Abstract
This study aimed to explore whether genetic polymorphisms in vitamin D receptor are correlated to the breast cancer prevalence in an Egyptian population. Polymerase chain reaction–restriction fragment polymorphism was used to genotype three frequently analyzed vitamin D receptor gene single-nucleotide polymorphisms (rs1544410, rs7975232, and rs731236) and were identified by sequencing analysis. This is the first study that recorded a new single-nucleotide polymorphism in ApaI genotype within an Egyptian population and was registered with the accession number KY859868. The authors found that TC in rs731236, and TG in KY859868 single-nucleotide polymorphism showed significant distribution differences with an increased risk of breast cancer (p < 0.05, odds ratio = 3.71, 95% confidence interval: 1.04–13.28 and p < 0.001, odds ratio = 7.05, 95% confidence interval: 2.02–24, respectively) compared with the wild-type TT genotype carriers in both single-nucleotide polymorphisms. In addition, the distribution frequencies of haplotypes ACT, GTT, and ATT in the patients group were significant, where ATT haplotype was associated with the highest breast cancer risk among all other haplotypes in the patients group (p = 0.0023, odds ratio = 1.72, 95% confidence interval: 1.24–2.437). In conclusion, vitamin D receptors ApaI and TaqI confer high breast cancer susceptibility, particularly in Egyptians females carrying haplotype ATT. However, further studies focusing on the vitamin D receptor variants and haplotypes effects on vitamin D and vitamin D receptor concentrations, activities, and functionalities are needed.
Keywords
Introduction
Breast cancer is one of the most investigated invasive malignancies and the second most known deadly disease for women around the world.1–3 A total of 80% and 24% of breast cancer patients above 50 and 75 years, respectively, were diagnosed in the United Kingdom between the years 2009 and 2011, based on the breast cancer incidence statistics (2013). 4 The most current statistical study which divided Egypt geographically into three sections—Lower Egypt: Behira, Gharbiah, Cairo, and Damietta; Middle Egypt: Minya Assiut; and Upper Egypt: Aswan—reported that women with breast cancer showed the highest frequencies (33.8%, 26.8%, and 38.7%, respectively) among all cancers. 5 The invasive properties of breast cancer in Egyptians are strongly associated with low survival rate and may exhibit specific “molecular feature that might be attributed to some genetic factors and environmental exposure.” 6
However, molecular mechanisms of breast cancer pathogenesis are still not clearly identified. Vitamin D participates in several biological processes including bone metabolism, regulation of calcium and phosphorus metabolism in bones and intestines, and modulation of immune response. 7 Moreover, it is involved in the regulation of cell proliferation and differentiation, as well as in the regulation of apoptosis,2,8 and the alteration of cancer risk, through binding to vitamin D receptors (VDR).9–12
In the liver, vitamin D from all sources undertakes hydroxylation and converts to 25-hydroxyvitamin D (25(OH)D, calcidiol), which is further hydroxylated, via 25-hydroxyvitamin D1a-hydroxylase in the kidneys and other tissues, to the most active form of vitamin D, 1,25-dihydroxyvitamin D (1,25(OH)2D3, Calcitriol).13,14 Calcitriol is expressed in most normal and cancerous cells. In cancer cells, it may bind to VDR and regulates cell development or progression, 15 enhance cell differentiation, and can suppress cell proliferation.16,17
VDR is an intracellular hormone receptor that binds to calcitriol specifically and interacts with specific nucleotide sequences of some specific genes to yield multiple biological effects. Laboratory investigations have suggested that VDR could play vital roles in breast cancer prevalence,7,18 and their low expression might be associated with an increased breast cancer susceptibility.19–21 This hypothesis was predicted from studies which have indicated the association of an increased risk of breast cancer with the low level of calcitriol in serum. 22 Moreover, several studies proposed that metabolite concentrations of vitamin D in serum were associated in reverse with the risk of breast cancer progression.23,24 It is well recognized that vitamin D effects are correlated with the nuclear VDR. 25 In addition, the alterations in expression and activity of VDR could cause downregulation of vitamin D uptake, metabolism, and decrease the serum levels of biologically active vitamin D.
VDR gene is found on chromosome 12q13 and includes more than 470 single-nucleotide polymorphisms (SNPs) that may affect the structure and function of VDR. Therefore, the relation between VDR gene variants and breast cancer susceptibility has attracted a huge attention in the last decade. The most studied SNPs were VDR FokI (rs2228570), VDR BsmI (rs1544410), VDR TaqI (rs731236), VDR ApaI (rs7975232), and VDR Poly A (rs17878969). 26 VDR BsmI, VDR TaqI, and VDR ApaI are all found near the 3’ untranslated region (UTR), where VDR BsmI and VDR ApaI are substitutions in intron 8, include G→ to A replacement for BsmI, and C→ to A replacement for VDR ApaI, whereas VDR TaqI reveals substitution of T→ by C in exon 9, which results in a synonymous change at codon 352. 27
The efficient role of VDR BsmI, TaqI, and ApaI is still mysterious. However, studies do propose that these polymorphisms may affect messenger RNA (mRNA) stability through the alteration of VDR mRNA polyadenylation. 28 Many reports have indicated the presence of an association between BsmI, TaqI, and ApaI polymorphisms and breast cancer risk,29–32 while others found negative associations.33–35 Thus, up till now, there is still a lot of arguments about this relationship. This study was proposed to estimate the role of the well characterized VDR gene polymorphisms, BsmI (rs1544410), ApaI (rs7975232), and TaqI (rs731236) in the pathogenesis of breast cancer in Egyptian women.
Materials and methods
Materials
All chemicals or enzymes used were with high purity or molecular biology gradient and were purchased from ThermoFisher scientific and Promega (Cairo, Egypt). All reagents were used according to the necessary health and safety procedures. All molecular kits are listed elsewhere.
Study subjects
A total of 100 cases of Egyptian females with breast cancer were collected by the Cancer Institute in Cairo from March 2013 to October 2014 and confirmed by pathology. Three cases were excluded from the study due to unavailable clinical data. A written informed consent was obtained from each participant. Their ages ranged from 40 to 73 years, with an average age of 53.2 years. A total of 50 cases of apparently healthy female’s volunteers with ages ranging from 35 to 75 years, with no clinical symptoms, no tumors, and no history of genetic diseases were used as a control group. The diagnosis of breast cancer was based on the history taking, clinical examination of breast and axilla, pathological diagnosis (fine needle aspiration cytology (FNAC) and true cut needle biopsy of breast mass), and estrogen and progesterone receptor (ER/PR) evaluation. Our study was approved by the Medical Ethics Committee of National Research Centre, and all study subjects were informed and agreed to participate.
Sample collection
A total of 3 mL of the peripheral venous blood from the control female samples was collected in a sterile ethylenediaminetetraacetic acid (EDTA) tube for DNA extraction. Tumor samples were collected from breast tissues of females with breast cancer and they were stored at −80°C for further analysis.
DNA extraction and genotyping
Genomic DNA was extracted from blood and tissue samples using DNA mini kits (ThermoFisher Scientific, K0781 and K0721) according to the manufacturer’s instructions. DNA concentration and purity was measured using Q5000 UV spectrophotometer (Quawell, USA) and DNA samples were stored at −20°C till the time of use.
VDR TaqI, ApaI, and BsmI genotypes were analyzed using polymerase chain reaction–restriction fragment polymorphism (PCR-RFLP). Regions of VDR gene flanking the studied SNPs were amplified by PCR using primers previously published36,37 and listed in Table 1.
The primer sequences for VDRs TaqI, ApaI, and BsmI.

VDR: vitamin D receptor; SNP: single-nucleotide polymorphism.
The amplification was accomplished in a 50-μL reaction mixture containing 2 μL of 50 ng template DNA, 1 μL of each primer (10 pmol), and 25 μL master mix Taq polymerase (Promega). The amplification was performed under the following conditions: initial denaturation at 94°C for 3 min (for TaqI and ApaI) and 5 min (for BsmI), then denaturation at 94°C for 30 s, followed by annealing at 60°C (for TaqI and ApaI) and 58°C (for BsmI) for 30 s, 72°C for 30 s extension for 30 cycles (for TaqI and ApaI) and 35 cycles (for BsmI), then final elongation at 72°C for 5 min (for TaqI and ApaI) and 8 min (for BsmI), and final hold at 4°C. The PCR product was digested with TaqI, ApaI, and BsmI (Thermo Scientific FastDigest FD0674, Thermo Scientific FastDigest FD1414, and Thermo Scientific FastDigest Mva 12691, respectively). To avoid star activity or inefficient digestion, the PCR concentration has been measured before starting the digestion and adjusted to 0.2 µg. The following mixture (1 µl restriction enzyme + 34 µl nuclease free water + 10 µl PCR product (0.2 µg) + 5 µl 10× buffer) was incubated at 37°C for 20 min. The PCR products were visualized using 2% agarose gel containing ethidium bromide under ultraviolet transillumination. The different sizes of DNA fragments were as follows: T allele (352 bp), C allele (292–59 bp) for VDR TaqI, T allele (352 bp), G allele (216–135 bp) for VDR ApaI, A allele (358 bp), and G allele (191,166 bp) for VDR BsmI. To confirm the accuracy of genotyping, partial samples representing each genotype were examined by DNA sequencing.
Sequencing analysis
For all SNPs, samples representing homozygous and heterozygous alleles for each genotype have been sequenced. Their PCR products were electrophoresed on 2% agarose gel and purified using the QIAquick® Gel Extraction Kit (QIAGEN, USA), according to the manufacturer’s recommendations. The samples were sequenced by the dideoxy chain termination method, using the original sets of primers. DNA sequencing was performed on an ABI Prism 3100 Genetic Analyzer (Applied Biosystems, USA), using ABI PRISM®BigDye™ Terminator v3.0 Ready Reaction Cycle Sequencing Kit (Applied Biosystems).
Statistical analysis
The results were statistically analyzed using Statistical Analyses Systems (SAS) (2009). SAS Program v9.2 (SAS Institute Inc., USA). The χ2 test and the Student’s t-test were used to analyze clinical data between groups. Hardy–Weinberg equilibrium was tested with a goodness of fit χ2-test. We compared genotype and allele frequencies of TaqI, ApaI, and BsmI in VDR gene between the two groups using the χ2 test and Fisher’s exact test when appropriate, and calculated odds ratio (OR) and 95% confidence intervals (CI) to assess the relative risk. p < 0.05 was considered significant.
Results
Genotypes and clinical data
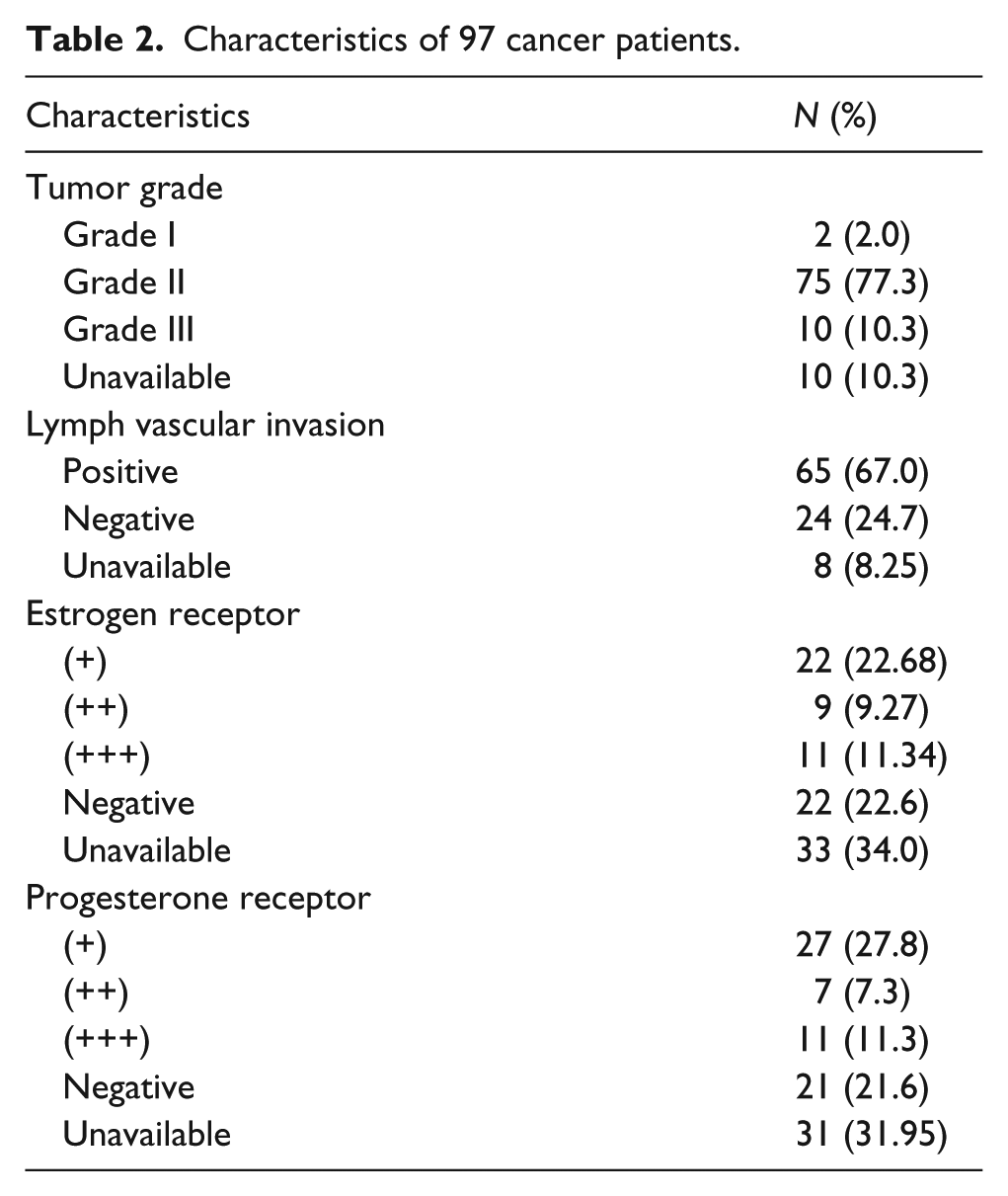
The number of patients for each clinical parameter has been indicated in Table 2. We investigated whether any of the VDR genotypes of the studies SNPs were associated with particular clinical/pathological characteristics of our breast cancer group. There was no association between any VDR genotypes and either ER or PR expression of the tumor, stages of tumor, or lymph node invasiveness (Table 3).
Characteristics of 97 cancer patients.
Association of VDR genotypes with clinical data.

VDR: vitamin D receptor; n.s.: not significant with p > 0.05.
Distributions of different genotypes
Genotypes for the three SNPs in the fragments digested by TaqI, ApaI, and BsmI were indicated C, G, and G alleles, respectively, in the presence of restriction site, while the T, T, and A alleles, respectively, denote lacking of restriction site, according to the sequencing data of our study (Figures 1–3). The new SNP discovered in this study demonstrates the replacement of the second nucleotide G→T within ApaI restriction site GG^GCCC, as indicated in our sequence deposited in the nucleotide database with the accession number KY859868, instead of replacement of the fifth nucleotide C to A as previously reported (rs7975232) in the other research studies.21,37
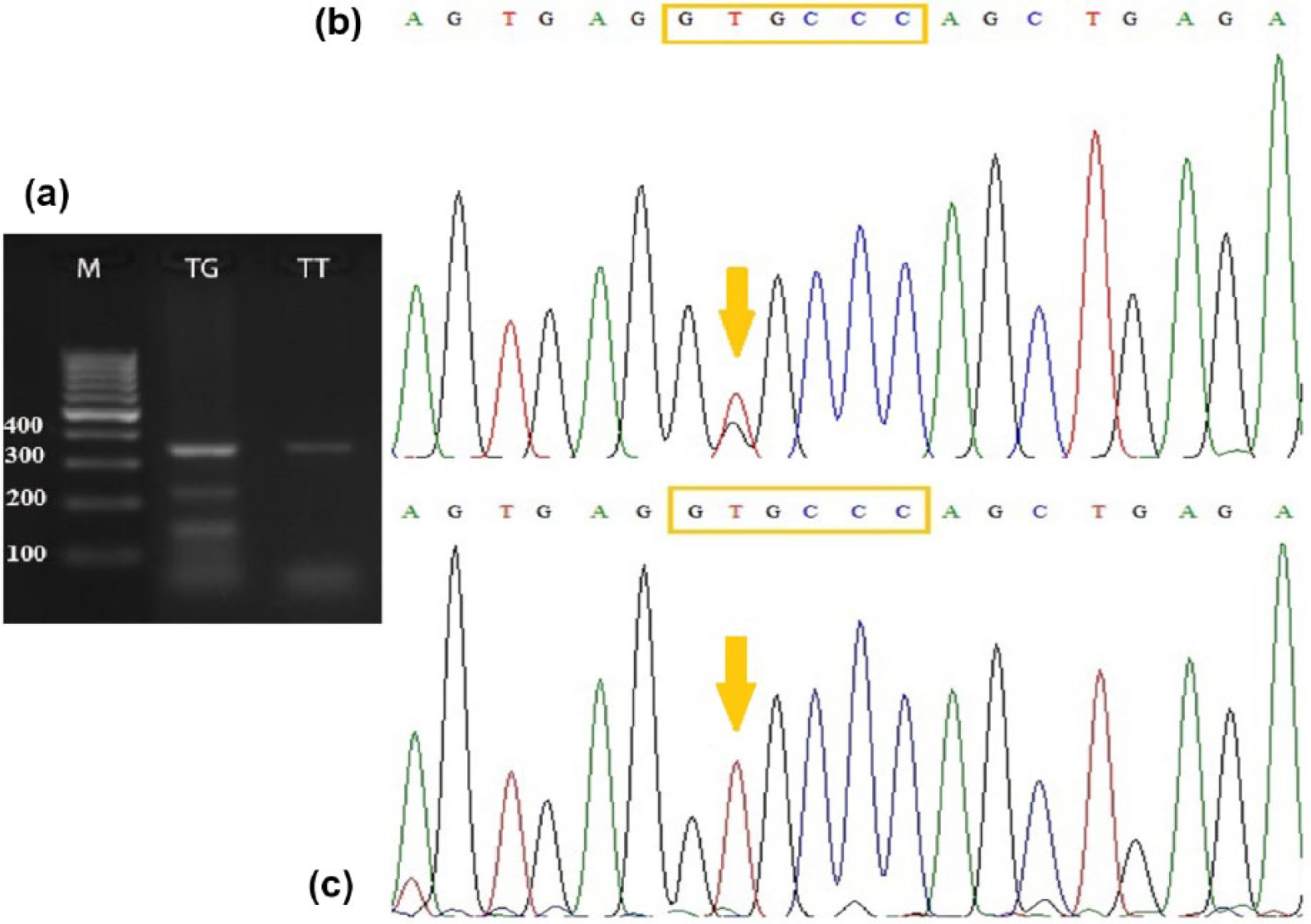
Genotyping of ApaI fragment of VDR gene. (a) The electrophoretic pattern obtained after digestion of PCR amplified part of VDR gene using ApaI enzyme showing heterozygous genotype TG (lane 1) and homozygous genotype TT (lane 2), where T allele appears at (352 bp) and G allele appears at (216–135) bp. M: 100 bp ladder marker. (b) DNA sequencing analysis representing TG genotype. (c) DNA sequencing analysis representing TT genotype.

Genotyping of TaqI fragment of VDR gene. (a) The electrophoretic pattern obtained after digestion of PCR amplified part of VDR gene using enzyme showing heterozygous genotype TC (lane 1) and homozygous genotype TT (lane 2), where T allele appears at 352 bp and C allele appears at 292–59 bp. M: 100 bp ladder marker. (b) DNA sequencing analysis representing TT genotype. (c) DNA sequencing analysis representing TC genotype.

Genotyping of BsmI fragment of VDR gene. (a) The electrophoretic pattern obtained after digestion of PCR amplified part of VDR gene using BsmI enzyme showing homozygous genotype AA and GG (lanes 1 and 3, respectively), and heterozygous genotype AG (lane 2), where A allele appears at 358 bp, and G allele appears at 191,166 bp. M: 100 bp ladder marker. (b) DNA sequencing analysis representing AA genotype. (c) DNA sequencing analysis representing AG genotype. (d) DNA sequencing analysis representing GG genotype.
The frequency of VDR polymorphisms in our sample populations is shown in Table 4. We found significant differences in two genotype frequencies between patients and controls (rs731236 and rs7975232 SNPs) which were associated with breast cancer risk. Compared to homozygous wild-type TT in both SNPs,38–40 the distribution differences of TC in rs731236 and TG in rs7975232 within the patients group and the control group were significant (p < 0.05 and p < 0.001, respectively) and after adjustment of other confounders, the difference remains significant (OR = 3.71, 95% CI: 1.04–13.28 and OR = 7.05, 95% CI: 2.02–24, respectively). There was significant deviation from the expected Hardy–Weinberg frequencies (p < 0.0001) for the three tested genotypes suggesting that collected samples don’t represent the group.
Distributions of three SNPs of VDR gene within breast cancer patients and control subjects.

CI: confidence interval; OR: odds ratio; SNP: single-nucleotide polymorphism; VDR: vitamin D receptor.
Significant results (p < 0.05).
Significant results (p < 0.001).
VDR gene haplotype analysis
STAT software showed that SNPs of VDR genes rs1544410, KY859868, and rs731236 had strong linkage disequilibrium. After population analysis, it was found that A-T-T is the most frequent haplotype found in both control and cases groups (13.59% and 16.76%, respectively), followed by G-T-T (13.07% and 14.62%, respectively), then equally A-C-T or A-T-G in control groups (12.77%), and only A-C-T in cases (14.23%). Meanwhile, G-C-G is found to be the least frequent haplotype among all tested groups (Table 5). Our study showed that the distribution frequencies of haplotypes A-C-T, A-T-T, and G-T-T in the patient group (14.23%, 16.76%, and 14.62%, respectively) were significantly higher (p < 0.05) than those in the control group (12.77%, 13.59%, and 13.07%, respectively). A-T-T haplotype resulted in a higher breast cancer risk (OR = 1.72, 95% CI: 1.24–2.437).
Haplotype distribution between patients and control subjects, n (%).

OR: odds ratio; CI: confidence interval.
Significant results (p < 0.05).
Discussion
The pathogenesis of breast cancer is not established yet, as it involves various genetic and environmental factors as well as several molecular signaling pathways. 41 A meta-analysis that included 15 epidemiological studies has verified an inversely statistically significant association between serum 25(OH)D concentration (which is affected mainly by VDR gene polymorphisms) and breast cancer risk in some geographical regions (North America), while this association was absent in some other regions (Europe). Results of ethnic distributions were in accordance with geographic distributions. This indicates a possible role of ethnic differences in genetic profile variation together with other factors such as people’s living environment and dietary habits that may be contributed as well. Thus, breast cancer risk can be changed according to the ethnicity which were categorized into Caucasian, Asian, African and American. Herein, we conducted our study in African-Egyptian population.
Interestingly, vitamin D does not appear to be functional by its own, but it is the resource of the vital hormone, 1,25-dihydroxy vitamin D (1,25(OH)2D), 42 that initiates growth arrest, activates cell death, and indorses differentiation of cancer cells counting breast cancer cells. 43 Hence, vitamin D may act as a protective agent in breast cancer risk. Furthermore, the active form of vitamin D, 1,25-dihydroxyvitamin D3 (1,25(OH)2D3), is a powerful inhibitor of a diversity of cancer cell categories especially breast cancer. 44 However, the exact mechanisms by which 1,25(OH)2D3 exerts its anti-proliferative effects are still not clear.
VDR mediates the regulatory effects of 1,25(OH)2D3 and acts as its ligand. 45 Furthermore, VDR function in cancer cells is important for 1,25(OH)2D sensitivity, so corruption of VDR function during carcinogenesis leads to loss of the anti-proliferative effects of 1,25D. In addition, VDRs are responsible for the transcriptional regulation of a number of hormone-responsive genes and thus effects breast cancer which is known to be strongly influenced by the hormonal milieu and variation in genes that are responsive to such hormones. 46
This study concerns about the most common VDR SNPs: BsmI (rs1544410), TaqI (rs731236), and ApaI (accession number KY859868). While VDR BsmI and ApaI polymorphisms are intronic, VDR TaqI leads to a silent codon change, meaning they do not change the transcriptional mRNA or the translated protein; instead, they are thought to be important in post-transcriptional regulation of gene expression. 47 From the previous studies, there seems to be a strong linkage disequilibrium between the three SNPs BsmI, ApaI, TaqI, and the variable length poly(A) in Caucasian populations.38,48
Over the last two decades, large number of population-based studies were conducted to assess the association of variations in the VDR gene SNPs and the breast cancer prevalence; however, the data are controversial. In this study, we found that TT genotype in ApaI (accession number KY859868) was related to breast cancer pathogenesis in an Egyptian population, whereas no GG genotype in our population sample could be detected which may be due to the limited number of samples. In accordance with our findings, Dalessandri et al. 49 observed an increased risk to breast cancer among women carrying VDR ApaI (TT) genotype within Marin County population, whereas Curran et al. 38 found a marked increased susceptibility of breast cancer with the other two genotypes of VDR ApaI (GT and GG). Moreover, many recent reports found a marked increased susceptibility to breast cancer between women carrying ApaI GG genotype compared with women carrying genotype TT.4,9,37,50
On the contrary, McCullough et al. and Mishra et al. reported no significant relation between the ApaI variants and breast cancer prevalence among Indian and Caucasian women.32,49,51–53
Interestingly, our findings indicated the existence of marked difference between patients and control samples of women carrying TC genotype of VDR TaqI (rs731236). A previous study revealed that death of patients carrying CC genotype (22%) is considerably lower than the TT genotype (41%) or the TC genotype (44%) patients, and the TT genotype patients have a marked higher risk for lymph nodes metastasis than patients carrying the C allele suggesting a strong indication for breast cancer progression. 48 Moreover, VDR TaqI polymorphism was associated with a significantly increased breast cancer risk in estrogen receptor positive patients.54,55 Similarly, an elevated but not significant susceptibility of breast cancer was reported in Asians heterozygous and homozygous for C allele of VDR TaqI, as well as in Caucasians.38,54 This is different from studies that indicated no association between VDR TaqI (rs731236) polymorphisms and susceptibility of breast cancer in Egyptian women.32,51,56,57
Our study resulted in no association between breast cancer and VDR BsmI; this result is consistent with the previous reports.21,57,58 On the contrary, many studies indicated an association between breast cancer risk and VDR BsmI polymorphism bb genotype.9,46,59
Anyway, there was a strong linkage disequilibrium between tested SNPs indicating that A-T-T haplotype conferred a higher breast cancer risk. Moreover, our data indicated no association between the tested VDR genotypes and ER status of women in our group. In contrast, Lundin et al. found a tendency toward increased survival in ER-positive, premenopausal women with tt genotype. However, the relationship between VDR variants and breast cancer risk may be very different in pre- and post-menopausal women. 48 The deviation from Hardy–Weinberg equilibrium that we observed for the tested polymorphism could be contributed to undetected ethnic diversity, biologic clarifications, or genotyping misclassifications.
In conclusion, while vitamin D and its analogues are being established as preventative and/or treatment agents in breast cancer, the assessment of VDR polymorphisms is considered as a vital step in the identification of at-risk groups and strategies for targeting and intervention. This study has introduced an additional evidence for a significant association between VDRs ApaI and TaqI variants with the prevalence of breast cancer. However, as in all research, our study has limitations, such as small sample size, insufficient analyzed loci. Subsequent high-quality epidemiological studies are encouraged to clarify the results identified in this study.
Footnotes
Acknowledgements
The authors thank all patients and controls who agreed to join in our study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Department of Zoology, Faculty of Science, Cairo University.
