Abstract
The role of circulating microRNAs as a promising tool for diagnosing cancer and monitoring anticancer therapies has been widely studied in the past decades. To date, no suitable reference microRNAs for normalizing quantitative real-time polymerase chain reaction assays has been identified in vulvar intraepithelial neoplasia lesions and vulvar squamous cell carcinoma. The purpose of this study was to select appropriate references for gene expression studies in plasma of patients with these lesions. Expression levels of six microRNAs—hsa-miR-425-5p, hsa-miR-191-5p, hsa-miR-93-5p, hsa-miR-423-5p, hsa-miR-103a-3p, and hsa-miR-16-5p—were analyzed by quantitative reverse transcription polymerase chain reaction in plasma samples obtained from 17 patients with vulvar intraepithelial neoplasia lesion and 27 patients with vulvar squamous cell carcinoma. The expression stability of these candidate normalizers was assayed using geNorm algorithm. hsa-miR-93-5p was revealed as the most stably expressed reference in plasma samples of both vulvar intraepithelial neoplasia lesion and vulvar squamous cell carcinoma patients. The results pointed at hsa-miR-93-5p and hsa-miR-425-5p as microRNAs that retained the greatest robustness in plasma of vulvar intraepithelial neoplasia lesion and vulvar squamous cell carcinoma patients, respectively. Our work is the first report on reference microRNA selection for quantitative real-time polymerase chain reaction applications in vulvar intraepithelial neoplasia lesion and vulvar squamous cell carcinoma. The candidate microRNA stability values for the two types of lesions are provided and might serve for normalization of the future novel microRNA biomarkers in these rare entities.
Keywords
Introduction
Vulvar carcinoma is a rare disease, with an estimated 27,000 new cases in 2008 worldwide, representing 4% of all gynecologic cancers. 1 An age-standardized rate is between 0.5 and 1.5 per 100,000. 2 There were 490 new cases diagnosed in 2013 in Poland (with standardized incidence rate: 1.61/100,000) and 303 deaths (with standardized mortality ratio: 0.86/100,000) caused by this malignancy. 3 The 5-year survival rates for vulvar cancer are approximately 70%, regardless of the stage (http://seer.cancer.gov/statfacts/html/vulva.html). Vulvar squamous cell carcinoma (VSCC) accounts for more than 90% of malignant tumors of the vulva. 4
It is widely recognized that VSCC develops on the basis of vulvar intraepithelial neoplasia (VIN) lesions. In the classification adopted in 2015 by the International Society for the Study of Vulvar Disease (ISSVD), vulvar precancerous lesions are divided into two types, differing in their pathogenesis and clinical significance: high-grade squamous intraepithelial lesion (HSIL, VIN usual type (uVIN)) and VIN, differentiated-type (dVIN). 5 The pathogenesis of VSCC can follow two different pathways: human papillomavirus (HPV)-dependent and HPV-independent.6,7 uVIN is nearly exclusively associated with HPV infection (98.8% of cases). 8 This pathway is more common in younger women and is associated with smoking, higher number of sexual partners, and compromised immune status. 6 In dVIN, HPV infectionis detected occasionally (8.3% of cases). 8 The risk of progression from uVIN to VSCC is low and ranges from 3% to 16%, while for dVIN, it reaches 32.8%. Van de Nieuwenhof et al. 9 recorded that median time of progression to VSCC was 22.8 months for dVIN and significantly longer, that is, 41.4 months, for uVIN.
The mechanisms leading to the development and progression of VIN to VSCC remain unclear. Nevertheless, a thorough appreciation of the natural history of this malignancy would be essential for the identification of molecular markers allowing the diagnosis of VSCC at early stage and/or indicating progression risk of premalignant lesions. The aim of this study was to identify the suitable references for microRNA (miRNA) expression analysis of plasma of VIN and VSCC patients using quantitative reverse transcription polymerase chain reaction (qRT-PCR).
Materials and methods
A total of 44 patients with VIN and VSCC treated between February 2005 and October 2014 were included in the study. Among 17 enrolled patients treated for VIN in Holycross Cancer Center in Kielce (HCC) and the Medical University of Warsaw (WUM), 12 and 5 were diagnosed with HSIL and dVIN, respectively. The median age of patients treated for VIN was 58.4 years (range: 33–79). Overall, 27 patients surgically treated for primary early-stage VSCC (T1-2, N0-2, M0) at the Maria Skłodowska-Curie Institute—Oncology Center (MSCI) were recruited. No patient received preoperative chemotherapy or radiotherapy. The selected characteristics of VSCC patients are presented in Table 1. The study was approved by the Independent Ethics Committee of all the institutions, MSCI (No. 44/2002, 16/2015), HCC (No. 15/2014), and WUM (No. 247/2015), and all patients gave their informed consent.
Epidemiological, anatomopathological, and clinical characteristics of the VSCC patients selected for the study.

VSCC: vulvar squamous cell carcinoma; FIGO: International Federation of Gynecology and Obstetrics.
Plasma samples were obtained from patients’ blood before their surgical treatment and stored at −70°C until RNA isolation. Total RNA was isolated from 200 µL of plasma using miRNeasy Mini Kit (QIAGEN), and miRNA was reverse transcribed using miScript II RT Kit (QIAGEN) according to the manufacturer’s protocol. For each reverse transcription, 3.2 × 108 copies of synthetic cel-miR-39-3p (miRNeasy Serum/Plasma Spike-In Control; QIAGEN) were added to monitor reaction efficiency. Expression levels of six candidate reference miRNAs, that is, hsa-miR-93-5p, hsa-miR-425-5p, hsa-miR-16-5p, hsa-miR-103a-3p, hsa-miR-191-5p, hsa-miR-423-5p, were analyzed by qRT-PCR using the miScript miRNA Arrays (QIAGEN) according to the manufacturer’s protocol. Reactions were performed in 7500 Fast Real-Time PCR System (Applied Biosystems) in a final volume of 25 μL reaction mix, according to the arrays manufacturer’s instructions (QIAGEN). Gene expression data were analyzed using geNorm™ algorithm (https://genorm.cmgg.be/) integrated into DataAssist™ Software (Applied Biosystems, Thermo Fisher Scientific). RefFinder, a web-based application which integrates three most commonly used normalization algorithms (geNorm, Normfinder, and BestKeeper) was used for cross-validation of results. 10
Results
The algorithm used allowed to determine the most stable reference molecules from a set of tested candidate normalizers in a given sample panel. It calculates a gene stability measure (M) according to formulas described by Vandesompele et al. 11 Genes with the lowest M values (<1.5) have the most stable expression. Our analysis of miRNA expression data with the algorithm identified hsa-miR-93-5p (M = 1.3276) as the gene that retained the greatest robustness in all the plasma samples analyzed, that is, of both VIN and VSCC patients. We identified hsa-miR-93-5p (M = 1.4481) as the most stable reference miRNA for VIN. In plasma samples of patients with VSCC, hsa-miR-425-5p (M = 1.017) followed by hsa-miR-93-5p (M = 1.044) and hsa-miR-191-5p (M = 1.3937) were the most stable reference miRNAs (Table 2). RefFinder analysis confirmed hsa-miR-425-5p and hsa-miR-93-5p as the most stably expressed normalizers, with comprehensive stability values of 1.56 and 1.68, respectively (Figure 1).
Putative reference miRNAs and their expression stability values in plasma samples of VIN (n = 17) and VSCC (n = 27) patients.

miRNA: microRNA; VIN: vulvar intraepithelial neoplasia; VSCC: vulvar squamous cell carcinoma.
The bold values represent the lowest stability values, i.e. indicate the best normalizers.
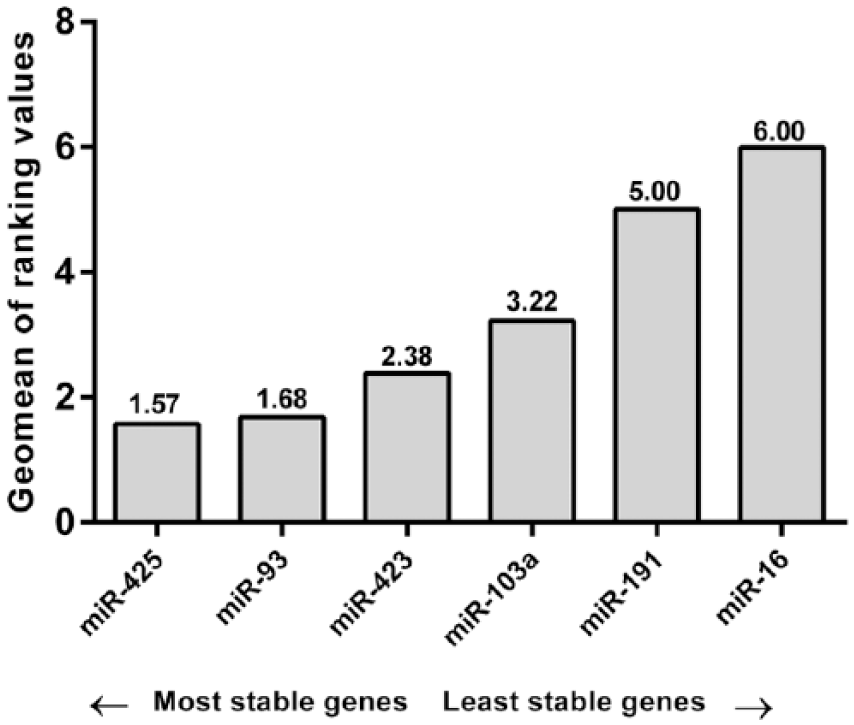
Expression stability values of the candidate references as calculated by the Geomean method of RefFinder.
Discussion
Extracellular miRNAs have been shown to be associated with numerous diseases, including cancer. 12 Tumor-derived miRNAs were found to be maintained in human plasma and serum in stable form pointing to their clinical utility as promising blood-based cancer detection strategy.13,14 Indeed, several studies reported that circulating miRNAs are potential biomarkers for the diagnosis of numerous cancers,14,15 including those of gynecological origin.16–19
The initial study of de Melo Maia et al. 20 shed some light on the miRNA signature in vulvar carcinoma identifying 79 miRNAs that showed markedly altered expression patterns in tumors compared to control tissues. Deregulated expression of several identified miRNAs correlated with clinicopathological features such as the presence of lymph node metastasis (downregulation of miR-223-5p and miR-19-b1-5p), vascular invasion (downregulation of miR-100-3p and miR-19-b1-5p), HPV infection (upregulation of miR-1274b and downregulation of miR-519b), or advanced International Federation of Gynecology and Obstetrics (FIGO) stage (miR-519b and miR-133a overexpression). Later, the upregulation of miR-20a and miR-106a was demonstrated to correlate with deeper tumor invasion in VSCC. 21 Moreover, the promising preliminary results of in vitro experiments on novel miRNA-based therapeutic option against VSCC merit further investigation and ultimately development for clinical translation. 21 Recently, other authors documented that miR-590-5p is involved in the VSCC carcinogenesis, and its upregulation in tumors is associated with lymphatic metastases. 22 To the best of our knowledge, the circulating miRNAs in patients with VSCC or VIN have not been examined to date.
qRT-PCR is a gold standard method employed to quantify miRNAs; however, no universal reference miRNA has so far been identified.23,24 There is no substitute for empirical validation of normalization that is appropriate for the particular experimental design and goals. 25 Therefore, we performed literature search to identify suitable reference genes for miRNA expression analysis by qRT-PCR in VSCC and VIN patients’ plasma. As this search revealed no validated endogenous controls for miRNA analysis in VSCC and VIN, we chose the miRNA basing on data documenting their presence and stable levels in a variety of biological tissues and body fluids. 26 MiR-93 has been identified as a suitable reference miRNA to normalize miRNA levels in studies of disparate diseases from gastric cancer to major depression disorders.27,28 Other investigations have proven that miR-93 is differentially regulated in cancer tissues and its levels may be predictive of disease state in ovarian and breast cancer,29,30 further highlighting the need to confirm the suitability of selected miRNAs to normalize miRNA expression data depending upon the investigated tissue. MiR-93 was found to be a suitable reference gene for serum miRNA analysis in gastric cancer patients and healthy controls. 28 Our data demonstrate that miR-93 is also a suitable reference miRNA for analyzing miRNA levels in plasma of VIN and both VIN and VSCC patients. MiR-425 was demonstrated as stably expressed in colorectal tumors and potentially suitable reference for miRNA studies. 31 Both miR-93 and miR-425 are also recommended by the manufacturer as usually stably expressed in serum/plasma (https://www.exiqon.com/ls/Documents/Scientific/serum-plasma-mirna-profiling.pdf, accessed 20 June 2016).
MiR-16 was used to normalize miRNA levels in serum samples of patients with various tumor types. 14 More recent studies have reported upregulation of plasma miR-16 in plasma due to several pathological conditions, including autoimmune thyroid disease, 32 experimental sepsis, 33 and gastric cancer. 34 Another study demonstrated that miR-16 is highly expressed in red blood cells so that hemolysis strongly affects its plasma levels. 35 In our study, miR-16 did not meet the criteria of a stable reference miRNA in VIN and VSCC as it had variable expression between plasma samples examined. MiR-103 proved to be the most stable miRNA in the normal and cancerous lung tissues. 36 MiR-191-5p was described as reference miRNA for serum qRT-PCR analysis in colorectal adenocarcinoma, colorectal adenoma, and healthy controls. 37 As shown in our work, miR-191-5p was stably expressed in sera of VSCC patients but not in the VIN samples analyzed in our study. MiR-423 was identified as the least variable among 345 miRNAs in the 40 normal tissues in the study of Liang et al. 38 hsa-miR-103a-3p, hsa-miR-191-5p, and hsa-miR-423-3p are also considered acceptable as normalizers in body fluids by the manufacturer, if information on good normalizers is lacking (www.exiqon.com/ls/Documents/Scientific/microRNA-serum-plasma-guidelines.pdf, accessed 20 June 2016).
Our overall analysis showed that hsa-miR-93-5p followed by hsa-miR-425-5p are the most suitable genes as internal controls for quantitative miRNA expression studies in these rare gynecological tumors. When stratified by disease, the results still pointed to hsa-miR-93-5p as the most stable single gene in VIN, where hsa-miR-425-5p showed the M value smaller than the algorithm default threshold of 1.5 in VSCC. However, we are aware of the study limitations. First, vulvar carcinoma and its precursor lesions are rare diseases, and for that reason, the number of clinical samples in our study was relatively low. Although it is assumed that for the algorithm used, analysis of 10 samples is the sufficient set to be analyzed, 11 and this recommendation was met in our study, we intend to validate our results in the plasma samples obtained from a larger cohort of VIN and VSCC patients as well as of healthy volunteers to confirm the clinical importance of these two reference miRNA.
Second, only six candidate reference genes were analyzed by the qRT-PCR. However, these miRNAs proved to have a good expression stability in the analyzed plasma sample set. Importantly, our results were also confirmed using RefFinder web-based application which integrates three most commonly used normalization algorithms (geNorm, Normfinder, and BestKeeper). 10 RefFinder also indicated hsa-miR-93-5p and hsa-miR-425-5p as the most stable miRNAs in VIN and VSCC, respectively. Although in all the patients, that is, with VIN and VSCC, hsa-miR-425-5p was more stable than has-miR-93-5p, still both these molecules were identified as the best normalizers among the six analyzed.
Conclusion
Our work is the first report on reference selection for miRNA profiling in plasma of VIN and VSCC patients. The use of more than one normalizer increases the accuracy qRT-PCR data analysis. 24 Therefore, a ranking of candidate miRNA stability values for these types of tumors is provided and might serve as a valuable guide for future gene expression studies of these rare entities. Explanation of the molecular expression signatures may lead to more accurate diagnostics, including differential diagnosis of preinvasive vulvar lesions (HSIL and dVIN), as well as identification of prognostic factors and ultimately targets for future therapeutics.
Footnotes
Acknowledgements
The authors thank the donors who volunteered to take part in this study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval and consent to participate
This study was conducted after obtaining a research ethics board approval and the institutional authorization of all the institutions, CO-I (No. 44/2002, 16/2015), HCC (No. 15/2014), and WUM (No. 247/2015), and all patients gave their informed consent.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the Polish National Science Centre (grant no. 2013/10/E/NZ5/00663).
