Abstract
Globally, colorectal cancer is the third most common type of cancer. Genetic instability leading to cancer development is one of the major causes for development of cancer. Alterations in mitochondrial genome, that is, mutations, single-nucleotide polymorphisms, and copy number variations are known to contribute in cancer development. The aim of our study was to investigate association of mitochondrial T16189C polymorphism and copy number variation with colorectal cancer in North Indian population. DNA isolated from peripheral blood of 126 colorectal cancer patients and 114 healthy North Indian subjects was analyzed for T16189C polymorphism and half of them for mitochondrial copy number variation. Genotyping was done using polymerase chain reaction–restriction fragment length polymorphism, and copy number variation was estimated using real-time polymerase chain reaction, numbers of mitochondrial copies and found to be significantly higher in colorectal cancer patients than healthy controls (88 (58–154), p = 0.001). In the regression analysis, increased mitochondrial copy number variation was associated with risk of colorectal cancer (odds ratio = 2.885, 95% confidence interval = 1.3–6.358). However, T16189C polymorphism was found to be significantly associated with the risk of rectal cancer (odds ratio = 5.213, p = 0.001) and non-significantly with colon cancer (odds ratio = 0.867, p = 0.791). Also, false-positive report probability analysis was done to validate the significant findings. Our results here indicate that mitochondrial copy number variation may be playing an important role in the development of colorectal cancer, and detection of mitochondrial copy number variation can be used as a biomarker for predicting the risk of colorectal cancer in North Indian subjects.
Introduction
Colorectal cancer (CRC) remains the third most common cancer worldwide. In the United States alone, it was estimated to cause 40,000 deaths in 2016. 1 After lung and liver cancer, CRC is also among the leading cause of cancer-related deaths worldwide. 2 CRC generally progresses slowly, initially as a polyp and later develops into cancer. 3 As per American Cancer Society, diet, obesity, overweight, smoking, and alcohol consumption have been associated with the development of CRC. Although 70% cases of CRC patient are reported to be sporadic, 30%–35% have been shown as familial and inheritable.4–6 Early diagnosis of different cancer types including CRC can help in better management of cancers. 7 Genetic variations associated with CRC would serve as an important diagnostic tool for such management. Different genetic variability including those of GSTT1, GSTP1, GSTM1, PTEN, and IGF-1, IGFBP-3 has been found to be associated with the risk of CRC.8–12 GST (glutathione S-transferase) is a well-established reactive oxygen species (ROS) detoxifier in our cells, and ROS detoxification is a common anticancer strategy in cellular system. 13 Cancer cells exhibits high levels of ROS generation which promotes genetic instability, where mitochondria remains the major site for ROS generation. 14 Changes in mitochondrial genome are known to promote ROS generation in cancer cells.14,15 Mitochondrial copy number variations (CNVs) and mitochondrial genome polymorphisms are also further reported to contribute ROS and cancer development.16,17 Previous literature have already shown the association of mitochondrial CNV in the peripheral blood leucocytes (PBLs) with risk of different cancers such as melanoma, breast cancer, hepatic carcinoma, and esophageal cancer.18–21 The poly cytosine tract (poly-C) of D-loop region in mitochondrial DNA (mtDNA) is especially important for replication and transcription of mitochondrial genome, thus variations in this region can directly affect function of mitochondria. 22 T16189C variation in this poly-C tract is one such variation which affects the replication of mitochondrial genome, although only limited information is available on the association of mitochondrial polymorphisms with CRC.23,24 In Indian population till date, only one study made on south Indian population showed that T16189C variant was associated with the risk of CRC. 25 In addition, no meaningful information is available on mitochondrial CNV from PBLs in CRC patients. As South Indian and North Indian populations are two distinct populations, in this present case-control study, we have tried to figure out whether T16189C variant and mtDNA CNVs from PBLs are somehow associated with the risk of CRC in North Indian population or not.
Material and methods
Study subjects
A total of 126 tumor subjects who regularly attended Medical Outpatient Department of Gastrointestinal Surgery GB Pant Hospital, Delhi, India were enrolled in this study. The patients were divided into two groups based on location of tumor either colon (n = 78) or rectal (n = 48). Diagnosis of cancer was based on the criteria of histological confirmation. The control group was composed of 114 subjects free from CRC with similar distribution of sex and age with that of tumor patients and having no familial history of CRC. The study was approved by ethical committee of Jamia Millia Islamia. Informed written consent was obtained from all subjects participating in the study.
Sample collection and DNA extraction
Taking full aseptic precautions, fasting blood samples were collected and genomic DNA was extracted from peripheral blood leucocytes using the standard phenol–chloroform method. All samples were gathered consecutively during the detailed time period. The quality and quantity of extracted DNA were estimated first by running it on 0.8% agarose and then using NanoDrop ND-1000 spectrophotometer. After quantification, appropriate dilutions were made for further experiments.
Genotyping
Genotyping was performed using polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) method where region 15911–16540 of mitochondria (630 bp) was amplified. For PCR, 25 μL reaction was setup in a 200 μL PCR tubes. The reaction mixture consisted as follows: 2.5 μL of 10× reaction buffer (with 1.5 mM MgCl2), 2 μL of deoxynucleotide triphosphate (dNTP; 2.5 mM), 2 μL of primer pair each of 12 μM, 50 ng/μL template, 1 U/μL of Taq polymerase (Himedia), and 14.5 μL Milli-Q water. The primer sequences used were, FP 5′-ACCAGTCTTGTAAACCGGAG-3′ and RP 5′-GTGGGCTATTTAGGCTTTAT-3′ and PCR cycle used for amplification was as follows 95ºC for 5 min and 32 cycles of 95ºC for 30 s, 59ºC for 30 s, 72ºC for 30 s, and final extension for 72ºC for 5 min. PCR product (15 μL) was digested overnight at 37ºC in a water bath with 1 U of the Mn1I enzyme in total reaction volume of 50 µL as per manufacturer’s protocol (BioLab). The digested products were resolved on 3% agarose gels at 60 V for 90 min. The T-allele gives a major product of 287 bp, while the C-allele gives a major product of 322 bp; 10% percent of samples were reconfirmed with direct sequencing to check the accuracy of our RFLP data. CRC cancer samples were compared with healthy controls, where “control ref” is a “baseline group.”
CNV
Real-time PCR was performed to estimate mtDNA CNV using PowerUp SYBR Green Master Mix (#A25742; Thermo Fisher Scientific) on Real-Time Detection System (CFX96, #1855195; BioRad). The relative mtDNA CNV for CRC and normal healthy samples was calculated as mitochondrial gene/nuclear genome; 5 ng of total genomic DNA was used as a template to carry out real time polymerase chain reaction (real time-PCR) reaction using primer for mitochondrial 16s ribosomal RNA (rRNA) gene as a mitochondrial gene and β-2 microglobulin (β-2M) gene (single copy) as a nuclear reference gene. Primers used for nuclear DNA (nDNA) β2M are FP 5′-GCTGTCTCCATGTTTGATGTATCT-3′ and RP 5′-TCTCTGCTCCCCACCTCTAAGT-3′ giving an amplicon size of 107 bp. Primers used for mtDNA, 16S rRNA are FP 5′-GCCTTCCCCCGTAAATGATA-3′ and RP 5′-TTATGCGATTACCGGGCTCT-3′ giving an amplicon of 97 bp. The real time-PCR cycle used consisted of initial denaturation at 95ºC for 10 min followed by 40 cycles of 15 s at 95ºC and 1 min at 60ºC. Each sample was studied in duplicate on a 96-well plate (BioRad). A negative control (without template) was included in each plate. Quantities of mtDNA and nDNA were estimated using the equation 2 − ΔCt, where ΔCt = ΔCt mtDNA − ΔCt nDNA. For copy number analysis, “reference” is the baseline group in which mitochondrial copy number is less than or equal to 56 in case of comparison by median and 36 in case of comparison by quartile.
Statistical analysis
Statistical analysis was carried out using standard statistical methods (SPSS software version 17.0). Data were expressed as mean ± standard deviation. In case of high inter-individual variability and non-normal distribution, data were expressed as median and inter-quartile range. Differences between means were compared by Student’s t-test or Mann–Whitney U test (where the data were not distributed normally). Multivariate analysis was done using Bonferroni adjustment to negate the effects for confounding factors like age and sex. Allele and genotype frequencies of CRC patients and healthy controls were compared with values predicted by Hardy–Weinberg equilibrium using the chi-square (χ2) test. Odds ratios (ORs) of tumor subjects associated with genotype with 95% confidence of intervals were calculated by logistic regression analysis assuming dominant mode of inheritance. Sample size for the single-nucleotide polymorphism (SNP) at 5% significance level and statistical power was calculated using OSEE online calculator (Table 3). False-positive report probability (FPRP) analysis was performed using “R” software for assessing our significant findings. FPRP threshold was set at 0.2 along with a prior probability of 0.1 for detecting the associated risk. Values obtained below 0.2 in FPRP were considered noteworthy. 26 Logistic regression was also performed to find out the association of copy number with the risk of cancer. For all statistical tests, p < 0.05 was considered as the level of significance.
Results
In this study, 114 healthy controls with no previous history of any type of malignancy and 126 CRC patients were included. The clinical characteristics of the study groups are summarized in Table 1. There was no significant difference in age and sex distribution between the study groups.
Clinical characteristics and laboratory data of cancer patients and healthy subjects.

CRC: colorectal cancer; SD: standard deviation.
Data are presented by mean ± SD.
Cancer patients versus healthy subjects.
Association of mitochondrial T16189C polymorphism with the risk of cancer development
We have analyzed T16189C polymorphisms in D-loop region of mitochondria to find out its association with the risk of cancer development. The allelic frequency distribution of T16189C variants in controls and tumor subjects is shown in Table 2. The frequency of C-allele was found to be higher in CRC patients (17.18%) than control (8.77%). Also, the frequency of C-allele in rectal cancer (33%) was almost four-fold higher when compared to control (Table 2). Although no significant difference was observed in C-allele frequency of colon cancer compared to control.
Genotype distribution of SNPs in mitochondrial T16189C polymorphism in cancer patients and healthy subjects.

CRC: colorectal cancer.
T represents wild form.
C represents polymorphic form.
Values are presented as n (%).
Genotype distribution was in Hardy–Weinberg equilibrium.
The binary logistic regression analysis revealed that C-allele provides risk for development of CRC (OR: 2.158, p = 0.058) and the p value was found to be very close to significant level. Interestingly when rectal and colon samples were analyzed individually with healthy controls, C-allele provided a very high risk (OR = 5.213, p = 0.001) for development of rectal cancer, while no risk was observed for colon cancer (Table 3). FPRP value was far below 0.2 for the rectal cancer risk suggesting that the association found was not a false positive. However, FPRP values revealed that the risk found for development of CRC was not noteworthy.
Odds ratios and FPRP values for association between mitochondrial T16189C polymorphism in different study groups.

CRC: colorectal cancer; OR: odds ratio; FPRP: false-positive report probability; CI: confidence interval.
Test of association was repeated in logistic regression model adjusting for age, duration, and gender.
Reference baseline group free from any type cancer.
p < 0.05 considered significant, FPRP values (bold values) below 0.2 are considered noteworthy.
Association between mtDNA copy number and CRC
The comparison between mtDNA copy number of CRC and healthy subjects is summarized in Table 4. The levels of mtDNA copy number were not normally distributed in study subjects; hence, they are presented by quartile range. It was found that the levels of mtDNA copies were significantly higher (p = 0.001) in CRC than control subjects (Table 4). Adjustment for potential confounding factors, that is, sex and age by the analysis of covariance using Bonferroni adjustment, revealed that difference in mtDNA copy number among study subjects also remained significant (p < 0.05) as shown in Supplementary Table 2.
Copy number data of tumor patients and healthy subjects.

CRC: colorectal cancer.
Comparison between the groups was performed with Mann–Whitney U Test.
Data are presented by inter-quartile range.
versus healthy subjects.
Separate analysis of mtDNA copy number of colon and rectal cancer with control subjects also showed the significant association with colon (81 (60–143), p = 0.002) and rectal (103 (32–178), p = 0.021) subjects, respectively (Table 4). On dividing the tumor subjects on the basis of their pathology into different grades, significant difference was also observed for mtDNA copy number in tumor grades I + II (83 (53–148), p = 0.005) and tumor grades III + IV (88 (60–190), p = 0.007) on comparing with control.
To find out the risk for development of CRC due to mtDNA copy number, binary logistic regression was carried out between tumor subjects and control (reference), and it was found that increased mtDNA copy number provides a significant risk for development of CRC having odd ratios of 2.885 (p = 0.009). Data analysis by quartile distribution of mtDNA copy number also revealed a dose-dependent association between copy number and CRC risk (highest vs lowest quartile: OR = 5, 95% confidence interval (CI) = 1.659–15, p = 0.004). The presence of greater than 73 mtDNA copies per nuclear genome provides highly significant risk for development of CRC in North Indian population (Table 5). Also, no significant effect of different parameters like smoking and alcohol was observed on CNV among the patients (Supplementary Table 1). While all parameters except smoking showed significant difference of copy number between CRC patients and control (Supplementary Table 2).
Association of copy number with cancer development.
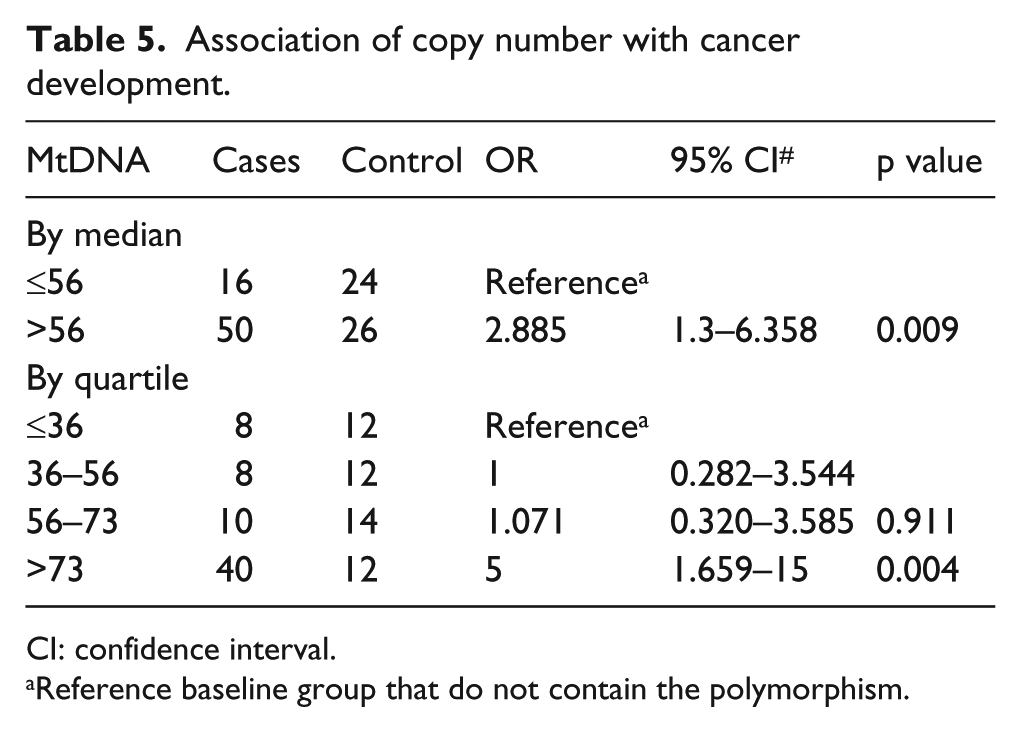
CI: confidence interval.
Reference baseline group that do not contain the polymorphism.
Association of mitochondrial T16189C polymorphism and mtDNA copy number
To confirm the role of T16189C polymorphism in controlling the mtDNA copy number, association of this polymorphism in different samples with their mtDNA copy number was tested. Interestingly, no association was found between the presence of T-/C-allele and mtDNA copy number in both CRC patients and controls (Table 6).
Association of mitochondrial T16189C polymorphisms and copy number data in cancer patients and healthy subjects.
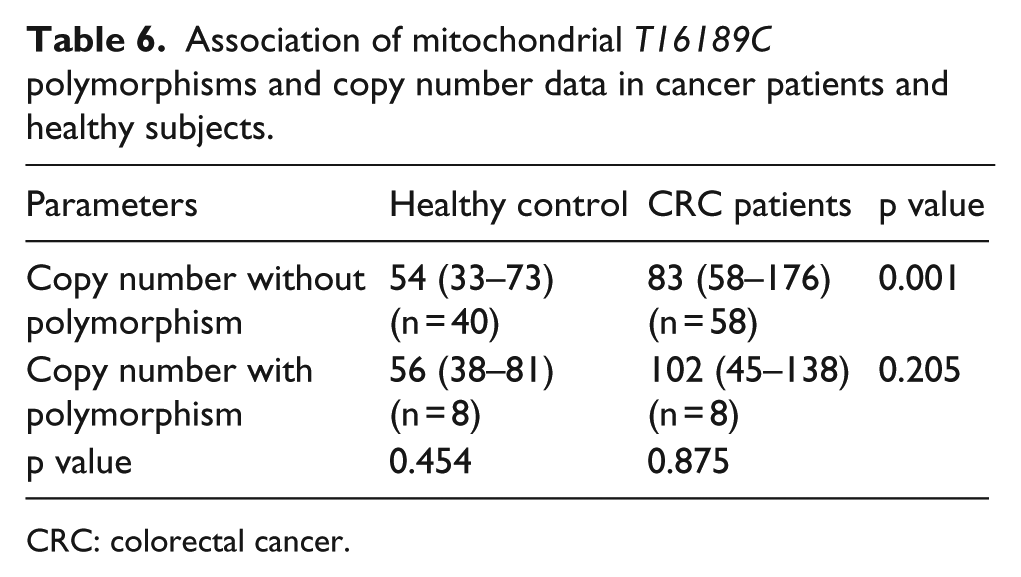
CRC: colorectal cancer.
Discussion
This study was carried out on 230 North Indian subjects including 126 CRC patients, diagnosed between 2013 and 2016, and age-, sex-, and ethnicity-matched healthy controls. Mitochondrial mutation polymorphisms and copy number have been shown to play an important role in development of many diseases including cancer. Usually, these variations in mtDNA destabilize intracellular ROS balance which leads to initiation or progression of the disease. 27 mtDNA contains hyper variable regions (HVR I, II, and III) which are the prime sites where such changes occur. In the D-loop region, this T to C transition at position 16189 generates poly-C tract. It has been suggested that this poly-C tract can regulate mtDNA copy number leading to disease phenotype. Continuous cytosine residues were associated with reduced mtDNA copy number.22,28 A common variant T16189C from HVR III region has been associated with different diseases such as Type II diabetes mellitus (DM) among Indians and other Asians,29,30 coronary artery disease among middle Europeans, 31 metabolic syndrome among Caucasians, 32 insulin resistance, 33 and dilated cardiomyopathy. 34 T16189C provides risk and is very significantly associated in South Indian CRC patients 25 as well as breast cancer 35 and found in higher frequency among melanoma patients in Europeans, 36 and the T-allele correlates with shorter life expectancy among prostate cancer patients. 37 The frequency of minor allele (C) among North Indian CRC patients and healthy subjects (17.18% in CRC patients and 8.77% in controls) matched exactly with the frequency reported in South Indian population (18.39% in CRC patients and 8.82% in controls). 25 Our data of North Indian subjects showed that minor allele provides a risk (OR = 2.158) for development of CRC which was consistent with the data published on South Indian population, although the significant value that we observed was borderline non-significant (p = 0.058) which may be due to lower sample size in this study. Since the FPRP values were not significant for CRC risk, the result obtained could be a false negative due to lower sample size. Therefore, we suggest for additional validations with larger sample size to further validate the risk of CRC development due to T16189C polymorphism. Individual analysis of colon and rectal samples revealed a greater minor allele ‘C’ frequency (33.33%) in rectal cancer cases than in colon cancer cases, which provided highly significant risk (p = 0.001, OR = 5.213) for development of rectal cancer. FPRP values for this comparison suggested that the result obtained was authentic and not a false positive. No risk or significance was observed for colon cancer cases. Association studies between mtDNA copy number from cancer and adjacent normal tissues has been described for many cancer types such as breast cancer, cervical cancer, leukemia, and CRCs. 38 The state of cancer cell depending on its needs selects an increased or a decreased copy number of mitochondrial genome. Evidence from multiple studies of mtDNA copy number from PBLs has described association of an increased mtDNA in melanomas, renal cell carcinoma, breast cancer, and prostate cancer18,19,39,40 and decreased mtDNA copy number with risk of cancers such as esophageal adenocarcinoma and soft tissue sarcomas.21,41 It has been reported that increased mtDNA from PBLs of CRC patients shows poor survival of patients. 42 Counter claims where patients with lower mtDNA show low survival have also been reported. 43 In this study, we tried to find out the scenario in peripheral blood of the patients. To the best of our knowledge, the association of peripheral blood mtDNA copy number as a marker and risk of CRC has not yet been ascertained in Indian population.
The comparison of mtDNA copy number revealed that copy number was significantly higher among CRC patients in comparison to healthy controls. It was also significantly higher both in colon cases versus healthy controls and in rectal cases versus healthy controls. As the C-allele is known to reduce the mtDNA copy number, 22 the reduced C-allele frequency and corresponding increased mtDNA copy number among colon cancer samples go hand in hand. But the same was not found true for rectal cancer, thereby suggesting involvement of other variations which might be involved with mtDNA replication. No significant difference was found between different parameters such as smoking, hypertension, and alcohol consumption among colon and rectal cases, thus limiting the involvement of such factors in regulating the mtDNA copy number between the two types. However, these factors showed significant deference in copy number between CRC patients and control in our samples. The factors such as hypertension, smoking, and alcohol consumption have been previously reported to affect the copy number in different studies44,45 and not among others.46,47 Increased copy number in CRC patients (median >56) provided a significantly increased risk (OR: 2.885, p = 0.009) for the development of CRC. Thus, there is a strong risk factor associated between increased mtDNA and development of CRC. The quartile distributions of copy number have further showed that greater than 73 copies of mtDNA have a high risk. Another blood samples related study from CRC patients revealed high ORs for both lower and higher quartiles of mtDNA copy number suggesting a U-shaped relationship, 48 therefore supporting our data in part for increased risk for CRC with increased mtDNA copy number from PBLs. Increased mtDNA may lead to a situation where ROS and other free radicals produced by mitochondria increase which is thought to be involved in carcinogenesis. 49 Mitochondrial ROS-mediated retrograde signaling has been shown to induce pro-cancerous metabolic features. 16 Whereas reduced mtDNA has been shown to promote resistance to apoptosis in cancer cells along with epithelial–mesenchymal transition (EMT).50,51 These alterations in mtDNA copy number (both increased and decreased) may represent diverse mechanisms by which mitochondria and mtDNA copy number play role in different malignancies. 17 As per the finding of this study, it would be appropriate to state that there is definitely some involvement of T16189C polymorphism and mtDNA copy number in providing risk for development of CRC among North Indian population. Interestingly, our data also suggest that C-allele at position 16189 may not be regulating mtDNA copy number alone since we could not find any statistically significant relationship between the two, which may be due to the smaller sample size employed for CNV analysis.
In conclusion, this study is the first report where the relationship between mtDNA copy number from PBLs of CRC and healthy subjects was established. Mitochondrial D-loop variation T16189C and copy number may constitute intrinsic risk factors for CRC development. Although a future study with larger sample size is required to fill in any gap that we faced. We therefore suggest that mtDNA CNV and alteration at 16189 could be utilized for identification of subjects who are at high risk for development or progression of CRC.
Footnotes
Acknowledgements
Zafar Iqbal Bhat acknowledges MANF-UGC for providing fellowship. M Moshahid Alam Rizvi is thankful to UGC for providing funds. Bhupender Kumar acknowledges Taruna Kumari from Department of Statistics, University of Delhi, for valuable help in FPRP analysis.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
