Abstract
Porcine circovirus type 2 (PCV2) is now recognized as the essential infectious component of porcine postweaning multisystemic wasting syndrome (PMWS). PMWS was first recognized in high-status, specific pathogen-free pigs in Canada in 1991 and is now an economically important disease that affects the swine industry around the world. Recently, reports of genomic studies on PCV2 viruses indicated that 2 distinctive genogroups of PCV2 exist. 4,10 This report involves the results of a study on the distribution of predominant PCV2 genogroups recovered from samples taken from PMWS-affected and PMWS-nonaffected farms on the island of Ireland over a 9-year period and the results of a study on PCV2 genogroup recovery from fecal samples taken from a farm in Northern Ireland from 2003 to 2005 that was first diagnosed as PMWS positive in August 2005. The results indicate that, although at least 2 distinct genogroups of PCV2 have been circulating on pig farms on the island of Ireland, there does not appear to be a direct relationship between infection with these different genogroups of PCV2 and the development of PMWS.
Introduction
Postweaning multisystemic wasting syndrome (PMWS) is now recognized as an epizootic disease characterized by clinical signs that can include growth retardation, dyspnea, enlargement of inguinal lymph nodes, diarrhea, and/or occasionally jaundice. 12 PMWS causes significant economic losses to pig producers around the world. Porcine circovirus type 2 (PCV2) is now recognized as being the essential infectious agent of PMWS. 1–3,5,7,8,10 The global explosion of PMWS in the last 5 to 10 years has raised many questions regarding the source and nature of the disease epizootic. Retrospective testing of sera and tissue samples from pigs has shown that PCV2 infections occurred in pigs from at least 1969, 9,11 and sporadic cases of classical PMWS have now been identified from 1985 onward. 11,13
However, field observations and epizootiologic studies in the United Kingdom and Denmark strongly suggest that the spread of PMWS since 1999 has been consistent with the introduction of a “new” infectious agent into a naïve population. 14,16 In contrast, a recent report on the PMWS epizootic in Sweden concluded that the spread pattern of the disease in that country does not support the introduction of a new infectious agent as the cause. 15 Recent reports on genomic analysis of PCV2 viruses recovered from swine herds in 3 states in the United States that suffered from “sporadic cases of an acute onset disease of high mortality” 4 detailed the identification of 2 distinct PCV2 genogroups circulating in that country. It is not known if these differences in the U.S. PCV2 isolates are truly significant in regard to disease severity. The present study reports the results of a temporal study on the prevalence of predominant PCV2 genogroups based on the criteria used for PCV2 isolates from U.S. pig herds. The PCV2 isolates were recovered from pigs on PMWS-affected and PMWS-nonaffected farms in Ireland and from samples from a farm in Northern Ireland collected at regular intervals from 2003 to 2005 that was initially diagnosed with PMWS in 2005.
PCV2 genogroups recovered from farms in Northern Ireland and Ireland with and without PMWS from 1997 to 2006.∗
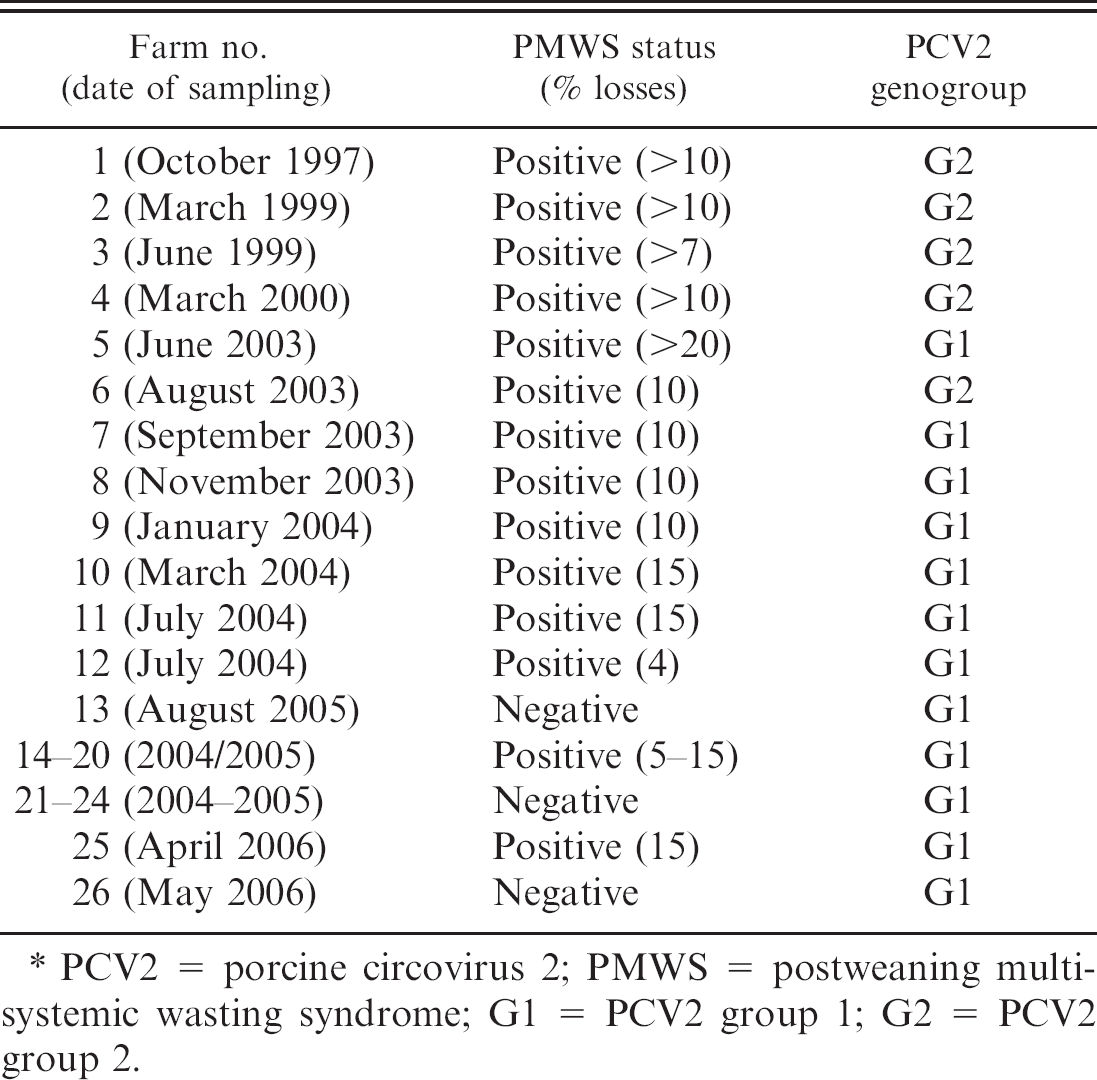
PCV2 = porcine circovirus 2; PMWS = postweaning multisystemic wasting syndrome; G1 = PCV2 group 1; G2 = PCV2 group 2.
Materials and methods
Sample collection
PMWS-affected and PMWS-nonaffected farms. Samples from pigs on 26 PMWS-affected and PMWS-nonaffected farms on the island of Ireland, collected over a 9-year period (1997–2006), were tested by PCR for the presence of PCV2 nucleic acid. The PMWS diagnosis on the selected farms was carried out by using the internationally accepted criteria of clinical disease, histologic lesions, and the PCV2 viral load. 12 A total of 20 PMWS-affected farms with losses attributed to the disease, in pigs primarily between 6 and 12 weeks of age, between 5% and 20%, and 6 PMWS-nonaffected farms were used (Table 1). All the farms studied were free of porcine reproductive and respiratory syndrome virus and swine influenza virus infections and the PMWS-affected farms were, before the outbreak of PMWS, considered to be of high health status. No other significant pathogens were recovered from, or were associated with, the disease outbreaks on the PMWS-affected farms sampled in this study. Where practical, and on all the PMWS-affected farms, the predominant PCV2 nucleic acid for sequence analysis was recovered by PCR from a tissue sample pool from up to 5 diseased pigs. On nondiseased farms, PCV2 nucleic acid was recovered from a fecal pool from up to 5 fecal samples collected from pigs between 6 and 12 weeks of age.
A longitudinal study on a single farm in Northern Ireland. In addition to the study carried out on a selection of farms in Ireland, a 3-year longitudinal study was also carried out on a single pig farm in Northern Ireland. This farm had been regularly monitored for PMWS over a 3-year period beginning in 2003, by analysis of tissue samples from pigs that died after weaning. No evidence of the disease was seen until August 2005 when the farm became affected with PMWS, with losses in excess of 10%. Eighty pooled fecal samples from 8- to 12-week-old pigs (each pool randomly derived from 5 individual samples) from a total of 550 individual fecal samples collected from this farm at regular intervals from 2003 to 2005 and stored at −20°C were analyzed by PCR for the predominant PCV2 DNA (Table 2), as were tissue samples from 10 PMWS-positive pigs euthanized between August and September 2005.
PCV2 genogroups recovered from a longitudinal study on a single farm in Northern Ireland carried out between 2003 and 2005.∗
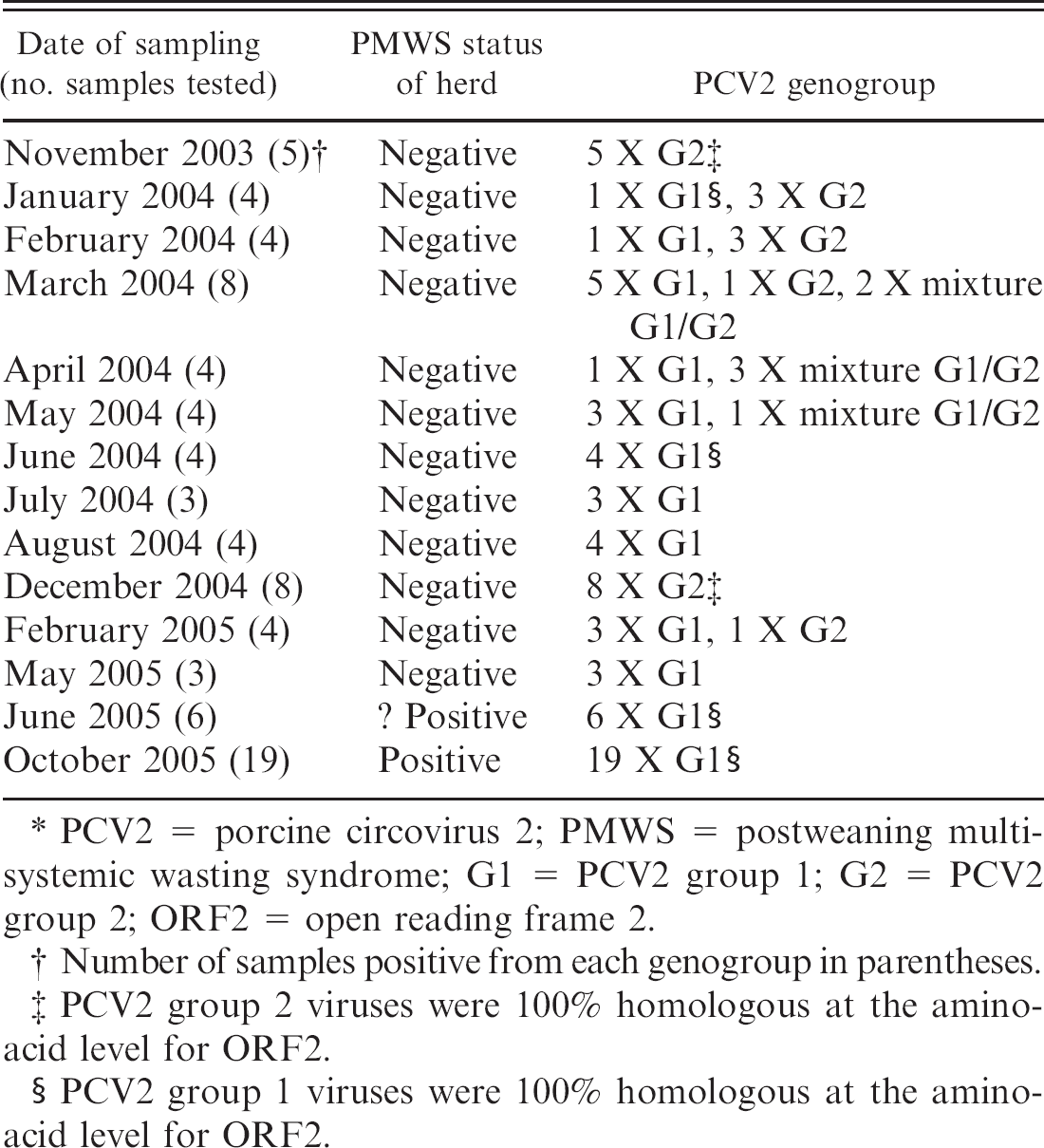
PCV2 = porcine circovirus 2; PMWS = postweaning multisystemic wasting syndrome; G1 = PCV2 group 1; G2 = PCV2 group 2; ORF2 = open reading frame 2.
Number of samples positive from each genogroup in parentheses.
PCV2 group 2 viruses were 100% homologous at the aminoacid level for ORF2.
PCV2 group 1 viruses were 100% homologous at the aminoacid level for ORF2.
Isolation of DNA from feces and tissues
DNA was extracted from both fecal and tissue samples with a QIAamp DNA Blood mini kit, a according to the manufacturer's protocol. This kit, although marketed for use on blood samples, has been found in the authors' laboratory (G. M. Allan, F. McNeilly, I. McNair, Virology Department, Agri-Food and Biosciences Institute, Belfast, Northern Ireland) to be equally effective when used on clarified fecal and tissue sample homogenates (G. M. Allan, F. McNeilly, I. McNair, unpublished data). For each extraction, 200 μl of a 10% suspension in a buffer of homogenized and clarified fecal or tissues samples were used. The resulting DNA was eluted in 100 μl of elution buffer supplied a and stored at −20°C until required.
Identification of PCV2 open reading frame 2. The PCV2-specific DNA was amplified by PCR in duplicate 50 μl reactions with HotStarTaq Master Mix a in a DNA Engine DYAD thermal cycler. b The PCR consisted of a single hot start step of 95°C followed by 40 cycles of denaturation at 95°C for 30 sec, annealing at 55°C for 30 sec, and extension at 72°C for 1 min, followed by a final extension at 72°C for 10 min. Primers PCV2 open reading frame 2 (ORF2) forward (5′-GGT AAT ACT TAC AGC GCA CTT C-3′) and PCV2 ORF2 reverse (5′-CCC CAT GCC CTG AAT TTC C-3′) were used to generate an 814-bp amplicon. 6
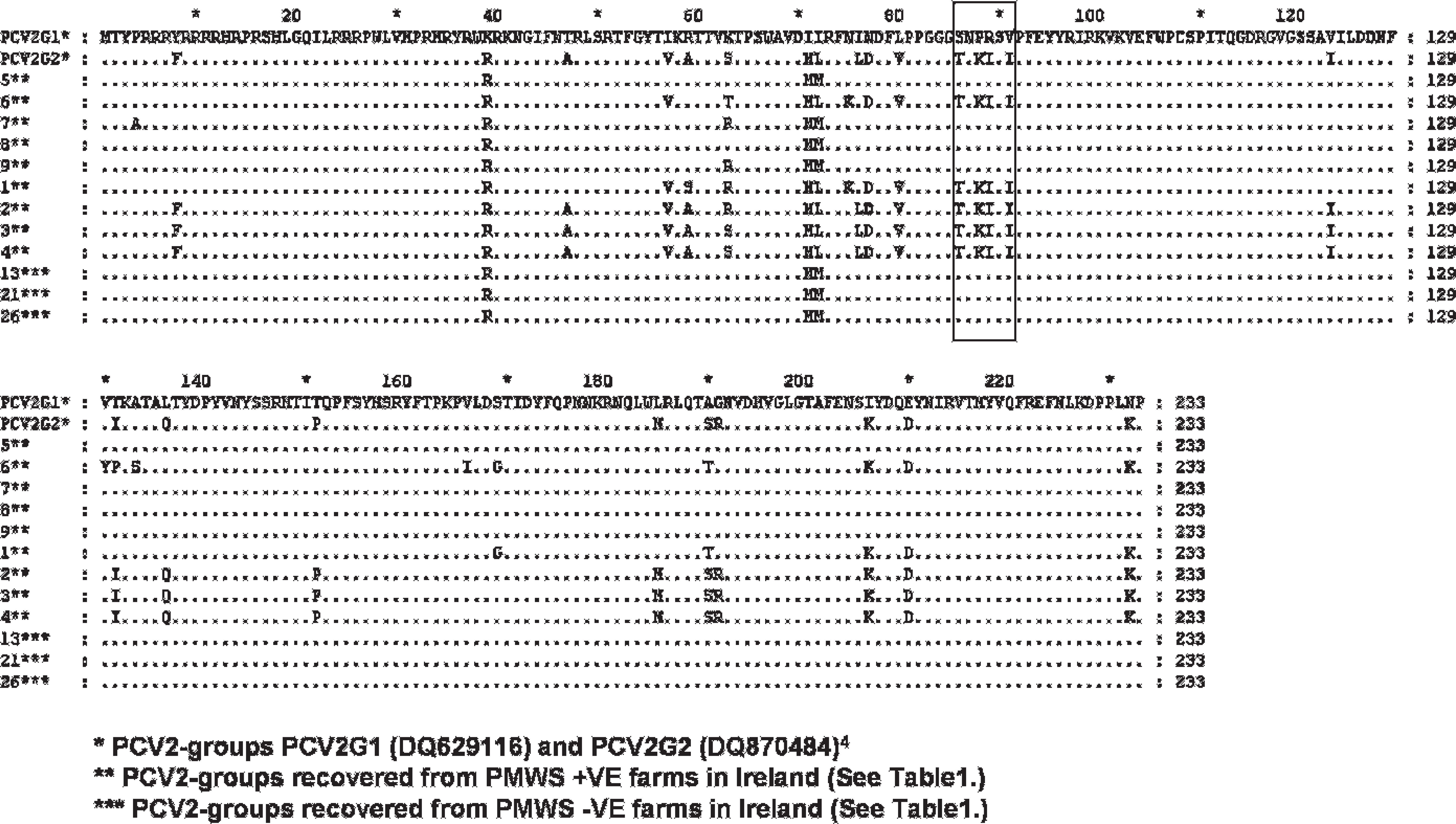

Alignment of predicted amino-acid sequence encoded for ORF2 of a selection PCV2 virus genomes recovered from PMWS-affected and nonaffected herds in the United States 4 (PCV2 G1 and PCV2 G2) and Northern Ireland (1–26). Note the signature region denoting the signature sequence motif is highlighted in box.
Confirmation of amplicon size and purification for sequencing. Amplified PCR products were electrophoresed in a commercially available 1.5% agarose gel c prepared in Tris-acetate-EDTA (TAE) buffer d for 1 hr at 100 V. Amplicon size was estimated by using a 100-bp DNA ladder c and known PCV2 ORF2 amplification controls. Those corresponding amplicons were excised, and PCV2 ORF2 DNA was isolated by using the Wizard SV genomic DNA purification system. f
Sequence analysis of PCR products. The PCV2 ORF2-specific DNA underwent cycle sequencing in triplicate with the previously mentioned gene specific primers (PCV2 ORF2 forward and PCV2 ORF2 reverse). Cycle sequencing was carried out with a BigDye Terminator v3.1 cycle sequencing kit g in a DNA Engine DYAD thermal cycler. b The cycle sequencing consisted of a single denaturation step of 96°C followed by 25 cycles of denaturation at 96°C for 10 sec, annealing at 50°C for 5 sec, and extension at 60°C for 4 min. Extension products were purified by using Performa DTR gel filtration cartridges, h according to the manufacturer's protocol. Cycle sequencing sample electrophoresis was carried out using an Applied Biosystems 3100 Genetic Analyzer. g Sequenced sample data were aligned and analyzed by using the ContigExpress and Align × applications included in Vector NTI software version 10.0 e and GeneDoc (www.psc.edu/biomed/genedoc).
Grouping of PCV2 viruses. The PCV2 nucleic acid sequences detected in this study were genogrouped by using the predicted amino-acid sequence encoded by their ORF2 as described previously for PCV2 genomes recovered from pigs in the United States. 4 In the U.S. study, the authors used cloning, sequence analysis, and alignments to show that the PCV2 ORF2 sequences generated from US samples could be classified into 2 distinct genomic groups. In-depth phylogenetic analysis of recovered complete genomic sequences from a previous study 10 suggested that each of these groups could be further divided into divergent clusters. The authors of the U.S. study proposed that sequence analysis of the capsid gene alone was sufficient to categorize PCV2 isolates as either PCV2 group 1 (PCV2 G1) or PCV2 group 2 (PCV2 G2). 4 The U.S. study proposed that the change in amino-acid sequence positions 86 to 91 found in the ORF2 (ORF2 nucleotide positions 256–273) from SNPRSV to TNKISI 4 acts as a signature motif to distinguish PCV2 G1 viruses from PCV2 G2 viruses, respectively (Fig. 1). This PCV2 G1 and PCV G2 nomenclature was applied to PCV2 genomes recovered from pigs in Ireland and Northern Ireland and analyzed in the present study.
Results
PMWS-affected and PMWS-nonaffected farms
The results of ORF2 genogrouping of PCV2 PCR products from samples from the farms studied in Ireland and Northern Ireland are presented in table 1 and Figure 1. None of the individual samples from pigs on these farms were shown to contain mixtures of PCV2 genogroups, as determined by PCR. However, the PCV2 virus genomes detected in pigs from farm 1 and farm 6 displayed unique aminoacid variations in comparison with other isolates, although these genomes did contain the proposed PCV2 G2 sequence motif. 4 As such, they were classified, for the purposes of this study, as PCV2 G2 viruses. The genomes detected in samples from these 2 farms differed from each other at amino-acid position 131. Sequences from pigs on farm 1 exhibited a threonine at this position and, from farm 6, a proline. At position 63, the genomes recovered from pigs from farm 1 exhibited a unique amino-acid change (compared with other PCV2 G2 isolates in this study) to an arginine, which was also evident in various other Irish PCV2 G1 genomes. Both genomes had a unique amino-acid change at position 169 to a glycine.
Only PCV2 G2 genomes were recovered from samples taken on 4 PMWS-affected farms between 1997 and 2000. Of the pigs sampled on the 4 PMWS-affected farms in 2003, 1 set of samples from a single farm contained PCV2 G2 genome and samples from the 3 other farms contained PCV2 G1 genomes. All the PCV2 genomes recovered from pigs on the 12 PMWS-affected farms sampled from 2004 to 2006 were classified as PCV2 G1. The PCV2 G1 genome only was also recovered from pigs on all 6 of the PMWS-negative herds tested in this study from 2004 to 2006.
The longitudinal study
The results of genogrouping of the PCV2 genomes recovered from the 80 samples collected over 3 years from a single farm in Northern Ireland from 2003 to 2005 are summarized in table 2. Initially (November 2003), the samples tested contained only PCV2 G2 genomes. However, in the samples tested from January 2004 to May 2004, both PCV2 G1 and PCV2 G2 genomes were detected. Occasionally, both PCV2 genogroups were detected in the same sample. With the exception of the 8 samples collected in December 2004 and 1 of the 4 samples collected in February 2005, which were shown to contain PCV2 G2 only, all other samples tested from June 2004 to 2005 were shown to contain only PCV2 G1. The predicted amino-acid sequence coded for ORF2 of the PCV2 G2 recovered throughout the study were 100% homologous, as were the predicted amino-acid sequences of the recovered PCV2 G1 genomes.
Discussion
PCV2 is recognized as a necessary infectious agent for the development of PMWS, an economically important and fatal disease of swine that was reported in all of the major swine-producing countries around the world. 12 The present study reported the results of 2 separate but related studies that were initiated to clarify if the infection of pigs on the island of Ireland with specific genogroups of PCV2 was related to outbreaks of PMWS over a period of up to 9 years.
In the field study, it is of note that, of the first 6 farms confirmed as PMWS positive from 1997 to 2003, the PCV2 G1 genome was detected in pigs in only one of these (farm 5), which indicated that infection of pigs on a farm with this genogroup may not be essential for the development of clinical PMWS. However, it is also of note that farm 5 in this study had, by far, the worst disease scenario of the first 6 farms studied, with losses from PMWS that averaged in excess of 20% and peaked at 35%, perhaps indicating an increased virulence attributable to the PCV2 G1 virus, compared with the PCV2 viruses detected on the other 5 farms.
The results from this field study also showed that all the PCV2 genomes detected from pigs on PMWS-affected and PMWS-nonaffected farms in Ireland sampled after September 2003 fall within the PCV2 G1 genogroup. This includes samples collected from animals on 6 farms deemed to be PMWS negative. 12 Overall, the findings reported in this study support those of the recent study in the United States on PCV2 genotypes currently circulating on diseased and nondiseased farms in that country. 4 In the U.S. study, the authors suggest that PCV2 can be effectively genogrouped on the basis of the ORF2 coding sequence. In another separate study of PCV2 whole genome ORF1 and ORF2 sequences of 148 sequences from the National Center for Biotechnology Information (NCBI) nucleotide database, the authors also identified 2 main groups of PCV2 (1 and 2). No correlation between groups and health status or between groups and country of isolation was reported in this study, only a relationship between group and year of submission of the sequence. Interestingly, 92% of the sequences reported by these authors that belonged to PCV2 G1 were submitted to the NCBI database after 2003, whereas the majority (62%) of the sequences that belonged to PCV2 G2 were submitted before 2003. The authors suggested that this difference might indicate that PCV2 G1 is a more recent emerging virus than PCV2 G2 virus. This suggestion is consistent with the findings in the present study and the findings recently reported in the United States. The current study also supports the suggestion in the U.S. study that the amino-acid changes at amino-acid positions 86 to 91 in ORF2 from SNPRSV to TNKISI 4 could be recognized as a signature motif to distinguish PCV2 G1 from PCV2 G2. In the previous study of 148 ORF2 sequences from the NCBI nucleotide database, 10 the authors did note the equivalent nucleotide changes that correspond to the proposed signature motif and did suggest the ORF2 as a potential phylogenetic marker. However, these signature changes were included as part of 33 distinct nucleotide changes throughout the ORF2 and were not distinguished as a unique identifier of either PCV2 G1 or PCV2 G2.
It is interesting to note that, between 1997 and the spring of 2003, only a few sporadic outbreaks of PMWS, with losses of between 5% and 10%, were seen on farms in Northern Ireland. The severe epizootic of PMWS in Ireland occurred from the end of 2003 to 2006. PCV2 G1 virus was not found on 4 of the 5 farms sampled for this study before the disease epizootic. In addition, it is of interest that PCV2 G1 was recovered from all the farms sampled after August 2003, perhaps indicating that the introduction or emergence and spread of a new PCV2 genogroup in Ireland corresponds with the initiation of a severe epizootic of disease. However, the recovery of PCV2 G1 viruses from nondiseased herds during the disease epizootic argues against the emergence of a new, more virulent genogroup of PCV2 that was, in itself, responsible for this disease scenario.
The results of the longitudinal study on a single farm in Northern Ireland also support the hypothesis that the emergence of a new PCV2 G1 virus was not, on its own, responsible for the breakdown of this farm with PMWS. The first indication of PMWS on this farm was seen in August 2005 when pigs were found to contain lesions consistent with PMWS associated with massive amounts of PCV2. The longitudinal study carried out on fecal samples from this farm clearly showed that 2 PCV2 genogroups circulated on the farm. It is of interest that only PCV2 G2 virus was detected in the 8 samples taken from this farm on December 2004 and that the predicted amino-acid sequences of these PCV2 G2 viruses encoded for ORF2 were 100% homologous to the predicted amino-acid sequences of the PCV2 G2 viruses encoded for ORF2 previously recovered from the farm. This perhaps indicates a single source of reintroduction of this virus onto the farm. It is also interesting to note that the samples collected after this possible incursion and recovery of PCV2 G2 virus contained both PCV2 G1 and PCV2 G2, indicating the possible reintroduction of PCV2 G1 from a common outside source into the herd. It is currently unknown if the immunologic response in pigs to infection with a specific genogroup of PCV2 (PCV2 G1 and PCV2 G2) confers protection against infection with a heterologous genogroup. In vitro cross-neutralization studies and in vivo cross-protection studies are currently underway within the current group of authors to determine if this is or is not the case. Early results of viral neutralization assays indicate a degree of cross neutralization between PCV2 G1 and PCV2 G2 viruses when tested against polyclonal rabbit hyperimmune antisera that would suggest at least a partial immune resistance to heterologous challenge (F. McNeilly unpublished data). All the samples taken from this farm from May 2005 onwards were shown to contain PCV2 G1 virus only, and the predicted amino-acid sequences of these PCV2 G1 viruses encoded for ORF2 recovered from the farm when PMWS occurred were 100% homologous to the predicted amino-acid sequences of the PCV2 G1 viruses encoded for ORF2 previously recovered from the farm when PMWS was not present. These results clearly indicate that the presence of PCV2 G1 virus on this farm was not in itself causal for the farm to develop PMWS. In addition, it would appear that, on this farm, the cocirculation of and presumably coinfection of pigs with 2 different genogroups of PCV2 was also not an important factor in the development of PMWS.
The present results from studies on farms in Ireland and Northern Ireland support this suggestion of a temporal relationship between the emergence of PCV2 viruses. In view of these supportive findings and in an attempt to clarify the confusing nomenclature currently emerging with respect to genogroups/genotypes of PCV2, consideration should be given to a simpler classification, essentially based on the differing amino-acid sequences at positions 86–91 observed on the ORF2 of PCV2 (Fig. 1). 4 Thus, it is proposed that PCV2-group 1 (PCV2 G1) viruses should be defined as having an ORF2 sequence characteristic of the majority of early isolates of PCV2 (group 210), and that PCV2-group 2 (PCV2 G2) viruses should be defined as having an ORF2 sequence characteristic of the majority of more recent PCV2 viruses (group 110). The use of a consistent and defined nomenclature, based on the overall documented temporal emergence of these PCV2 reported to date and discussed above, would avoid the use of misleading and inaccurate terms such as “European” and “American” strains of virus.
In conclusion, the results generated in this study do not support the hypothesis that infection of pigs with PCV2 G1 virus as determined by the predicted amino-acid sequence encoded for ORF2 was, in itself, responsible for the development of epizootic PMWS on pig farms on the island of Ireland. However, further in vivo and in vitro studies are required on the newly documented genogroups of PCV2 virus to determine if biological differences in these viruses exist and, if so, do these reflect differences in pathogenicity in controlled experimental infections and field cases of disease.
Acknowledgements
This research was partially supported by the Higher Education Authority of the Republic of Ireland, the EU contract PCVD-FP6 513928, and by Merial.
Footnotes
a.
Qiagen Ltd., Crawley, United Kingdom.
b.
MJ Research Inc., Waltham, MA.
c.
Mast Group Ltd., Merseyside, United Kingdom.
d.
Sigma-Aldrich Ltd., Gillingham, United Kingdom.
e.
Invitrogen Ltd., Paisley, United Kingdom.
f.
Promega UK Ltd., Southampton, United Kingdom.
g.
Applied Biosystems, Foster City, CA.
h.
Edge Biosystems, Gaithersburg, MD.
