Abstract
A missense mutation of the guanine nucleotide binding protein alpha stimulating activity polypeptide 1 (GNAS) gene, typically Arg201Cys or Arg201His (R201H/R201C), leads to constitutive activation of the Gsα-cyclic AMP (cAMP) signaling pathway that causes several diseases. However, no germline mutations of GNAS have been identified to date, likely due to their lethality, and no robust human cell models have been generated. Therefore, the aim of this study was to generate GNAS-mutated disease-specific induced pluripotent stem cells as a model for these diseases. We then analyzed the functionality of this induced pluripotent stem cell model and differentiated epithelial cells. We generated disease-specific induced pluripotent stem cells by introducing a mutation in GNAS with the clustered regularly interspaced short palindromic repeats (CRISPR) nickase method, which has lower off-target effects than the conventional CRISPR/Cas9 method. We designed the target vector to contain the R201H mutation in GNAS, which was transfected into human control induced pluripotent stem cells (Nips-B2) by electroporation. We confirmed the establishment of GNASR201H-mutated (GNASR201H/+) induced pluripotent stem cells that exhibited a pluripotent stem cell phenotype. We analyzed the effect of the mutation on cAMP production, and further generated teratomas for immunohistochemical analysis of the luminal epithelial structure. GNAS-mutated induced pluripotent stem cells showed significantly higher levels of intracellular cAMP, which remained elevated state for a long time upon hormonal stimulation with parathyroid hormone or adrenocorticotropic hormone. Immunohistochemical analysis revealed that several mucins, including MUC1, 2, and MUC5AC, are expressed in cytokeratin 18 (CK18)-positive epithelial cells. However, we found few CK18-positive cells in mutated induced pluripotent stem cell–derived teratoma tissues, and reduced MUCINs expression in mutated epithelial cells. There was no difference in CDX2 expression; however, mutated epithelial cells were positive for CEA and CA19-9 expression. GNASR201H-mutated induced pluripotent stem cells and GNASR201H-mutated epithelial cells have distinct phenotypic and differentiation characteristics. We successfully established GNASR201H-mutated human induced pluripotent stem cells with increased cAMP production. Considering the differentiation potential of induced pluripotent stem cells, these cells will be useful as a model for elucidating the pathological mechanisms of GNAS-mutated diseases.
Keywords
Introduction
Guanine nucleotide binding protein alpha stimulating activity polypeptide 1 (GNAS) stimulates the activity the adenylate cyclase enzyme, which controls the activity of many hormones and cytokines.1,2 The GNAS gene on the long arm of human chromosome 20 produces multiple gene products, including the alpha subunit of the stimulating guanine nucleotide binding protein (Gsα), a signaling protein that mediates effects through the generation of the secondary messenger cAMP, extra-large Gsα (XLαs), A/B (also referred to as 1A), and neuroendocrine secretory protein (NESP55).3–7GNAS also results in transcripts for coding and non-coding regions that are derived from particular alleles of a parent. The transcriptional and splicing profiles of GNAS are very complex. In the process of transcription initiation and RNA processing, NESP55, XLαs, and A/B individually initiate transcription from their own exon 1 using unique promoters, and the transcript is further spliced onto exons 2–13. Gsα is biallelically expressed in most tissues. However, GNAS gives rise to other gene products, most of which exhibit exclusively monoallelic expression.
Multiple differentially methylated regions (DMRs) are located within the GNAS locus, which contains different promoters. Due to silencing of the GNAS promoter at the methylated allele, XLα, A/B, and GNAS-AS1 transcripts are expressed only in the paternal line, whereas NESP55 transcripts are expressed only in the maternal line. 8 The underlying mechanism of the tissue-specific maternal expression of Gsα is not well understood. However, it is clear that this epigenetic event significantly contributes to the parent-of-origin–specific phenotypic changes of GNAS mutation.
The GNASR201H and GNASR201C mutations are associated with several highly specific pathological molecular phenotypes. However, both mutations activate the GNAS-cAMP pathway, and no major differences in disease development have been reported. McCune–Albright syndrome (MAS) is one of the most well-characterized GNAS mutation-related rare genetic disorder that was originally characterized as a disease with a triad of polyostotic fibrous dysplasia (FD) of the bone, gonadotropin-independent precocious puberty, and café-au-lait skin pigmentation. 9 However, there does not appear to be a parental allele bias in patients with MAS. 10
GNAS R201H/C mutations have been linked to low-grade and benign tumors, especially gastrointestinal or pancreatic lesions.11,12 Intraductal papillary mucinous neoplasms (IPMNs) often have a somatic GNASR201H/C mutation. Gastric lesions that harbor GNAS mutations have also been observed in pyloric gland adenomas, oxyntic gland adenomas, gastric heterotopia, and gastric mucin cell metaplasia of the stomach.13,14 Since a GNAS point mutation can cause different diseases, generation of induced pluripotent stem cells (iPSCs) with this mutation would be useful for studying these diseases in more detail. 15
Emerging gene editing technology using the clustered regularly interspaced short palindromic repeats (CRISPR) system has provided new investigative opportunities.16–20 CRISPR-associated protein 9 (Cas9) nuclease guided by short single guide RNAs (sgRNAs) that recognize the target DNA can generate double-strand breaks (DSBs) at specific DNA target sites. 20 Although the CRISPR/Cas9 system is effective, off-target cleavage of Cas9 has been observed. 21 This problem of off-target cutting has been mitigated with development of an engineered dual sgRNA system and DNA “nickase” that was derived from Cas9 and creates a single-stranded break (SSB). 19 Thus, Cas9 nickases (Cas9n) can be exploited for more specific CRISPR editing, 19 and this strategy could be applied to engineer isogenic embryonic stem cell (ESC) and iPSC disease models with specific mutations introduced or corrected, respectively. Furthermore, this technology could be applied in vivo as well as ex vivo. 22
The purpose of the present study was to establish GNASR201H-mutated (GNASR201H/+) iPSCs to obtain a model for GNASR201H-related diseases. Toward this end, we generated GNASR201H/+ iPSCs with Cas9n.
Materials and methods
Cell culture
The wild-type human iPSC line Nips-B2 (HPS0223) 23 was purchased from Riken Bioresource Center (Tsukuba, Japan). Before gene editing, iPSCs were maintained with SNL76/7 feeder cells in human ESC medium (Dulbecco’s modified Eagle medium/Ham’s F-12 medium (DMEM/F12); Invitrogen, Carlsbad, CA, USA) with 20% KnockOut Serum Replacement (Thermo Fisher Scientific, Waltham, MA, USA) supplemented with 1× non-essential amino acids (Merck, Billerica, MA, USA), 2-mM l-glutamine (Thermo Fisher Scientific), 0.11-mM 2-mercaptoethanol (Wako Pure Chemical Industries, Ltd., Osaka, Japan), 1% penicillin/streptomycin (Thermo Fisher Scientific), and 5-ng/mL human basic fibroblast growth factor (bFGF; ReproCELL Inc., Yokohama, Japan). We cultured the iPSCs without feeder cells. For feeder-free culture, iPSCs were plated on an iMatrix-511 (Nippi Inc., Tokyo, Japan)-coated 60-mm dish and cultured in StemFit AK02N medium (Ajinomoto Co., Inc., Tokyo, Japan) supplemented with 1% penicillin/streptomycin. The culture medium was replaced daily.
Targeting donor vector construction
The targeting donor vectors (pDONR-P3P1R, pDONR-P2rP4, pENTR, and pDEST) were kindly provided by Dr Takefumi Sone and Dr Hideyuki Okano. The left arm of the target vector was designed to contain exon 7 and an intron between exons 6 and 7 of the GNAS gene (NG_016194.1). The right arm was designed to contain exons 8–10 with the R201H mutation. The right and left DNA fragments were artificially synthesized (Genewiz, Suzhou, China) and inserted into pDONR-P2rP4 (right arm) and pDONR-P3P1R (left arm) using Gateway BP clonase II enzyme mix (Thermo Fisher Scientific). pDONR and pENTR vectors containing phosphoglycerate kinase (PGK), the murine PGK promoter, a puromycin-resistance gene (Puro), and an N-terminal shortened version of herpes simplex virus type 1 thymidine kinase (ΔTK) were cloned into pDEST using Gateway LR clonase II enzyme mix.
SgRNA design and cloning
The sgRNAs were designed for the human GNAS locus located on chromosome 20 with the CRISPR design tool (http://crispr.mit.edu/). A pair of sgRNAs was designed targeting the intron between exons 7 and 8, so that they were sufficiently close (within 4-bp distance) to generate a DSB. The complete sequences and locations of the sgRNA target sites are shown in Figure 1(a) and (b). Oligo pairs encoding the 20-nucleotide guide sequence were annealed and ligated into the plasmid pSpCas9 (BB) (formerly pX460; Addgene plasmid ID: 48873).

Generation of GNASR201H iPSCs using the CRISPR/Cas9 system. (a) gRNA sequences (5′–3′). (b) CRISPR-mediated genome editing and genotyping strategy for the GNAS gene. The targeting vector containing a puromycin-resistance gene flanked with loxP. The left arm contains the exon 7 sequence, and the right arm contains the R201H mutation. The primers for target clone selection are indicated with red and blue arrows. (c) Protocol for gene editing of GNAS+/+ iPSCs.
CRISPR/sgRNA transfection in iPSCs
We transfected the CRISPR/sgRNA plasmids and targeting vectors into Nips-B2 by electroporation using the Neon Transfection System (Thermo Fisher Scientific) according to the manufacturer’s instructions. Briefly, iPSCs (1 × 106) were suspended in 100-μL electroporation buffer and electroporated at 950 V for 20 ms with 2 pulses. Seven days after electroporation, puromycin selection (0.25 μg/mL, #A1113802, Thermo Fisher Scientific) was started. Individual colonies were picked and expanded after 14 days (Figure 1(c)).
Sequencing
DNA extraction was performed with the DNeasy Tissue Kit (Qiagen AG, Switzerland) according to the manufacturer’s instructions. Primers used for the polymerase chain reaction (PCR) are listed Table 1. Sequencing reactions were carried out using the BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems, Rotkreuz, Switzerland) and analyzed with the ABI Prism 3130 Genetic Analyzer (Applied Biosystems, Rotkreuz, Switzerland).
Sequence primer.

Cell proliferation assay
We measured cell proliferation using WST-1 proliferation assay kit (MK 400: Roche Diagnostics GmbH, Mannheim, Germany). We performed WST-1 proliferation assay according to the manufacturer’s protocol. Briefly, we cultured iPSCs at the density of 1 × 104 in 96-well plate and incubate for overnight. WST-1 solution was added to each well and incubated for another 30 min at 37°C. The absorbance at 450 nm was measured and the reference wavelength was 650 nm.
Embryoid body formation
After the medium was removed, the cells were rinsed twice with phosphate-buffered saline (PBS), and the iPSCs were dissociated using a cell scraper and transferred to low-attachment Petri dishes to generate embryoid bodies (EBs). EBs were maintained in human ESC medium without bFGF for 21 days. The culture medium was replaced every 3 days.
RNA isolation and reverse transcription
Total RNA was extracted using QIAzol reagent (Qiagen Inc., Hilden, Germany) according to the manufacturer’s instructions. Complementary DNA (cDNA) was synthesized using a high-capacity cDNA reverse transcription kit (Applied Biosystems, #4368814). All cDNA samples were stored at −20°C. PCR was performed with GoTaq (Takara, Japan). The KD human lip fibroblasts (JCRB9103; JCRB Cell Bank, Osaka, Japan) and distilled water (DW) were used as negative control. Primer sequences used for the PCR are listed in Table 2.
RT-PCR primer.

PCR: polymerase chain reaction.
Alkaline phosphatase staining and immunofluorescent microscopy
The iPSCs were washed twice with PBS, fixed in 4% paraformaldehyde in PBS for 10 min at room temperature, and washed twice with DW. The fixed cells were stained with an alkaline phosphatase (ALP) substrate solution (Roche Diagnostics, Basel, Switzerland) for 30 min at room temperature.
For immunocytochemistry, after fixation and washing with PBS, the cells were incubated with the primary antibodies against the following molecules: NANOG (Wako, Japan; 1:200 dilution), SSEA-4 (R&D Systems; 1:100 dilution), and TRA1-60 (R&D Systems; 1:100 dilution) for 60 min at room temperature. The secondary antibodies were fluorescein-conjugated anti-rabbit IgG (H + L) (VECTOR; 1:200 dilution) and fluorescein-conjugated anti-mouse IgG (CAPPEL; 1:1000 dilution).
Teratoma formation
The iPSCs cells (1 × 106 cells in 20-μL PBS) were injected into the testis of 8- to 10-week-old male CB17 SCID mice (Charles River). Teratomas were resected 9–12 weeks after injection, fixed with 10% neutral-buffered formalin for 24 h, and paraffin-embedded sections (4 μm) were processed and stained with hematoxylin and eosin. All mouse studies were carried out according to protocols approved by the Animal Research Committee of Tokyo Dental College (No. 270401).
cAMP assay
cAMP accumulation was measured using homogeneous time-resolved fluorescence (HTRF) cAMP dynamic kits (CisBio, Codolet, France) according to the manufacturer’s protocol. iPSCs were cultured in 96-well plates (20,000 cells/well) and treated with parathyroid hormone (PTH) (100 nM; Sigma-Aldrich, Buchs, Switzerland) 24 or adrenocorticotropic hormone (ACTH) (10 nM; Sigma-Aldrich, Buchs, Switzerland) 25 along with 500-nM 3-isobutyl-1-methylxanthine (IBMX; Cayman Chemical, Ann Arbor, MI, USA). At each time point, the cells were lysed with 1% Triton X-100. cAMP accumulation was detected by HTRF (λex = 330 nm, λem = 665 and 620 nm) using a microplate reader (Synergy, BioTek, Winooski, VT, USA).
Protein extraction and immunoblotting
The iPSCs were lysed in the RIPA Lysis Buffer with 2-mM PMSF, 1-mM sodium orthovanadate and inhibitor cocktail (Santa Cruz Biotechnology) and protein concentrations were measured by a Qubit™ Protein Assay kit (Invitrogen). Equivalent protein concentrations were resolved by electrophoresis on SuperSep™Ace 5%–20% gels (Wako) and transferred to a PVDF membrane. The membrane was probed with anti-GNAS and Anti-beta Actin antibody (1:2000; Proteintech and Abcam), followed by horseradish peroxidase-conjugated goat anti-rabbit IgG and horseradish peroxidase-conjugated anti-mouse IgG. Bound antibodies were visualized using a chemiluminescent substrate (ECL™ Primer Western Blotting Detection Reagent; GE Healthcare Ltd., Buckinghamshire, United Kingdom) and the ImageQuant LAS4000 mini instrument (GE Healthcare).
Immunohistochemistry
Mouse monoclonal antibodies (anti-CK18: clone DC-10, 1:500; anti-MUC1: clone Ma695, 1:500; anti-MUC2: clone Ccp58, 1:500; anti-MUC5AC: clone CLH2, 1:500; anti-MUC6: clone CLH5, 1:500; anti-CEA: clone 12-140-10, 1:500; anti-CA19-9: clone C241:5:1:4, 1:500, Novocastra, Leica Biosystems, Newcastle, United Kingdom) and mouse and rabbit monoclonal antibodies (anti-beta III tubulin: clone 2G10, 1:200; anti-NeuN: clone EPR12763, 1:200, Abcam, Cambridge, Massachusetts, USA) as neural tube labeling were used for immunohistochemistry. Four-micrometer-thick sections were deparaffinized and pre-treated with Epitope Retrieval Solution (EDTA, pH: 8.8) at 98°C for 20 min. Peroxidase blocking was carried out for 10 min using the Bond Polymer Refine Detection Kit DC9800 (Leica Microsystems). The tissues were incubated with the primary antibody for 4°C overnight, washed four times with Tris-buffered saline, and incubated with peroxidase-labeled anti-mouse antibody (Histofine SimpleStain MAX PO, Nichirei, Tokyo, Japan) at 37°C for 30 min. Peroxidase activity was detected with diaminobenzidine (DAB) for 10 min. Qualitative and semi-quantitative analyses of MUC1, MUC2, MUC5A, MUC6, CK18, CDX2, CEA, and CA19-9 expression were performed for the epithelia of ductal structures (ductal structures with a single epithelial layer). Tissue sections were examined under a light microscope using a 4× objective. The pictures were edited with the AxioVision imaging program (version 4.7.1). Thirty ductal structures in each section from teratomas derived from control and mutated GNAS iPSCs (four teratomas each) were appraised using the following scoring system. Semi-quantitative analysis was performed based on the percentage of positive ductal structures according to the following criteria: 0, no detectable staining; 1+, <25% positive ductal structure; 2+, 25%–49%; 3+, 50%–74%; 4+, >75%. In addition, immunostaining intensity was classified according to the following parameters: 1 (absent or weak expression) or 2 (strong expression). The staining score was then obtained by multiplying the percentage and intensity scores as the final immunoreactive score (IRS), which was subjected to statistical analysis.
Statistical analysis
Means of semi-quantitative scoring were analyzed using the Student t-test or the Mann–Whitney U test (SPSS for Windows, version 17). All data are expressed as the mean ± standard deviation (SD, p < 0.05).
Results
Establishment of GNASR201H iPSCs using CRISPR/Cas9
We performed iPSC gene editing and puromycin selection of edited clones and confirmed that the shape of the edited iPSC colonies was similar to that of the GNAS+/+ iPSCs (Figure 2(a)). The single base substitution that introduced the R201H mutation in the GNAS gene was confirmed in 2 out of 24 clones that survived after puromycin selection (Figure 2(b) and (c)). There were no significant differences in cell proliferation rates among these clones (Supplemental Figure 1).

Confirmation of genome editing using RT-PCR and Sanger sequencing. (a) Morphology of GNASR201H iPSCs. Parental iPSCs (Nips iPSCs) used in the experiment and gene-edited GNASR201H/+ iPSCs. Scale bars: 200 μm. (b) We selected two clones with targeting vector sequences in the designated positions. The PCR product of the left arm was 950 bp, and the PCR product of the right arm was 900 bp. (c) We confirmed the designed mutations by Sanger sequencing. The R201H (cgt → cat) mutation in exon 8 was confirmed in both colonies.
Characterization of GNAS-mutated iPSCs
Expression of ESC marker genes (NANOG, OCT3/4, SOX2, REX1, KLF4, and c-MYC) and ALP positivity were confirmed in GNASR201H/+ iPSCs (Figure 3(A) and (B)). Immunofluorescence staining demonstrated that all cells were positive for SSEA4, NANOG, and TRA1-60 (Figure 3(C)). We grew GNASR201H/+ iPSCs as free-floating cultures to form EBs (Figure 3(D)). The EBs maintained in free-floating culture conditions expressed markers of all three germ layers: MAP2 and PAX6 as ectodermal markers, MSX1 and BRACHYURY as mesodermal markers, and AFP and SOX17 as endodermal markers (Figure 3(E)). Teratomas were removed, fixed, paraffin-embedded, and sectioned (4 μm) for hematoxylin and eosin staining, which also identified tissues representing all three germ layers, including neural tube-like structures and melanocytes as in the ectoderm (Figure 3(F)(a) and (b)), cartilage as in the mesoderm (Figure 3(F)(c)), and a gut-like epithelium as in the endoderm (Figure 3(F)(d)). Neural tube-like structures revealed positive staining for both beta-tubulin and NeuN, confirming that these structures were indeed neural tubes (Supplemental Figure 2).
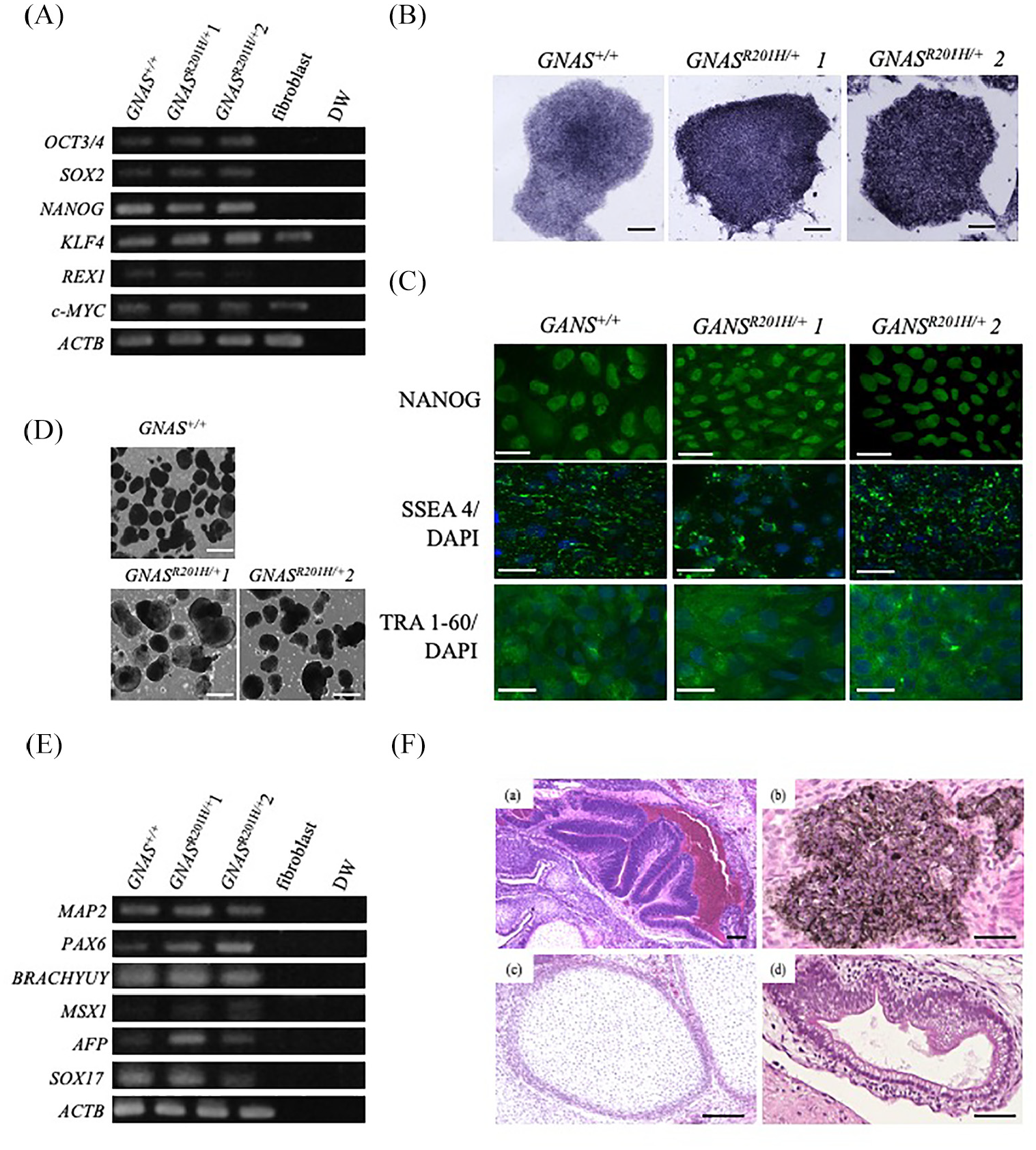
Confirmation of GNASR201H iPSC pluripotency. (A) RT-PCR for the expression of embryonic stem cell markers in GNASR201H iPSCs. ACTB was used as an internal control. KD fibroblasts and distilled water DW were used as negative controls. (B) Alkaline phosphatase (ALP) activity of GNASR201H. (C) Immunofluorescence analysis of pluripotency markers (NANOG, SSEA4, and TRA1-60) for GNASR201H iPSCs. Nuclei were stained with DAPI. Scale bars: 200 µm. (D) Embryonic bodies were generated from GNASR201H/+ iPSC cells expressing the marker genes of the three germ layers. Scale bars: 100 μm. (E) MAP2 and PAX6 were used as ectodermal markers, MSX1 and BRACHURY were used as mesodermal markers, and AFP and SOX17 were used as endodermal markers. ACTB was used as an internal control. KD fibroblasts and DW were used as negative controls. (F) Representative tissues of the three embryonic germ layers in teratomas, including neural tube-like structures (a) and melanocyte-like cells (b) (ectoderm), cartilage (c) (mesoderm), and gut-like epithelium tissues (d) (endoderm).
Measurement of intracellular cAMP accumulation
Gsα is expressed ubiquitously and there appears to be no parental allele bias in patients with MAS or other GNASR201H-related diseases. Thus, it is assumed that the GNAS activating mutation GNASR201H must cause a dominant effect in generating cAMP due to the constitutive activation of Gsα. Accordingly, we examined the changes in cAMP levels upon PTH and ACTH stimulation. We measured the expression of intracellular cAMP levels at the steady state by HTRF. Compared with that of GNAS+/+ iPSCs (Nips), the intracellular cAMP levels of GNASR201H/+1 and GNASR201H/+2 were significantly increased (Figure 4(a)). In addition, the cAMP levels of GNASR201H/+1 iPS and GNASR201H/+2 were elevated upon ACTH and PTH treatment (Figure 4(b)). No significant difference in Gsα protein level was observed between all iPS clones (Figure 4(c)).

cAMP production in GNASR201H/+1 and GNASR201H/+2 iPSCs treated with G- protein coupled receptor (GPCR) ligands. (a) cAMP production was examined in GNASR201H/+1 and GNASR201H/+2, and in GNAS+/+ Nips iPSCs. The iPSCs were cultured in conventional ESC medium. Even without GPCR ligand stimulation, the basal level of cAMP in GNASR201H iPSCs was higher than that of parental iPSCs (Nips). Six independent experiments were performed. Data are shown as the means ± standard deviation. Student’s t-test was performed to validate the differences between two groups. *p < 0.05. (b) cAMP production upon GPCR treatment. We examined the effects of stimulation with two hormones: parathyroid hormone (PTH) and adrenocorticotropic hormone (ACTH). cAMP generation increased upon hormone stimulation in GNASR201H iPSCs. Data are the means ± standard deviation from two independent experiments, each measuring six samples. *p < 0.05 GNASR201H iPSCs vs GNAS+/+ iPSCs at the same time point. (c) Gsα protein levels were examined by immunoblotting.
Ductal structures in MAS iPSC-derived teratomas have less mucin but more CEA and CA19-9
Despite the fact that GNAS mutations are frequently observed in many gastrointestinal benign tumors, including IPMN, the lack of adequate cell models has restricted the knowledge and studies on its pathological roles in tumor development. Therefore, to investigate whether GNAS mutations cause any changes in functional or differentiation phenotypes, we investigated the protein levels of MUCINs (MUC1, 2, 5AC, and 6), CK18, CDX2, CEA, and CA19-9. We used MUCIN expression as a functional marker of gastrointestinal or pancreatic epithelial cells. Mucins are secreted from ductal epithelial cells to cover its mucosa surface, and play an important role in protecting the mucosa surface, and are also used as lubricants and surfactants. There are several subclasses of mucins whose localizations are mostly tissue-specific. Thus, they are recognized as tissue-specific markers. Indeed, mucins are used as a differentiation marker as well as a functional marker in immunohistochemical examinations for the diagnosis of several tumors. CDX2 is another marker of intestinal and colon epithelium, CK18 is a differentiated epithelial cell marker, and CEA and CA19-9 are standard tumor markers.
We observed many MUC1-, MUC2-, and MUC5A-positive ducts in control iPSC-derived teratomas (Figure 5(a)). Every subclass of mucins was detected at much lower levels in GNAS-mutated iPSC-derived teratomas. However, there was no difference in the level of CDX2 positivity in ductal structures between the control and GNAS-mutated iPSC-derived teratomas. We observed weak CEA and CA19-9 positivity in control iPSC-derived teratomas and strong CEA and CA19-9 positivity in ductal structures from GNAS-mutated iPSC-derived teratomas (Figure 5(a)–(c)). Most of the ductal structures in the control teratomas were CK18-positive, whereas the structures from GNAS-mutated iPSC-derived teratomas were not (Figure 5(c)).

Immunohistochemical analysis of teratomas generated by transplantation of iPSCs into immunocompromised mice. (a) Fifty luminal structures surrounded with simple epithelial cells were selected. Immunohistochemical staining of mucins, CK18, CDX2, CEA, and CA19-9 was performed. Luminal structures in control iPSC-derived teratomas were positive for MUC1, MUC2, and MUC5AC. Almost half of the luminal structures in control iPSC-derived teratomas were CDX2-positive. Most of the epithelial cells were CEA-positive and faintly CA19-9 positive. Scale bars: 500 μm. (b) Fifty luminal structures surrounded with simple epithelial cells were selected and immunohistochemical staining was performed. We found less MUC1-, MUC2-, and MUC5AC-positive cells in GNASR201H iPSC-derived teratomas. CK18-positive cells were seldom observed. Almost half of the luminal structures of simple epithelial cells were CDX2-positive. There was strong positivity for CA19-9 and CEA in luminal structures. Scale bars: 500 μm. (c) Immunoreactivity scores (IRS). Data are the means ± standard deviation from GNASR201H iPSCs-derived teratoma and control GNAS+/+ iPSC-derived teratomas, with 30 luminal structures measured per group.
Discussion
In this study, we generated disease-specific iPSCs with the GNASR201H mutation using the CRISPR nickase system. Pathological mechanisms regarding GNASR201H mutations are yet to be clarified due to the lack of human cell model systems. 26 However, the GNASR201H/+ mutation is a rare somatic mutation, and it is often not feasible to establish cultured cells from patients. Although the establishment of iPSCs derived from FD patients was recently reported, they did not generate the “revertant” iPSCs, in which the mutation is corrected to the wild type with the same genetic background. 27 To overcome this obstacle, we generated artificially mutated GNASR201H/+ iPSCs. We used a double nicking strategy with Cas9n instead of CRISPR/Cas9 to avoid off-target effects. 28 This strategy uses paired Cas9ns that introduce nicks in each DNA strand, causing site-specific DSBs. This technique is highly specific because DSBs occur only when two nickases work simultaneously (Figure 1(b)). To identify pluripotent cells, we measured the expression of stem cell markers and teratoma formation after transplanting iPSCs into immunocompromised mice, and confirmed that the GNASR201H/+-mutated iPSCs exhibit pluripotency.
As GNAS is an imprinted gene, 29 it is possible that introduced mutations may not be transcribed and translated. Therefore, we confirmed the presence of the GNAS mutation with sequencing and functional GNAS activation by examining the effects of hormones that bind to GPCRs. We focused on two hormones as GPCR ligands, PTH and ACTH. PTH is often a target of diseases caused by inactivating mutations within Gsα-coding GNAS exons. ACTH is an ACTH, and its overactivation causes Cushing disease, one of the phenotypes in MAS. 30 As expected, we observed a rapid production of cAMP after ACTH or PTH stimulation and a higher level of sustained cAMP production in mutated iPSCs compared to parental iPSCs.
GNAS encodes a G protein α subunit that is associated with GPCRs. Over 800 different GPCRs are encoded in the human genome, and nearly a third of all drugs target this type of receptor.31,32 Their ligands include hormones, neurotransmitters, and odor molecules or proteins identified by sequence homology, but there are also so-called orphan receptors without known ligands. 31 Many diseases associated with GNAS hyperactivation have been reported due to its involvement in a wide spectrum of biological activities. 33 GNASR201H/+ mutations are associated with gastrointestinal neoplasms, including pancreatic IPMNs and invasive adenocarcinomas arising from IPMNs, colonic adenocarcinomas, and mucinous appendiceal tumors.34–36GNASR201H+ mutations are reported in up to two-thirds of pancreatic IPMNs. 37 In addition, approximately 2% of colonic adenocarcinomas harbor GNAS mutations, and are associated with a villous morphology. Therefore, GNASR201H mutations seem to play an important role in tumorigenesis.
Both villous adenoma and IPMN, which are benign tumors prone to malignant transformation, frequently harbor GNAS mutations. However, progress in elucidating the pathological conditions and developing therapeutic methods has been very slow. iPS cells can be induced into the digestive tract and pancreas; thus, our iPS cells can be used as a tool for examining the tumorigenesis mechanism and discovering effective treatment strategies. A recent study demonstrated that medulloblastoma was frequently detected in teratomas produced by transplanting iPSCs derived from patients with Gorlin syndrome, where medulloblastoma frequently occurs. 38 Thus, even though a teratoma is overall benign, it is possible that some parts of the teratoma will harbor a trait that makes them prone to cancer.
To evaluate the epithelial structures often affected in GNAS-mutated tumors, we examined differentiation markers such as mucins (MUC1, 2, 5A, and 6), CK18, and CDX2. We also investigated the distribution of CEA and CA19-9 as markers of malignant tissues, which offered insight into the differential state that could contribute to malignant potential.
Mucin is a functional protein secreted by many ductal epithelial cells and has many tissue-specific subtypes. The distribution of different mucin subclasses varies depending on the pancreatic or colon tumor pathological phenotype; thus, it is useful for pathological and prognostic diagnosis39–41 Owing to its specificity, mucin has been widely applied for pathological diagnosis using immunohistochemical staining. The distribution of mucin subclasses in lesions differs from that in normal tissues. For example, in Barrett’s esophagus, the pattern of mucin expression differs from that of the esophagus. Moreover, the distribution of mucin expression reflects the origin of the adenocarcinoma (esophageal type, gastric type, and intestinal type). Gastric goblet cells are MUC2-positive, and in gastric cancer tissues, MUC5AC is positive on the apical side, whereas MUC6 is positive on the basement membrane side. GNASH201C mutation is common in villous adenoma, which is considered to be a low-grade malignancy, and is also characterized by MUC5AC and MUC2 expression. The most typical IPMN is characterized by both MUC5AC and MUC2 expression. However, no comprehensive study has been conducted to evaluate the difference in staining properties of mucin from those of normal tissues. Therefore, we examined the immunohistochemical staining of mucin between normal and GNAS-mutated teratoma tissues. Based on the fact that the GNAS mutation was frequently observed in gastric villous adenoma and pancreatic IPMN, immunohistochemical staining patterns of mucin were compared between tissues with wild-type and mutated GNAS. The results showed decreased staining for every subtype of mucin in the samples with mutated GNAS, suggesting that the production of mucin, which is a functional protein, might be decreased in tumors caused by GNAS mutations. Therefore, we examined mucin expression in luminal structures from iPSC-derived teratoma tissues. The same number of luminal structures was examined in control and GNAS-mutated iPSC-derived teratomas; luminal epithelial cells of control teratomas were positive for MUC1, MUC2, and MUC5A as well as for CK18. GNAS-mutated luminal structures had much less mucins and CK18 positivity; almost no CK18-positive epithelial cells were found in GNAS-mutated iPSC-derived teratomas. In contrast to the negative staining observed for mucins and CK18, CEA and CA19-9 frequently show strong positive staining in GNAS-mutated teratomas.42,43
Taken together, these findings indicate that most of the mucins investigated and CK18 are epithelial cell markers, and are expressed on most of the luminal epithelial cells of the normal iPS-derived teratoma but are weakly expressed in GNASR201H+ iPS-derived teratomas. It is speculated that the extreme decrease in the expression levels of all types of mucins as well as CK18, and the more intense positivity of CEA and CA19-9 may indicate that these luminal structures have distinct differentiation patterns from normal structures. This may be caused by undifferentiated or dysplastic properties, developmental abnormalities, or low epithelial functional properties.
To date, only postzygotic mutations (R201C and R201H) have been reported in humans, such as in MAS; in contrast, germline mutations (R201C and R201H) have seldom been reported. 44 Lack of inheritance of the disease in humans is considered to reflect the embryonic lethality of germline-transmitted activating Gsα mutations, which would only survive through somatic mosaicism. Thus, properly controlled GNAS function is likely critical for a proper epithelial phenotype, further supporting that GNAS dysfunction leads to dysplastic or malignant tumor development. Inducing gastrointestinal or pancreatic organoids may provide a clue to reveal the pathogenesis of GANS R201H mutation-related tumors.
Conclusion
GNAS R201H mutations are rare but play a crucial role in tumorigenesis. Unfortunately, it has not been feasible to establish primary cell cultures from patients with these mutations. We succeeded in generating artificially mutated GNASR201H iPSCs. The model developed herein holds promise for determining the detailed pathological mechanisms of diseases with mutated GNAS. Although further studies, such as the generation of gastrointestinal or pancreatic organoids, are warranted, we believe that the new cell line generated in our study has potential for use as a model in the identification of novel therapies to combat these diseases.
Supplemental Material
sup_figure_1 – Supplemental material for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors
Supplemental material, sup_figure_1 for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors by Katsuhito Watanabe, Takashi Nakamura, Shoko Onodera, Akiko Saito, Takahiko Shibahara and Toshifumi Azuma in Tumor Biology
Supplemental Material
sup_figure_2 – Supplemental material for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors
Supplemental material, sup_figure_2 for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors by Katsuhito Watanabe, Takashi Nakamura, Shoko Onodera, Akiko Saito, Takahiko Shibahara and Toshifumi Azuma in Tumor Biology
Supplemental Material
sup_figure_legend_1 – Supplemental material for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors
Supplemental material, sup_figure_legend_1 for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors by Katsuhito Watanabe, Takashi Nakamura, Shoko Onodera, Akiko Saito, Takahiko Shibahara and Toshifumi Azuma in Tumor Biology
Supplemental Material
sup_figure_legend_2_ – Supplemental material for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors
Supplemental material, sup_figure_legend_2_ for A novel GNAS-mutated human induced pluripotent stem cell model for understanding GNAS-mutated tumors by Katsuhito Watanabe, Takashi Nakamura, Shoko Onodera, Akiko Saito, Takahiko Shibahara and Toshifumi Azuma in Tumor Biology
Footnotes
Acknowledgements
We would like to thank Drs Mahito Nakanishi (National Institute of Advanced Industrial Science and Technology –AIST), Manami Ohtaka (AIST), and Ken Nishimura (University of Tsukuba) for providing technical assistance. We would like to thank Dr Hideyuki Okano and Sone (Keio University) for kindly providing the targeting donor vectors (pDONR-P3P1R, pDONR-P2rP4, pENTR, pDEST). We thank Editage (![]() ) for English language editing.
) for English language editing.
Contributorship
W.K. contributed to data curation, writing—original draft, writing—review and editing. N.T. contributed to data curation, writing—original draft, writing—review and editing, conceptualization, funding acquisition, supervision, data curation. O.S. contributed to data curation, formal analysis, funding acquisition, writing—review and editing. S.A. contributed to data curation, formal analysis, funding acquisition. S.T. contribute to writing—review and editing. A.T. contributed to supervision, funding acquisition, writing—review and editing. All authors interpreted the results and reviewed and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All mouse studies were carried out according to protocols approved by the Animal Research Committee of Tokyo Dental College (No. 270401).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a JSPS KAKENHI Grant (no. 19K10063 and 18H03007) and by a Tokyo Dental College Research Branding Project.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
