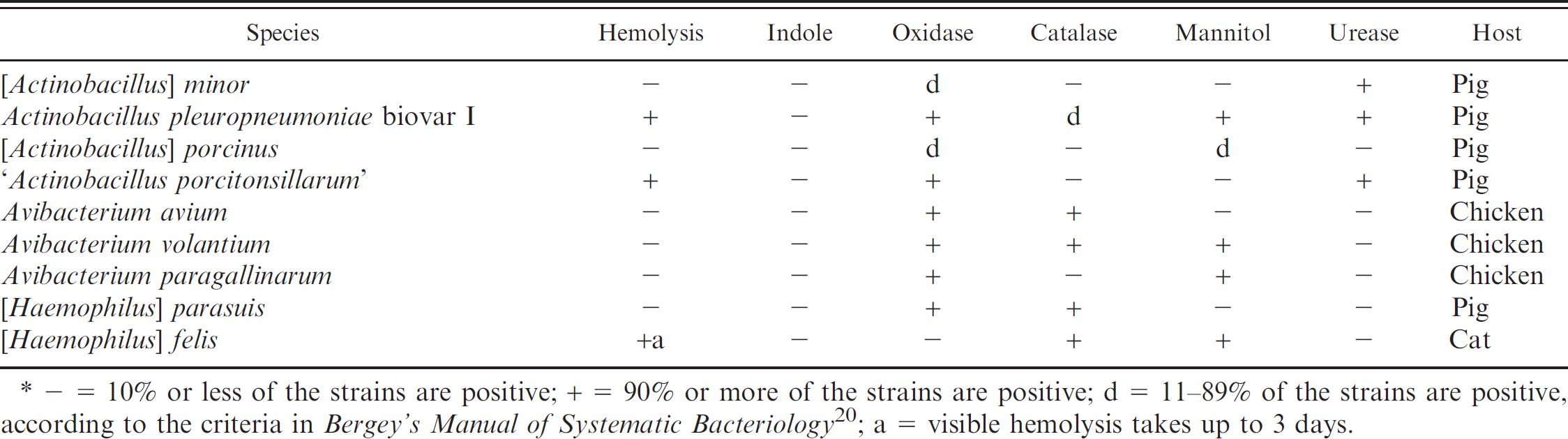

− = 10% or less of the strains are positive; + = 90% or more of the strains are positive; d = 11–89% of the strains are positive, according to the criteria in Bergey's Manual of Systematic Bacteriology 20 ; a = visible hemolysis takes up to 3 days.
Identification of β-NAD-independent hemolytic strains
Among the β-NAD-independent, hemolytic Pasteurellaceae, 8 different species, subspecies, or particular biovars could be distinguished using standard criteria for phenotypic characterization 5 (Table 3). The reference strain of Actinobacillus equuli subsp. haemolyticus and 10 field strains isolated from horses resulted in the phenotypic parameters expected for this subspecies. 8 The A. pleuropneumoniae biovar II serotype 2 reference strain N273, and 9 biovar II field strains of various serotypes isolated from pigs with pleuropneumonia showed the characteristic indole-negative, urease-positive, saccharose-positive, and trehalose- and sorbitol-negative phenotypes and could all be clearly identified. The hemolytic reference strains of [Actinobacillus] rossii 22 the type strain, and 3 field strains of Actinobacillus suis that were previously analyzed for the presence of apxI and apxII toxin genes, 23 revealed the characteristic phenotypes that allow for their identification (Table 3). For Gallibacterium anatis, biovar haemolytica, the internal reference strain analyzed thoroughly by rrs and rpoB sequence analysis and MLSA, 12 , 16 was used in the current study. It should be noted that field strains of this species show variable phenotypes regarding trehalose and sorbitol catabolism as noted in Table 3. 7 However, G. anatis shows a host specificity that is restricted to poultry, which facilitates routine identification.
Mannheimia varigena is not frequently encountered in clinical samples. The few strains from the authors' collection were scattered in the various groups and showed variable phenotypes regarding hemolysis and indole production (Tables 3–5). Its unambiguous identification would require a genotypic analysis. However, the phenotypes used differentiated M. varigena from the other species in this group (Table 3). All Mannheimia haemolytica strains used in the current study were grouped among β-NAD-independent, hemolytic, indole-negative strains and could be identified unambiguously using the phenotypic tests in Table 3. [Pasteurella] testudinis type strain CCUG19802T and 8 [Pasteurella] testudinis field isolates from tortoise could be identified at the species level, whereas 1 strain isolated from turtle, an unknown Pasteurella sp., could not be identified further. In addition, an unknown Actinobacillus sp. isolated from llama could not be typed to the species level. The non-Pasteurellaceae species Aeromonas hydrophila, hemolytic E. coli, and Pseudomonas aeruginosa resulted in phenotypes as expected from standard identification characteristics 5 (Table 3).
Identification of β-nicotinamide adenine dinucleotide (β-NAD)-independent, hemolytic strains in the current study.*
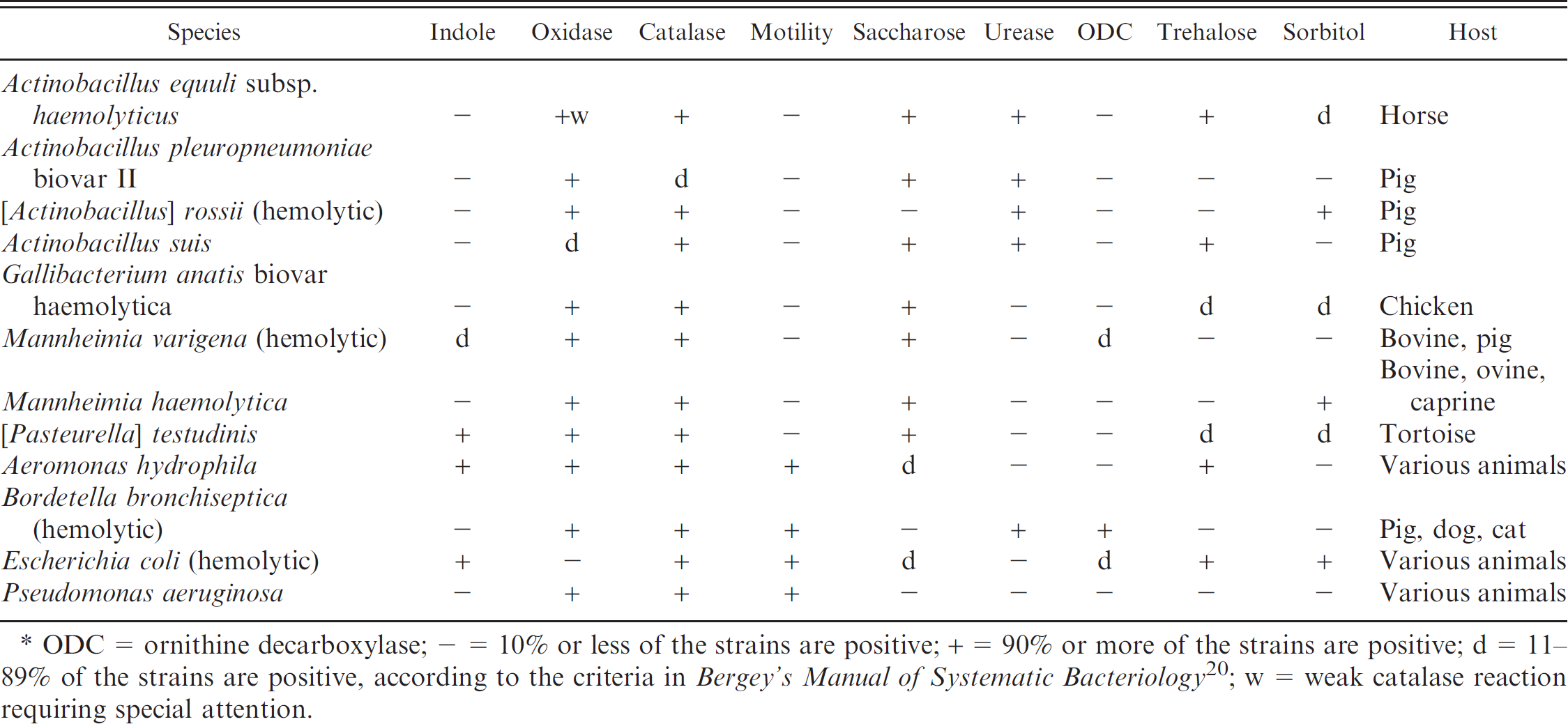
ODC = ornithine decarboxylase; - = 10% or less of the strains are positive; + = 90% or more of the strains are positive; d = 11–89% of the strains are positive, according to the criteria in Bergey's Manual of Systematic Bacteriology 20 ; w = weak catalase reaction requiring special attention.
Identification of β-nicotinamide adenine dinucleotide (β-NAD)-independent, nonhemolytic, indole-negative strains in the current study.*
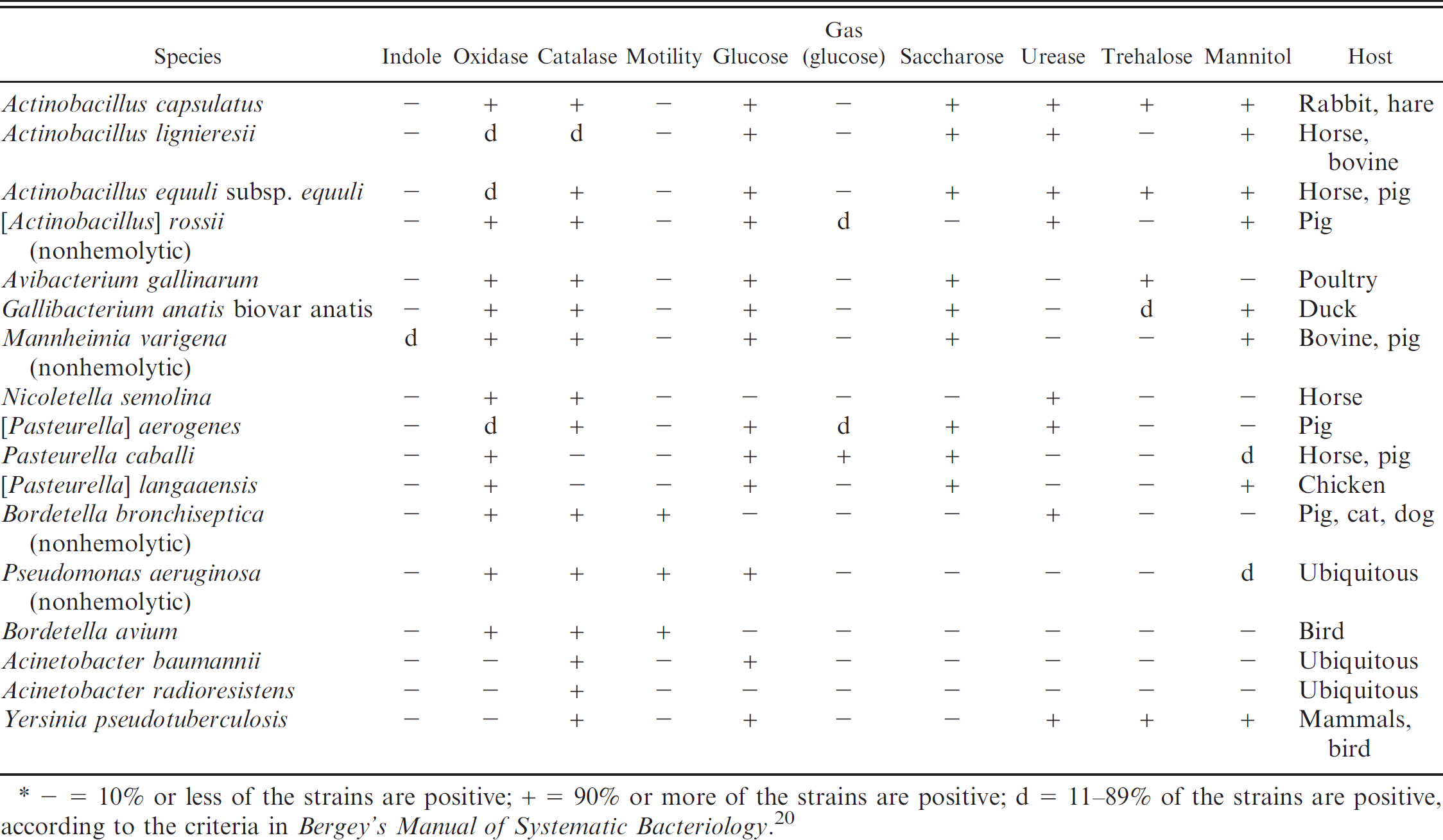
− = 10% or less of the strains are positive; + = 90% or more of the strains are positive; d = 11−89% of the strains are positive, according to the criteria in Bergey's Manual of Systematic Bacteriology. 20
Identification of β-nicotinamide adenine dinucleotide (β-NAD)-independent, nonhemolytic, indole-positive strains in the current study.*
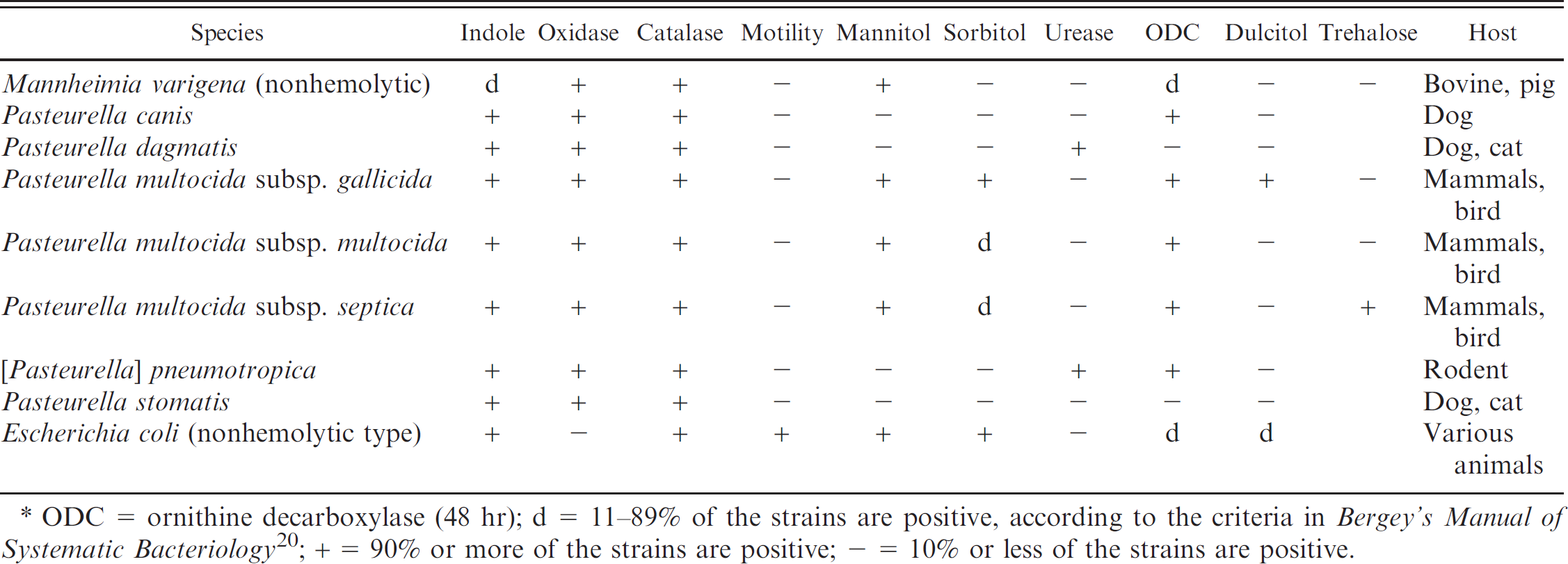

ODC = ornithine decarboxylase (48 hr)
d = 11−89% of the strains are positive, according to the criteria in Bergey's Manual of Systematic Bacteriology 20
+ = 90% or more of the strains are positive
− = 10% or less of the strains are positive.
Identification of β-NAD-independent, nonhemolytic, indole-negative strains
Among the β-NAD-independent, nonhemolytic, indole-negative strains analyzed, 9 species, 1 subspecies, and 1 particular biovar could be differentiated using the criteria of Table 4, and resulted in the criteria given in Bergey's Manual of Systematic Bacteriology. 5 For the species Actinobacillus capsulatus, Avibacterium gallinarum, Gallibacterium anatis biovar anatis, and [Pasteurella] langaaensis, only type strains were tested because these species' respective biovars are rarely encountered in routine veterinary diagnostic laboratory. The species gave the expected phenotypic pattern as listed in Table 4. It should be noted, however, that A. capsulatus and A. equuli subsp. equuli reveal the same phenotypes and differentiation between the 2 relays on the animal host species, unless genetic analysis are undertaken for their distinction. Two nonhemolytic [Actinobacillus] rossii strains could readily be identified. The type strain of Nicoletella semolina, and 3 N. semolina field strains from horses with respiratory distress, resulted in the phenotypes as published previously 15 (Table 4). The [Pasteurella] aerogenes type strain ATCC27883T, and 3 strains isolated from aborted piglets, all characterized as carrying the RTX toxin gene operon paxABCD 14 which resulted in the expected phenotype profile (Table 4). Furthermore, the procedure shown in Table 4 allowed the differentiation of the [Pasteurella] caballi type strain, 4 field strains isolated from horses and a pig, the species Actinobacillus lignieresii (type and 1 field strain), as well as nonhemolytic A. equuli subsp. equuli, which lacks known RTX toxin genes and was isolated from horses and pigs (type and 3 field stains). Two nonhemolytic M. varigena strains from cattle and chamois (Rupicapra r. rupicapra) that were identified by rrs and rpoB sequence analysis could be clearly differentiated from the other species in this group and confirmed that nonhemolytic M. varigena are isolated from ruminants. 20 Eight β-NAD-independent, nonhemolytic, indole-negative strains could not be classified by the phenotypic tests using the identification procedure outlined in the current study. These strains all revealed to be Pasteurella spp. by rrs and rpoB sequencing: 2 were identified as Bibersteinia trehalosi, 3 as Pasteurella multocida subsp. septica, 1 as Pasteurella multocida subsp. multocida, and 2 were unidentifiable Pasteurella species. Interestingly, the 3 P. multocida subsp. septica and the P. multocida subsp. multocida (all isolated from cattle) showed an uncommon indole-negative phenotype. The non-Pasteurellaceae species, Bordetella bronchiseptica, Pseudomonas aeruginosa, Bordetella avium, Acinetobacter baumannii, Acinetobacter radio-resistens, and Yersinia pseudotuberculosis resulted in the expected phenotypes (Table 4).
Identification of β-NAD-independent, nonhemolytic, indole-positive strains
For the differentiation of β-NAD-independent, nonhemolytic, indole-positive species of Pasteurellaceae, dulcitol fermentation was used as an additional differentiation attribute (Table 5). Moreover, the ornithine decarboxylase reaction was evaluated after 48 hr instead of the standard 24 hr for this group of Pasteurellaceae. The important species in this group is P. multocida represented by its 3 subspecies gallicida, multocida, and septica. The respective type and reference strains of the 3 subspecies all resulted in the expected phenotype (Table 5). Among 48 field strains that were previously identified by phenotypic and genotypic methods, 13 3 strains of Pasteurella multocida subsp. gallicida could be readily differentiated to the subspecies level. Furthermore, 28 strains of P. multocida subsp. multocida, 11 strains of P. multocida subsp. septica, and 15 strains of P. multocida that were not subtyped to the subspecies level and that were isolated from various animals (Table 1), could be identified as Pasteurella multocida subsp. multocida-septica without differentiation between the 2 subspecies multocida and septica. This is mainly due to the dual appearance of P. multocida subsp. septica, which can show both sorbitol-positive and sorbitol-negative phenotypes. An indole-positive M. varigena that is rarely isolated from cattle was differentiated accurately from the other species of this group (Table 5). The type strain and 13 field isolates of Pasteurella canis, all isolated from dogs except 1 that was isolated from a European hedgehog, were clearly identified with the phenotypes given in Table 5. The identification procedure allowed researchers to identify the type strain and 15 isolates from dogs as Pasteurella stomatis, although 7 isolates were wrongly identified previously as P. canis by others. The strains were retyped by genetic means and by the identification characteristics in the current study as P. stomatis. Furthermore, the procedure allowed for the unambiguous identification of the type strain and a clinical isolate from a rat of [Pasteurella] pneumotropica, as well as the type strain and 8 strains of Pasteurella dagmatis from parrots, cats, and a dog. In addition, nonhemolytic E. coli, which is often isolated from clinical samples, was clearly distinguished from the Pasteurellaceae species in this group.
Discussion
Because of the new taxonomic classification of many species of the Pasteurellaceae family, a new, rational, and accurate identification procedure for the veterinary diagnostic routine laboratory needed to be established to identify the most relevant and most often encountered species. In this respect, the economic aspect of veterinary routine diagnosis had to be considered. Genetic methods using DNA sequence analysis are currently used for species identification of Pasteurellaceae for research activities and reference purposes, and they allow for very precise species and subspecies identification, in most cases. However, these methods are usually considered too expensive for the routine diagnostic laboratory because they require specialized personnel and equipment. The phenotypic characteristics exploited in the current study for the differentiation of species of the Pasteurellaceae family are readily tested in routine laboratories, and many of the tests are also used as standard tests for the differentiation of species from other bacterial families, such as Enterobacteriaceae. In general, the phenotypic reactions analyzed, as presented in Tables 2–5, correspond to the phenotypic characteristics of Pasteurellaceae as published in the most recent edition of Bergey's Manual of Systematic Bacteriology. 20 However, all of the strains of [Haemophilus] parasuis analyzed in the current study, including type strain representatives CCUG3712T and NCTC4557T, showed a clear, positive reaction using oxidase test kits from 2 different suppliers. b,c Therefore, an amendment to this parameter in current identification grids stating [Haemophilus] parasuis as an oxidase-positive species of the Pasteurellaceae family is recommended. Taking into account this amendment, the method presented in the current study was able to identify all 40 type or reference strains tested (Table 1) and to differentiate 240 out of 267 (90%) strains of the Pasteurellaceae family that were isolated from various animals during routine diagnoses in a national veterinary diagnostic and reference center, when using rrs and rpoB sequence analysis as a reference. 12 , 16 Because of the high costs of differentiating P. multocida subsp. multocida from P. multocida subsp. septica, the authors suggest typing them as P. multocida subsp. multocida-septica for routine purposes because further differentiation is only of epidemiological, not medical, importance. The strains that could not be identified by the described method included 25 strains of Pasteurella spp. isolated mostly from chamois, deer, ibex, and turtle. Most were also unidentifiable by genomic methods using rrs and rpoB sequence analysis. 12 A possible explanation could be that neither the pathogens nor the commensal bacteria of these animal species have been investigated in detail. Hence, these Pasteurella spp. might represent novel, undesignated, bacterial species. One isolate from a llama could not be identified further than the genus level Actinobacillus. Also, the genomic typing for this strain was inconclusive. The same was true for an isolate of Haemophilus sp. from a cat. In conclusion, the results of the current study suggest that this phenotypic identification procedure allows for the accurate, reliable, and cost-efficient identification of Pasteurellaceae in routine veterinary diagnostic laboratories.
Acknowledgements
The authors are grateful to Andrea Hitz and Edy Vilei for outstanding editorial help. This project was financed by the Research Fund of the Institute of Veterinary Bacteriology, Bern, and by the Federal Veterinary Office, Bern.
Footnotes
a.
SmartGene Ltd., Zug, Switzerland.
b.
Oxoid AG, Basel, Switzerland.
c.
BioMérieux S.A., Geneva, Switzerland.
d.
Becton Dickinson AG, Basel, Switzerland.
