Abstract
Cholangiocarcinoma is a rare type of cancer which is an increasingly discernible health threat. The disease is usually very difficult in diagnosis and various treatment modalities are typically not effective. Cholangiocarcinoma is a complex and very heterogeneous malignancy characterized by tumor location, different risk factors, molecular profiling, and prognosis. Cancer cell lines represent an important tool for investigation in various aspects of tumor biology and molecular therapeutics. We established two cell lines, KKU-452 and KKU-023, which were derived from patients residing in the endemic area of liver fluke infection in Thailand. Both of tumor tissues have gross pathology of perihilar and intrahepatic mass-forming cholangiocarcinoma. Two cell lines were characterized for their biological, molecular and genetic properties. KKU-452 and KKU-023 cells are both adherent cells with epithelium morphology, but have some differences in their growth pattern (a doubling time of 17.9 vs 34.8 h, respectively) and the expression of epithelial bile duct markers, CK7 and CK19. Cytogenetic analysis of KKU-452 and KKU-023 cells revealed their highly complex karyotypes; hypertriploid and hypotetraploid, respectively, with multiple chromosomal aberrations. Both cell lines showed mutations in p53 but not in KRAS. KKU-452 showed a very rapid migration and invasion properties in concert with low expression of E-cadherin and high expression of N-cadherin, whereas KKU-023 showed opposite characters. KKU-023, but not KKU-452, showed in vivo tumorigenicity in xenografted nude mice. Those two established cholangiocarcinoma cell lines with unique characters may be valuable for better understanding the process of carcinogenesis and developing new therapeutics for the patients
Introduction
Cholangiocarcinoma (CCA) is a rare type of cancer worldwide. However, incidences of intrahepatic CCA are increasing in many parts of the world. 1 In particular, some regions like Northeast Thailand, Laos, Cambodia, Southern China and Korea where liver fluke (Opisthorchis viverrini (Ov) and Clonorchis sinensis) infection is endemic, high to very high incidence of CCA (7 to 85 /100,000) has been observed.1,2 Prognosis of the disease is dismal due to difficulties in diagnosis and poor response to drug therapy.1,3 CCA is a group of heterogeneous malignant tumors. The classification of CCA is based on anatomical location including intrahepatic and extrahepatic CCA, where extrahepatic CCA is subdivided into perihilar and distal CCA. 4 The frequent forms of intrahepatic CCA in Thailand are intraductal growth, periductal infiltrating and mass-forming types, while the vast majority of perihilar and distal CCA are mucinous adenocarcinoma.1,5
CCA has been suggested to be stemmed from diverse cell origins, such as hepatic stem cells, various stages of cholangiocytes and peribiliary gland cells.6,7 Intrahepatic CCA probably arises from hepatic progenitor cells, while perihilar CCA may be originated from mucin-producing biliary epithelium or peribiliary glands located in hilar bile ducts.1,6 Histological and phenotypic subtypes of the tumor seem to be associated with tumor aggression and prognosis, where papillary and gastric or intestinal phenotype were less aggressive than those of tubular and pancreatobiliary phenotype.8,9 Recent investigations over genomic and epigenetic alterations in CCA have revealed that genomic heterogeneity is not only related to tumor location but also to risk factors associated to carcinogenesis of CCA. 1 It is noted that mutation profile of CCA is different between tumors from liver fluke Ov-associated and non-Ov associated patients, in such that frequencies of mutation in TP53 and SMAD4 are high in Ov-associated CCA, while mutation of BAP1 and IDH1/2 is high in non-Ov-associated CCA. KRAS is a common mutation in both groups.10,11 Furthermore, in CCA within the same background risk, mutational profiles may be different depending on the tumor location. For instances, among Ov-CCA, intrahepatic CCA has shown high frequency of mutations in KRAS and ARID1A than extrahepatic CCA; however, among non-OV CCA, intrahepatic CCA has shown high frequency of mutations in BAP1 and IDH1/2 than extra-hepatic CCA.10,11 These represent a pronounced heterogeneity of malignancies probably resulting in variability of prognosis and resistance to chemotherapy.
The rising incidence of CCA, very poor prognosis of the disease, and lack of effective treatment prompt researchers to investigate molecular pathogenesis. Cancer cell lines represent an important tool for investigation in various aspects including tumor biology and putative molecular therapeutics. Several CCA cell lines have been established from Ov-associated and non-Ov-associated CCA, where tumor tissues were from various gross pathological and histological subtypes. Some representative cell lines of Ov-associated CCA are KKU-100, HuCCA-1, and RMCCA-112–14 and of non-OV-associated CCA cells are KMBC, MT-CH01, SNU-245, SNU-308, and so on.15–17 Cancer cell lines are simple models, representing some characteristics of their originated tumors. It is, therefore, very useful to employ several lines of cells in studies on some common as well as distinct molecular and phenotypic characteristics for better understanding of molecular basis of cancer development and for the advancement of the design of an effective therapeutics. CCA cell lines have been employed in the study for the role of putative target enzymes or transcription proteins, including Nrf2, 18 NF-κB, 19 HO-1, 20 NQO1,21,22 and PI3K/Akt, 23 or for investigation of pathways involving cancer invasion and metastasis24–26 and drug therapy.27–29
As CCA is a very heterogeneous malignancy and there are very few cancer cell lines with extensive characterization originated from Ov-associated CCA widely available, new cell lines with different origins may be very useful for study insight of cancer. We established two CCA cell lines from patients who resided in northeast Thailand, where are the endemic area of liver fluke infections. The each cell line was derived from extrahepatic and intraheptic CCA with different histopathology. Those two cell lines showed different characteristics in immunocytochemistry, growth rate, migration, and proteins that play roles in cancer invasion. KKU-023 cells also showed tumorigenicity in the nude mice model.
Materials and methods
Establishment of tumor cell culture
Cell lines were established from pathologically proven bile duct cancer tissues of two patients who were the residents in the Northeast Thailand. The study protocol has been approved by the Khon Kaen University Ethics Committee for Human Research (HE571283) and experiments were performed in accordance with the Declaration of Helsinki. The written informed consents were obtained from each subject. The surgically resected liver specimens were rinsed several times in Hank’s balanced salt solution. Tumor mass was minced into small pieces, transferred into centrifuge tubes containing 1 mg/mL collagenase, and digested for 30–45 min at 37°C. The cell suspensions were washed by centrifugation and resuspended in Ham’s F12 medium (Gibco BRL, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (FBS), antibiotic mixture (cefazolin 50 µg/mL, ciprofloxacin 10 µg/mL, and amphotericin B 2.5 µg/mL), Minimum Essential Medium (MEM) non-essential amino acid (Gibco BRL), 12.5 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES), and 5 µM ROCK inhibitor, Y-27632 (Santa Cruz Biotechnology, Santa Cruz, CA, USA). Cell suspensions were filtered through a cell restrainer with 70 µm pore mesh and cultured in CO2 incubation at 37°C. Doubling time of cultured cells was estimated during cell growth in an exponential phase. Cells were seeded at varying density in six-well plate and cultured for a certain period of time, and then, cells were trypsinized and counted the number. Primary cultured cells were observed periodically and contaminating fibroblasts were aseptically scraped off until CCA cells were free from fibroblasts. Cultured cells were initially subcultured every week until they grew at a stable rate and then subcultured when they reached 50%–70% confluence using trypsin–ethylenediaminetetraacetic acid (EDTA). Y-27632 was removed from full-culture media after few passages of primary cultures. The authentication of the cell lines was performed using AmpFLSTR® Identifiler® PCR Amplification Kit (Applied Biosystems, Foster City, CA, USA) and comparing their short tandem repeat (STR) profiles with the ICLAC Database. 30 Contamination with mycoplasma has been monitored periodically using a test kit of MycoAlert Plus Mycoplasma® (Lonza, Walkersville, MD, USA). The cultures were always free from mycoplasma during this series of experiments.
Cell migration and invasion assays
CCA cells were cultured in full media overnight. Scratch wounds were made with a sterile 200 µL pipette tip. After removal of any detached cells by washing with phosphate-buffered saline (PBS), images of scratched wound were recorded at time 0 and 24 h. The closing of wound gap representing cell migration was calculated using Image-Pro Plus software (Media Cybernetics, LP, Rockville, MD, USA). Cell invasion assay was performed by Transwell chamber assay using a 24-well plate. The Transwell inserts with pore size of 8 µm were coated with Matrigel (BD Bioscience, Bedford, MA, USA) overnight. The KKU-452 and KKU-023 cells in serum-free medium were seeded at density of 20,000 cells/well onto the precoated inserts, while the lower part of the chamber was filled with culture medium containing 10% FBS and MEM non-essential amino acids. After incubation for 24 h, the cells on the upper surface were gently removed with a cotton swab. The invaded cells were fixed with absolute methanol, stained with crystal violet, and then photographed. Area of cell invasion was assessed using Image-Pro Plus software.
Immunocytochemistry
CCA cells were cultured in a chamber slide for 24 h, washed with PBS, fixed with cold methanol for 20 min, and air-dried before subjected to immunostaining with mouse monoclonal antibodies against cytokeratin, CK-7, and CK-19 (Cell Marque; Sigma-Aldrich, St. Louis, MO, USA) using the standard horseradish peroxidase (HRP) method. CK7 and CK19 staining was assessed according to the intensity from no staining = 0 and increase in intensity from +, weak staining; ++, moderate staining; and +++, strong staining in the whole cytoplasm.
Western blotting
CCA cells were harvested and lysed with cell lysis buffer. The protein concentration was determined by the Bradford dye binding method. The protein was resolved in sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) using 10% polyacrylamide gel and transferred onto a polyvinylidene difluoride (PVDF) membrane. The protein bands for E-cadherin, N-cadherin, matrix metalloproteinase (MMP)-9, vascular endothelial growth factor (VEGF)-C, p53, and β-actin were detected using specific primary antibodies (Santa Cruz Biotechnology) followed by the appropriate HRP-conjugated secondary antibodies and Amersham™ ECL™ Prime detection reagent.
Chromosome analysis
CCA cells were subjected to genetic analysis when cell lines have become stable in culture. KKU-452 cells and KKU-023 cells of 50 and 40 passages, respectively, were used. The established cells were treated with 1 µg/mL colcemid. The cells were treated with a KCl hypotonic solution and harvested according to the standard cytogenetic procedures. Slides of fixed cells were trypsin-Giemsa-banded to identify individual metaphase chromosomes. Representative chromosome sets were photographed for karyotype analysis. The karyotypes of CCA cells were analyzed by two cytogeneticists (S.L. and Y.C.) with Ikaros MetaSystems (MetaSystems Group, Newton, MA, USA). The modal chromosome number was determined from 25 cells.
Sanger sequencing for TP53 and KRAS mutations
To explore the mutation of TP53 and KRAS in newly established cell lines, KKU-100 was used as a positive control with known mutations. For KRAS (NM_004985) mutation on exon 2, 100 ng of genomic DNA (gDNA) was used as templates for the polymerase chain reaction (PCR) reactions. For the detection of TP53 (NM_000546) mutation in the coding region, RNA of three cell lines was collected for preparation of complementary DNA (cDNA), which were used as PCR templates. The cDNA was synthesized using iScript™ Reverse Transcription Supermix for reverse transcription quantitative polymerase chain reaction (RT-qPCR) according to the manufacturer’s instructions (Bio-Rad, Hercules, CA, USA) and 40 ng of cDNA was used as PCR template. PCR was performed using specific primers (Supplementary Table S1) with Platinum Tag DNA Polymerase system (Invitrogen, Carlsbad, CA, USA) on the T100™ Thermal Cycler (Bio-Rad, Hercules, CA, USA). PCR cycling parameters included 1 cycle at 94°C for 2 min, 35 cycles at 94°C for 30 s, 50°C–58°C for 30 s, 72°C for 45 s, and 1 cycle at 72°C for 5 min. The PCR products were further analyzed using agarose gel electrophoresis.
For Sanger sequencing, PCR products were cleaned up by Wizard® SV Gel and PCR Clean-Up System (Promega, Fitchburg, WI, USA). The purified PCR products were subjected for nucleotide sequencing by service of the First BASE Laboratories (Selangor, Malaysia). Sequencing traces of amplified products were aligned to reference sequences using Lasergene 10.1 (DNASTAR) and analyzed by visual inspection.
Tumor formation in nude mice
Male BALB/cAJcl-nu/nu mice (6 weeks old) were purchased from CLEA Japan (Tokyo, Japan). Animals were housed under specific pathogen-free conditions at the Northeast Laboratory Animal Center, Khon Kaen University. The animal study protocol was approved by the Animal Ethics Committee, Northeast Laboratory Animal Center of Khon Kaen University (AEKKU-NELAC 7/2558), and all experiments were performed in accordance with the institutional guidelines. Mice were injected subcutaneously with 2–10 × 106 cells/injection for KKU-452 cells or 2–5 × 106 cells/injection for KKU-023 cells at the posterior lateral thoracic wall. The tumor volume was calculated using the formula: ½ × width 2 × length. Mice transplanted with KKU-023 were terminated at day 21. Tumor tissue and the liver, lung, kidneys, and intestine were fixed in buffered formalin for pathological analysis for hematoxylin- and eosin-stained sections. Mice transplanted with KKU-452 cells were observed for 6 weeks, as no tumor mass was apparent at the grafted site. After 6 weeks, animals were sacrificed and the tissue surrounding injection site and major organs were fixed and processed as above.
Results
Establishment of the primary CCA cell lines
Newly established cell lines were derived from tumor tissues of CCA patients, where both patients resided in the endemic area of liver fluke (Ov) infection. The clinical and pathological characteristics of the patients are shown in Table 1. KKU-452 cells were derived from the tumor resected from perihilar area with histology of poorly differentiated CCA, whereas KKU-023 cells were from intrahepatic CCA of mass-forming type with histology of well-differentiated adenocarcinoma. Serum tumor markers of the patient of the origin of KKU-452 were well within normal ranges, whereas those of the patient of the origin of KKU-023 had a marked increase in CA-19-9 level.
Clinical and pathological profiles of patients and the characteristics of the established cell lines.

AFP: alpha-fetoprotein; CEA: carcinoembryonic antigen.
Reference value.
Morphological features of CCA cells
Both CCA cell lines grew in cell culture as adherent monolayer. KKU-452 cells exhibited characteristic spindle shape with multiple processes and seldom multinucleated, while KKU-023 cell showed ovoid to cuboid shape, forming compact monolayer with occasionally multinucleated cells (Figure 1).

Morphology of cultured CCA cells. Typical morphology of stable culture of KKU-452 (left) and KKU-023 (right) at the 50th and 40th passages, respectively.
KKU-452 cells have a doubling time much shorter than KKU-023 (17.9 vs 34.8 h). When KKU-452 cells undergone cell division, daughter cells are segregated and spread surround not grow as a continuous adherent colony. However, KKU-023 cells form adherent compact colonies. The cytokeratins CK-7 and CK-19 are normally expressed in the lining of the gastrointestinal (GI) tract including bile duct epithelial cells and their neoplasm. KKU-452 cells were stained CK-7 with relatively low intensity and very low with CK-19. In contrast, KKU-023 cells were intensely stained with both CK-7 and CK-19 (Figure 2).

Immunocytochemistry of CK-7 and CK-19 in CCA cells. KKU-452 and KKU-023 cells were cultured in slide chambers for 24 h before cells were fixed and stained with CK-7 and CK-19 antibodies.
Karyotypes of CCA cells
KKU452 and KKU023 cells were subjected to karyotype analysis. The characteristics of KKU452 cells were a hypertriploid cell line with the modal chromosome number of 76, range = 59–81, occurring in 25% of cells. Cells with 74 (22%), 59, 79, 80, and 81 chromosome counts also occurred at relatively high frequencies. In all, 10 marker chromosomes, trisomic 3, pentasomic 10, iso(12)(q10q10), and rob(15)(q10q10) are present in all metaphases; nullisomic 13 and monosomic 14 are present in 70% of the karyotyped cells. Chromosomes 11 and 16 are generally trisomic. A half of cells had XX sex chromosome and the rest had XXX (Figure 3) sex chromosome. For KKU023, the analysis revealed that the stem line chromosome number is hypotetraploid with the modal chromosome number of 85, range = 70–92, occurring in 25% of cells. In cells with 88, 90, and 91 (15%), 72 chromosome counts also occurred at relatively high frequencies. Tetrasomy 3, tetrasomy 6, del(1)(p22.3p32), and add(12)(p13.3) are present in all metaphases. Generally, chromosome 18 is monosomic; chromosomes 13, 16, and 19 are tetrasomic; and chromosomes 15 and 17 are hexasomic. The marker chromosomes are seen in 50% of the karyotyped cells. Most cells had XXX chromosomes (Figure 3).

Representative of karyotypes of CCA cells. The karyotype of KKU-452 cells (left) showed 76, XX, +3, +5, +6, +7, +9, +10, +11, +12, +13, −14, −15, +16, +17, −18, +19, −21, +22, +iso(12)(q10q10), +rob(15)(q10q10), +rob(21)(q10q10), and +mar. KKU-023 cells (right) showed 85, XXX, −1, +2, +3, +4, +5, +6, +7, +8, +9, +10, +11, +12, +13, +15, +16, +17, −18, +19, +20, +21, +22, del(1)(p22.3p32), +add(12)(p13.3), and +mar.
Mutational signatures of CCA cells
Recent exome sequencing of the liver fluke–associated CCA patients revealed that TP53 and KRAS are the most frequent mutations. The two established cell lines and reference cell line KKU-100 were subjected for a mutational analysis. KRAS mutation was apparent only in KKU-100 cells with altered codon G12D as a missense mutation (Figure 4). For TP53, both KKU-100 and KKU023 cells exhibited missense mutations at codons P72R in exon 4, while KKU-452 cells showed a missense mutation at codons P72R and nonsense mutation at codon E349•. The expression of p53 protein by western immunoblot was performed. It was revealed that KKU-452 cells exhibited high expression of p53 protein, while KKU-023 and KKU-100 cells showed a very low expression under basal condition (Supplementary Material, Supplementary Figure S1). Data of sequencing of KRAS and TP53 are available in Supplementary data sequencing_KKU-100, 023, and 452.
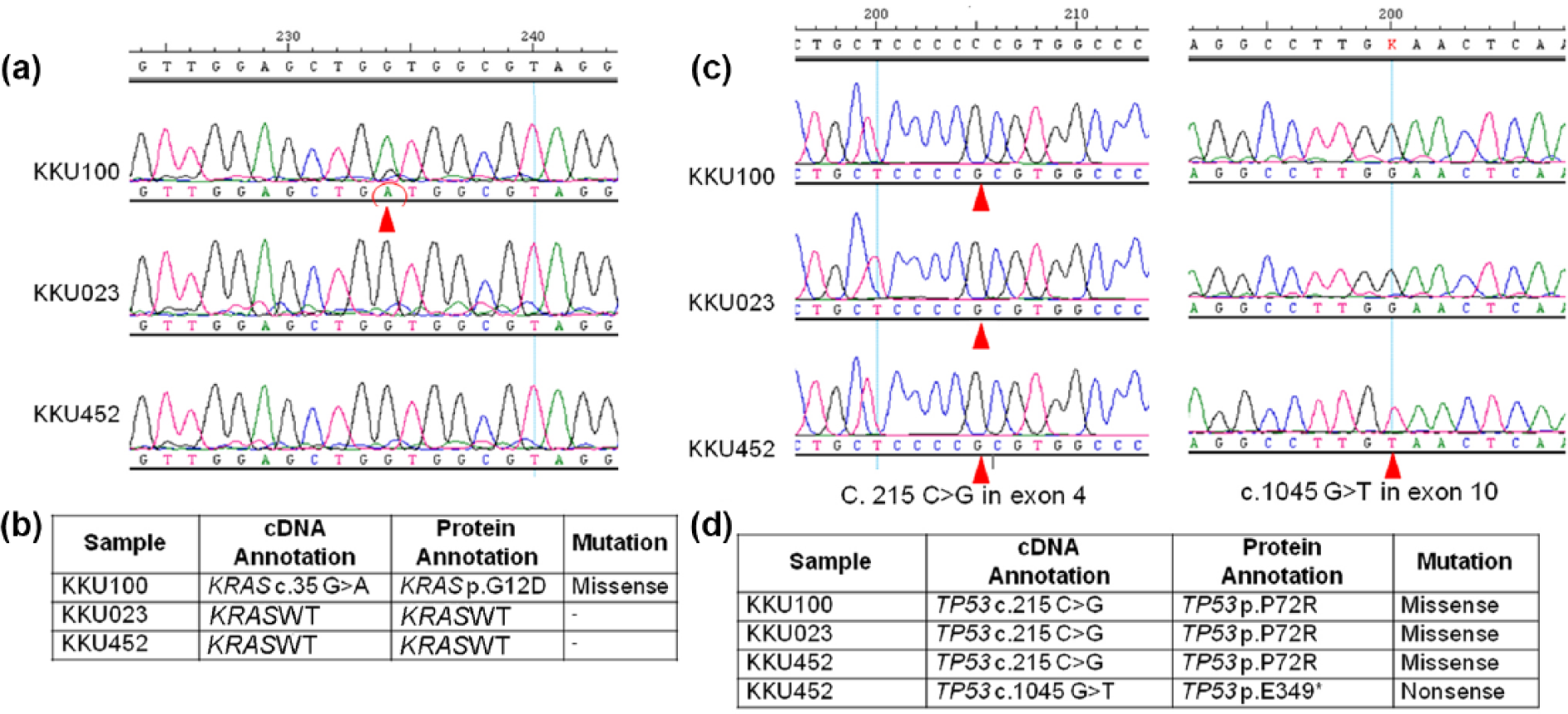
KRAS and TP53 mutations in CCA cells. (a) The representative image of sequencing trace of amplified KRAS in KKU-100, KKU023, and KKU452. The red circle represents point mutation. (c) The representative image of sequencing trace of amplified TP53 in KKU-100, KKU023, and KKU452. (b) The common affected nucleotides of KRAS and (d) entire coding sequence of TP53 were sequenced. Arrow heads represent point mutations in the sequences.
In vitro cell migration and invasion
CCA cells demonstrated a varied ability in migration and invasion. The migration ability was assessed using wound-healing assay and invasion using Transwell invasion assay. KKU-100 cells, a previously established CCA cell line, were employed as reference CCA cells. KKU-452 cells showed a remarkably high migration and invasion properties. The scratch wound on KKU-452 cells was almost completely closed after 24 h incubation, while KKU-023 and KKU-100 cells showed only a modest wound closure at 24 h (Figure 5(a) and (c)). Likewise, KKU-452 cells invaded Transwell chamber aggressively, whereas much less degree of invasion was observed with KKU-100 and KKU-023 cells (Figure 5(b) and (d)).
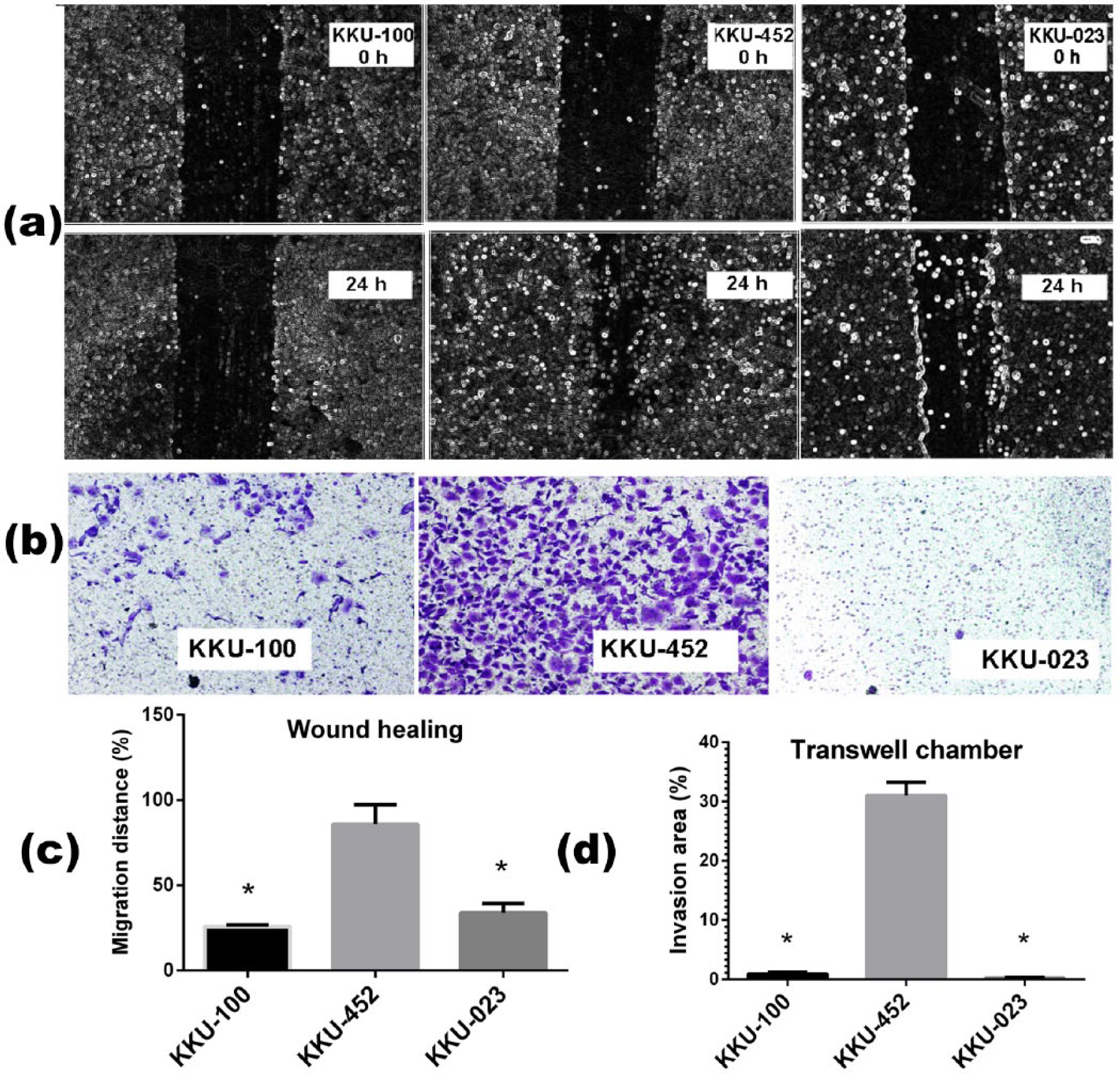
Migration and invasion property of KKU-452 and KKU-023 cells. For cell migration assay, KKU-100, KKU-452, and KKU-023 cells were seeded onto 24-well plates. (a and c) After cells forming confluent monolayer, a scratch line was made and the distance of the scratch wound was measured 24 h after incubation. (b and d) In Transwell invasion assay, CCA cells were seeded onto a Matrigel-coated inserts and incubated for 24 h before the invaded cells through the membrane were stained and enumerated. Each bar was the mean ± standard deviation (SD) from triplicate assays (*significantly different from KKU-452 group, p < 0.05).
Western blotting of proteins involved in cell migration and invasion
The expression levels of proteins involved in cancer cell migration and invasion were evaluated using western blot analysis. For KKU-452 cells, N-cadherin protein expression was relatively high but E-cadherin expression was very low. In case of KKU-100 cells, a reference CCA cells showed a similar protein expression pattern as that of KKU-452 cells. However, KKU-023 cells showed opposite expression pattern (Figure 6) in that there was very high E-cadherin expression and very low N-cadherin expression. MMP-9 protein was expressed in a comparable levels in all three CCA cells. However, VEGF-C expression was relatively high in KKU-452 and KKU-100 cells, while that was low in KKU-023 cells.

Western blot analysis of proteins involved in cell migration and invasion of two new CCA cell lines KKU-452 and KKU-023, with reference to established KKU-100 cell line. Expression of E-cadherin, N-cadherin, MMP-9, and VEGF-C was subjected to western blot analysis, using β-actin as laoding control. Protein bands shown in this figure were the representative from two experiments with similar results.
Tumorigenicity of CCA cells
The tumor forming ability of these established CCA cell lines was examined by in vivo grafting into nude mice. KKU-023 cells transplanted at the density of 2–5 × 106 cells/mice readily formed subcutaneous tumors in all nude mice within a week after transplantation and grew exponentially after the second week (Figure 7(a)). Tumor weight was extracted and measured at the end of study. There was a trend of increased tumor mass when the number of cell transplantation was increased (Figure 7(b)). Surprisingly, KKU-452 cells failed to form tumor mass even the cells were injected at wide range of densities (2, 3, 5, and 10 × 106 cells/mice) and observed up to 6 weeks. Necropsy of the mice xenografted with KKU-452 revealed no malignant cells at the injection site and no evidence of metastasis to other major organs. Histological examination of tumor mass from mice xenografted with KKU-023 cells showed characteristics of tubular adenocarcinoma with epithelial folding foci similar to the original tumor tissue from donor patient (Figure 7(c) and (e)). Moreover, xenografted tissue and donor tissue were markedly stained with CK19, a bile duct biomarker (Figure 7(d) and (f)). There was no evidence of tumor metastasis to nearby lymph nodes or distant organs including lung, liver, kidneys, and spleen of mice xenografted with KKU-023.

Tumor formation in mice and histopathology of CCA tissues from xenograft mice and patients. KKU-023 cells were injected subcutaneously into the shoulder at cell densities of 2 × 106, 3 × 106, and 5 × 106 cells (n = 3/group). (a) Tumor mass was formed within a week and tumor size was measured periodically. (b) On day 21, tumors were extracted, weighted and subjected to histological examination. Each bar represents the mean ± SD (n = 3). Micrographs of CCA from (c and d) xenograft mice and (e and f) tumor tissue of patient donor demonstrated tubular adenocarcinoma in (c and e) H&E staining and staining of CK19 (d and f), bile duct marker (10 × 20 magnification).
Discussion
CCA is a complex and very heterogeneous malignancy characterized by different risk factors, tumor locations, molecular profiles, and different prognosis. 1 The prognosis of the disease is generally very poor owing to asymptomatic or nonspecific symptoms in nature until disease progressing to an advanced stage and ineffective drug or radiotherapy.1,3 Cancer cell line is a preclinical model for study of various aspects of carcinogenesis and drug discovery. As there are very limited number of CCA cell lines available, we sought two lines of cells, KKU-452 and KKU023, established from extra-hepatic and intra-hepatic CCA with histological grade of poorly and well-differentiated adenocarcinoma, respectively. We characterized two cell types for their biological, molecular, and genetic properties. Two cell types exhibited different characteristics of proliferation, immunocytochemical staining, migration, invasion, and tumorigenicity. However, prognosis of both patients was quite similar, as donors of the cell lines were metastatic tumors with survival time after diagnosis of 150 and 191 days, respectively. It is, therefore, phenotypes of cell lines may not be readily associated with prognosis of the patients.
Both KKU-452 and KKU-023 cells grew as adherent monolayer cells with morphology of epithelial cells. KKU452 cells proliferate at much higher doubling time than KKU-023 cells (17.9 vs 34.8 h). KKU-452 and KKU-023 cells were derived from original tumor tissues of different histological grades, that is, poorly differentiated and well-differentiated adenocarcinoma, respectively. However, histological grade of differentiation does not seem to be associated with prognosis of the patients, unlike the histological types where papillary carcinoma has better prognosis than non-papillary type. 31 In this study, both cell lines are tubular types (non-papillary type) with different histological grading. The immunophenotypic expression of cytokeratins has been used as epithelial cell markers for the characterization of GI tract tumors, and CCA are reportedly positive for CK7, CK17, and CK19. 32 In this study, both cell lines were positive for CK7 and CK19, although KKU-023 showed much stronger staining than KKU-452 cells. Expression of CK19 is associated with prognosis of hepatocellular carcinoma (HCC) and combined HCC and CCA, 33 although this association is unclear for CCA.
In tumorigenesis, the multistage alteration of genomic composition leads to complex changes in genome. Cytogenetic analysis of KKU-452 and KKU-023 cells revealed the highly complex karyotypes with hypertriploid and hypotetraploid (modal chromosome number of 74 and 85, respectively) and multiple chromosomal aberrations. Both CCA cells show complex numerical and structural aberrations similar to the previously established CCA cell lines KKU-10014 and RMCCA-1. 13 The multiple structural aberrations in tumor cells have been associated with the presence of mismatch repair defects, with a replication error (RER) negative phenotype. 34 This is in contrast with a minority of colon cancers which is characterized by stable near-diploid karyotypes that showed the RER positive phenotype. 34 However, exact causes of chromosomal aberration are not known yet, and these may reflect the genomic instability in the tumor trigger by various mechanisms.
In mutation analysis, we screened KRAS and TP53 genes, as they are the frequent mutations observed in Ov-associated CCA. In KRAS gene, we sequenced exon 2, as mutations at codons 12 and 13 on exon 2 could be accounted for about 99% of mutations in KRAS. 35 KRAS is a member of a large family of small guanosine triphosphate (GTP)-binding proteins, where activating mutation is a key early event of RAF/mitogen-activated protein kinase mitogen-activated protein kinase kinase (MEK)/extracellular signal–regulated kinase (ERK) and phosphatidylinositol-4,5-bisphosphate 3-kinase (PI3K)/Akt signaling cascades controlling processes such as cell growth and survival. 36 KRAS mutations are found over one-third of colorectal cancer and 12%–14% in CCA.10,35 The two newly established CCA cell lines in this study have no detectable mutation, while the established reference KKU-100 cells have mutation at codon 12 (G12D). However, TP53 mutation was found in all three cell lines examined, particularly at codon 72 (Pro > Arg). This position is located within a proline-rich region and may affect the structure of DNA-binding domain in TP53. There was a suggestive evidence that polymorphism in TP53 at this position may influence individual responsiveness to cisplatin-based chemo-radiotherapy, where cancers expressing 72R mutants show lower response rates than those expressing 72P. 37 In addition, KKU-452 cells have a nonsense mutation on codon E349, suggesting a small truncation of the protein at the tetramerization domain (TD). The TD plays important role in facilitating p53 DNA binding and interactions with other proteins which enhances the stability of the overall protein–DNA complex. Mutations of TD including missense and nonsense mutations have been reportedly impaired p53 wild-type function. 38 Expression of p53 protein in KKU-452 cells was very high, while that of KKU-023 and KKU-100 cells was very low under basal condition. Nonetheless, the expression of p53 in KKU-100 cells was shown to be highly inducible by gemcitabine and dicoumarol. 22 It suggests that the single-nucleotide polymorphism (SNP) may also affect stability of p53 protein. However, the effect of mutation on codon E349 on p53 function is still unknown.
KKU-452 cells showed a remarkable migration and invasion activities when compared to KKU-023 cells or reference KKU-100 cells. There is no relationship in between histological grade of CCA and invasion/migration activities, since KKU-100 and KKU-452 cells are both the poorly differentiated adenocarcinoma. The rapid growing and extensive cell seeding of KKU-452 cells may be related to the aforementioned phenotypes. A key step in tumor progression is the transition of stationary epithelial cells to become motile by the loss of cell–cell adhesion and matrix degradation. The process of epithelial–mesenchymal transition (EMT) exhibits molecular hallmark by downregulation of E-cadherin, upregulation of matrix-degrading enzymes, and mesenchymal-related proteins such as MMP-9, N-cadherin, and vimentin. 39 The phenotype of cell invasion is consistent with western blot analysis which has revealed low and high expression of E-cadherin and N-cadherin, respectively, in KKU-452 cells, when compared with KKU-023 cells. However, KKU-452 cells did not form an apparent tumor in nude mice in current settings, while KKU-023 cells rapidly form tumor at injection site. The reference cell line, KKU-100 cells, is also shown to form tumor in nude mice. 40 The athymic nude mice used in the study have a T-cell deficiency; however, other immune components are still intact. The tumorigenic property of KKU-452 cells may have to evaluate in other animal models. It is also apparent that current biomarkers (E-cadherin, N-cadherin, and vimentin) employed in the study are not sufficient in predicting tumorigenicity of cultured cells.
Histology of the xenografted tumor tissue in mice is in agreement with that of the original tumor from CCA patient. Both tumor tissues exhibit tubular adenocarcinoma and both tissues were also stained with CK19, a marker of bile duct epithelium. This may confirmed a cell line derivation from human tumor tissue. We did not find tumor metastasis in distant organs within the study period in mice transplanted with KKU-023 and KKU-452 cells. The relatively short observation time after transplantation (3–6 weeks) may not allow an obscure metastatic tumor to be detectable.
In conclusion, we established two CCA cell lines from the tumor tissues of perihilar and intrahepatic mass-forming CCA of the patients who resided at the endemic area of carcinogenic liver fluke infection. The cancer cell lines represent a valuable tool in research on this disease, particularly CCA cells from various origins, for better understanding the process of carcinogenesis and developing new therapeutics for the patients.
Footnotes
Acknowledgements
The authors would like to acknowledge Dr Hasaya Dokduang of Department of Biochemistry, Faculty of Medicine for performing mouse xenograft experiment and Prof. Yukifumi Nawa for language editing of the manuscript via Publication Clinic, Khon Kaen University. Laddawan Senggunprai, Nisana Namwat, Sarinya Kongpetch, Puangrat Yongvanit, and Veerapol Kukongviriyapan designed the study; Sunitta Saensa-ard, Saman Leuangwattanawanit, Yaovalux Chamgramol, Watcharin Loilome, Walaiporn Khansaard, Apinya Jusakul, Auemduan Prawan, and Chawalit Pairojkul performed experiments; Narong Khantikeo provided clinical specimens; and Veerapol Kukongviriyapan and Sarinya Kongpetch wrote manuscript. All authors approved the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
The study protocol has been approved by the Khon Kaen University Ethics Committee for Human Research (HE571283) and experiments were performed in accordance with the Declaration of Helsinki. The animal study protocol was approved by the Animal Ethics Committee, Northeast Laboratory Animal Center of Khon Kaen University (AEKKU-NELAC 7/2558), and all experiments were performed in accordance with the institutional guidelines.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Cholangiocarcinoma Research Institute, Khon Kaen University and grant-in-aid from National Research Council of Thailand via Khon Kaen University.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
