Abstract
Long-term persistent infection of HPV16 E6/E7 is frequently associated with lung cancers, especially in non-smokers and in Asians. However, molecular mechanisms of HPV16 E6/E7 induction of lung cancer are not fully understood. Using bi-directional genetic manipulation and four well-established lung cancer cell lines, we showed HPV16 E6/E7 downregulated expression of liver kinase B1 at both protein and messenger RNA levels; liver kinase B1 downregulated hypoxia-inducible factor 2α at protein level but not at messenger RNA level, and hypoxia-inducible factor 2α upregulated vascular endothelial growth factor at both protein and messenger RNA levels. This is the first study to show hypoxia-inducible factor 2α as a downstream effector of liver kinase B1 in lung cancer cells. Our results indicate that HPV16 E6/E7 indirectly upregulated the expression of vascular endothelial growth factor by inhibition of liver kinase B1 expression and upregulation of hypoxia-inducible factor 2α expression, thus propose a human papillomavirus–liver kinase B1–hypoxia-inducible factor 2α–vascular endothelial growth factor axis for the tumorigenesis of lung cancer. Our study also provides new evidence to support the critical role of liver kinase B1 in the pathogenesis of human papillomavirus–related lung cancer and suggests novel therapeutic targets.
Keywords
Introduction
Lung cancer is the leading cancer killer worldwide, and the pathogenesis of this deadly disease is far from established. Although smoking is the established major etiologic factor, association of human papillomavirus (HPV) and lung cancer has been proposed by a series of reports, especially in non-smokers and in Asians.1–4 In actual fact, a recent meta-analysis indicated that 38% lung cancers are positive for HPV, with HPV16 as the most prevalent subtype.5,6 High titers of IgG antibodies against HPV16 and HPV18E7 are also detected in 16% of non-small-cell lung carcinomas (NSCLCs). 7 Molecular carcinogenesis of HPV has been extensively studied in cervical as well as in head and neck carcinomas, 8 but studies to address mechanism(s) of HPV induction of lung cancer are just emerging. It is known that HPV16 E6/E7 inactivates tumor-suppressor genes p53, pRB, and other host proteins to induce genomic instability and to abrogate cell-cycle regulation. HPV16 promotes angiogenesis in NSCLC by enhancing tumor cell expression of hypoxia-inducible factor 1α (HIF-1α), vascular endothelial growth factor (VEGF), and interleukin-8 (IL-8). 9 While above mechanisms are sufficient to induce cell immortalization and proliferation, current literature suggest that additional factors are necessary for HPV to transform human epithelial cells into cancers.
Liver kinase B1 (LKB1) is a serine/threonine kinase and tumor suppressor, it plays critical role in cell growth, polarity, and metabolism. LKB1 mutation causes Peutz–Jeghers syndrome 10 and increases risk of several types of cancer. 11 Mutational inactivation of LKB1 has been identified in 20%–30% NSCLC and 10% atypical adenomatous hyperplasia of lung.12,13 LKB1 loss has also been found to promote lung cancer progression and resistance to treatment.14,15 However, the downstream effectors of LKB1 in cancers including lung cancer are not fully elucidated.
Hypoxia-inducible factor 2α (HIF-2α) is a transcription factor of HIF class, which also include HIF-1α, HIF-3α, and HIF-βs. Similar to HIF-1α, HIF-2α induces the genes promoting angiogenesis including VEGF. 16 In comparison to HIF-1α, HIF-2α also involves regulation of genes important for tumor growth, cell-cycle progression, and maintenance of stem cell pluripotency. 17 Hypoxia is the main stimulator of HIF activity, and proteasomal degradation of HIFs is also oxygen dependent. Both HIF-1α and HIF-2α are upregulated in lung cancer and HIF-2α has been reported to be associated with poor prognosis. 18 Our previous study found higher VEGF messenger RNA (mRNA) expression by bronchial brushing cells in lung cancer patients. 19 However, the relationship between LKB1 and HIF-2α has not been previously reported.
In this study, we demonstrated that HPV16 E6/E7 inhibited the expression of LKB1, and downregulation of LKB1 increases the expression of HIF-2α and subsequent VEGF in several lung cancer cell lines. Our results are the first to illustrate that HIF-2α is a downstream effector of LKB1. Functional aspects of HIF-2α upregulation by HPV/LKB1 in lung cancer cells, including angiogenesis, are being further investigated.
Materials and methods
Our study protocol is approved by the Institutional Ethnical Review Committee.
Cell culture
Four human NSCLC cell lines H1299, NCI-H460, A549, and LK2 were used in this study. H1299, NCI-H460, and A549 cells, purchased from ATCC (Manassas, VA, USA), were cultured in HyClone RPMI 1640 medium. LK2 cells, obtained from the cell bank of Chinese Academy of Sciences (Shanghai, China), were grown in HyClone (Logan, UT, USA) Dulbecco’s modified Eagle’s medium (DMEM). The culture mediums were supplemented with 10% fetal bovine serum, and the cells were cultured at 37°C in a 5% CO2 humidified atmosphere. For transfections, cells were seeded in a six-well plate 24 h before the experiments.
Plasmid construction and transfection
The pEGFP-N1-HPV16 E6, pEGFP-N1-HPV16 E7, and pEGFP-N1 plasmids were kindly provided by Professor Xudong Tang (Institute of Biochemistry and Molecular Biology, Guangdong Medical College, China). Plasmids for LKB1 (pcDNA3-LKB1-His and pcDNA3-His) were kindly provided by Professor Xin Hou (College of Life Sciences, Inner Mongolia University, Huhhot, Inner Mongolia, China). The plasmid of Ha-HIF-2alpha pcDNA3 (Plasmid #18950) was purchased from Addgene (Cambridge, MA, USA). Plasmid with target was transiently transfected into the cells accordingly using Lipofectamine 3000 Transfection Kit (Invitrogen, Carlsbad, CA, USA). Transfection with empty vector and mock transfection served as the controls. The transfection efficiency was evaluated by visualizing the cells expressing Green Fluorescent Protein (GFP) directly under fluorescence microscopy. The protein analysis was assessed 48 h after transfection by western blotting, and the mRNA analysis was assessed 24 h after transfection by quantitative real-time reverse transcriptase-polymerase chain reaction (qRT-PCR).
Small-interfering RNA
HPV16 E6 small-interfering RNA (SiRNA; sc-156008), HPV16 E7 SiRNA (sc-270423), LKB1 SiRNA (sc-35816), and Control SiRNA (sc-37007) were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). EPAS1 SiRNA (HIF-2α SiRNA) and scrambled siRNA were purchased from RiboBio (Guangzhou, China). Scrambled siRNA was used as nonspecific siRNA control. For the siRNA transfection, cells were seeded at 5 × 104 cells/35-mm well for 24 h according to manufacturer’s instructions, and then, the siRNA was transfected into the cells using Lipofectamine RNA iMAX reagent (Invitrogen). Transfected cells were incubated for another 48 h and then they were ready for the analyses.
Western blot analysis
Cells were lysed in the lysis buffer to obtain the total protein. After the concentration of the protein was measured by the Bradford method, 60 µg of protein was separated on a 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and then transferred to a polyvinylidene difluoride (PVDF) membrane (Millipore, Bedford, MA, USA). The membrane was blocked with 5% skimmed milk and incubated with the following primary antibodies separately overnight at 4°C: HPV16 E6 (1:100; Bioss, Beijing, China), HPV16 E7 (1:100; Bioss, Beiing, China), LKB1 (1:1000; Cell Signaling Technology, Boston, MA, USA), HIF-2α (1:600; Abcam, Cambridge, MA, USA), VEGF (1:600; WanLei, China), and glyceraldehyde 3-phosphate dehydrogenase (GAPDH; 1:1000; Cell Signaling Technology, Boston, MA, USA).
After incubated with horseradish peroxidase (HRP)-conjugated secondary antibody (anti-rabbit IgG), the membrane was developed in enhanced chemiluminescence (ECL). The density of each protein band was measured and normalized to the density of GAPDH by BioImaging System.
qRT-PCR
Total RNA was extracted from cells using TRIzol Reagent (Life Technologies, Carlsbad, CA, USA) and RNeasy RNA isolation kit (QIAGEN, Hilden, Germany). First-strand cDNA was synthesized from 500 ng total RNA using high-capacity cDNA RT Kit (Applied Biosystems, Carlsbad, CA, USA). The qRT-PCR was performed using SYBER GreenMaster Mix on 7900HT Fast Real-Time PCR system (Applied Biosystems). Non-template controls every time were prepared for each primer pair to detect nonspecific amplification. Average threshold cycle (Ct) values for GAPDH (used as housekeeping gene) were used to normalize average Ct values of the gene of interest. These values were used to calculate the average Ct values between groups, and the relative quantity (power of −ΔΔCt) was used to calculate fold change between each group. The detailed information of the primers is given in Table 1.
Sequences and features of primers used for qRT-PCR.

mRNA: messenger RNA; qRT-PCR: quantitative real-time reverse transcriptase-polymerase chain reaction.
The amplified products of E6, E7, LKB1, HIF-2α, VEGF, and GAPDH were confirmed by DNA gel with correct sizes. The products were extracted, purified from the gel, and sent for DNA sequencing. The sequencing results were 100% correct.
Statistical analysis
All statistical analyses were performed using SPSS for Windows 13.0 (SPSS Inc, Chicago, IL, USA). All data presented three or more experiments with similar results. Statistical significance was determined by Student’s t-test, and a p value <0.05 was considered significant.
Results
The screening of lung cancer cell lines
Based on our previous results, H1299 and NCI-H460 cells are low E6 and E7 expression cell lines, and A549 and LK2 cells are high E6 and E7 expression cell lines. 20 Further assays were designed and performed based on these results.
E6 and E7 downregulated the expression of LKB1 and upregulated HIF-2α and VEGF expression
Due to transient transfection of pEGFP-N1-HPV16 E6 or pEGFP-N1-HPV16 E7 to H1299 and NCI-H460 two low expression cell lines, the expression levels of E6 or E7 increased significantly. The elevated levels of E6 and E7 downregulated the expression of LKB1 at protein level and at mRNA level, increased HIF-2α expression at protein level only, as well as upregulated VEGF expression at both protein and mRNA levels. There were no significant changes in HIF-2α mRNA expression level in both H1299 and NCI-H460 cells. The results are presented in Figure 1.

Overexpression of E6, E7, HIF-2α, and VEGF, but low expression of LKB1 in H1299 and NCI-H460 cells. Detection of the expression of protein and mRNA of E6, E7, LKB1, HIF-2α, and VEGF in lung cancer cells as well as in control cells by western blot and qRT-PCR. GAPDH served as an internal control.
Knocking down E6 and E7 upregulated the expression of LKB1 and downregulated the expression of HIF-2α and VEGF
Based on our screening results, A549 and LK2 cells were selected as high E6 and E7 expression cell lines. The expression of LKB1 was low, but HIF-2α and VEGF were high in these two cell lines. Using E6-specific siRNA or E7-specific siRNA techniques, we knocked down the expression of E6 or E7 in these two cell lines, which result in upregulation of the expression of LKB1 at both protein and mRNA levels, downregulation of HIF-2α expression at protein level only, and downregulation of VEGF expression at both protein and mRNA levels. There were no significant changes in HIF-2α mRNA expression level in both A549 and LK2 cells. The results are presented in Figure 2.

Low expression of E6, E7, HIF-2α, and VEGF, but overexpression of LKB1 in A549 and LK2 cells. Detection of the expression of protein and mRNA in lung cancer cells as well as in control cells by western blot and qRT-PCR. GAPDH served as an internal control.
Overexpression of LKB1 downregulating the expression of E6, E7, HIF-2α, and VEGF
Transient transfection of pcDNA3-LKB1-His to the low LKB1 expression cells A549 and LK2 was performed. Overexpression of LKB1 decreased the expression of E6, E7, and VEGF at both protein and mRNA levels as well as HIF-2α at protein level only. There were no significant changes in HIF-2α mRNA expression level in both A549 and LK2 cells. The results are presented in Figure 3(a).

(a) Overexpression of LKB1, but low expression of E6, E7, HIF-2α, and VEGF in A549 and LK2 cells. Detection of the expression of protein and mRNA of LKB1, HIF-2α, and VEGF in lung cancer cells as well as in control cells by western blot and qRT-PCR. GAPDH served as an internal control (mock: mock transfection; vector: empty vector; ns: no significance (*p < 0.05; **p < 0.01)). (b) Low expression of LKB1, but overexpression of E6, E7, HIF-2α, and VEGF in H1299 and NCI-H460 cells (Mock: mock-specific siRNA; NS: nonspecific siRNA; SiLKB1: LKB1-specific siRNA (*p < 0.05; **p < 0.01)). Detection of the expression of protein and mRNA of them was in lung cancer cells as well as in control cells by western blot and qRT-PCR. GAPDH served as an internal control.
Knocked down LKB1 upregulated the expression of E6, E7, HIF-2α, and VEGF
Using LKB1-specific siRNA techniques, we knocked down the expression of LKB1 in both H1299 and NCI-H460 cell lines, which resulted in upregulation of E6, E7, and VEGF at both protein and mRNA levels as well as HIF-2α at protein level only. There were no significant changes in HIF-2α mRNA expression level in both H1299 and NCI-H460 cells. The results are presented in Figure 3(b).
HIF-2α upregulation of the expression of VEGF
Transient transfection of Ha-HIF-2alpha pcDNA3 to H1299 and A549 two cell lines overexpressed HIF-2α significantly. Which result in upregulation VEGF expression at both protein and mRNA levels. The results are presented in Figure 4(a).
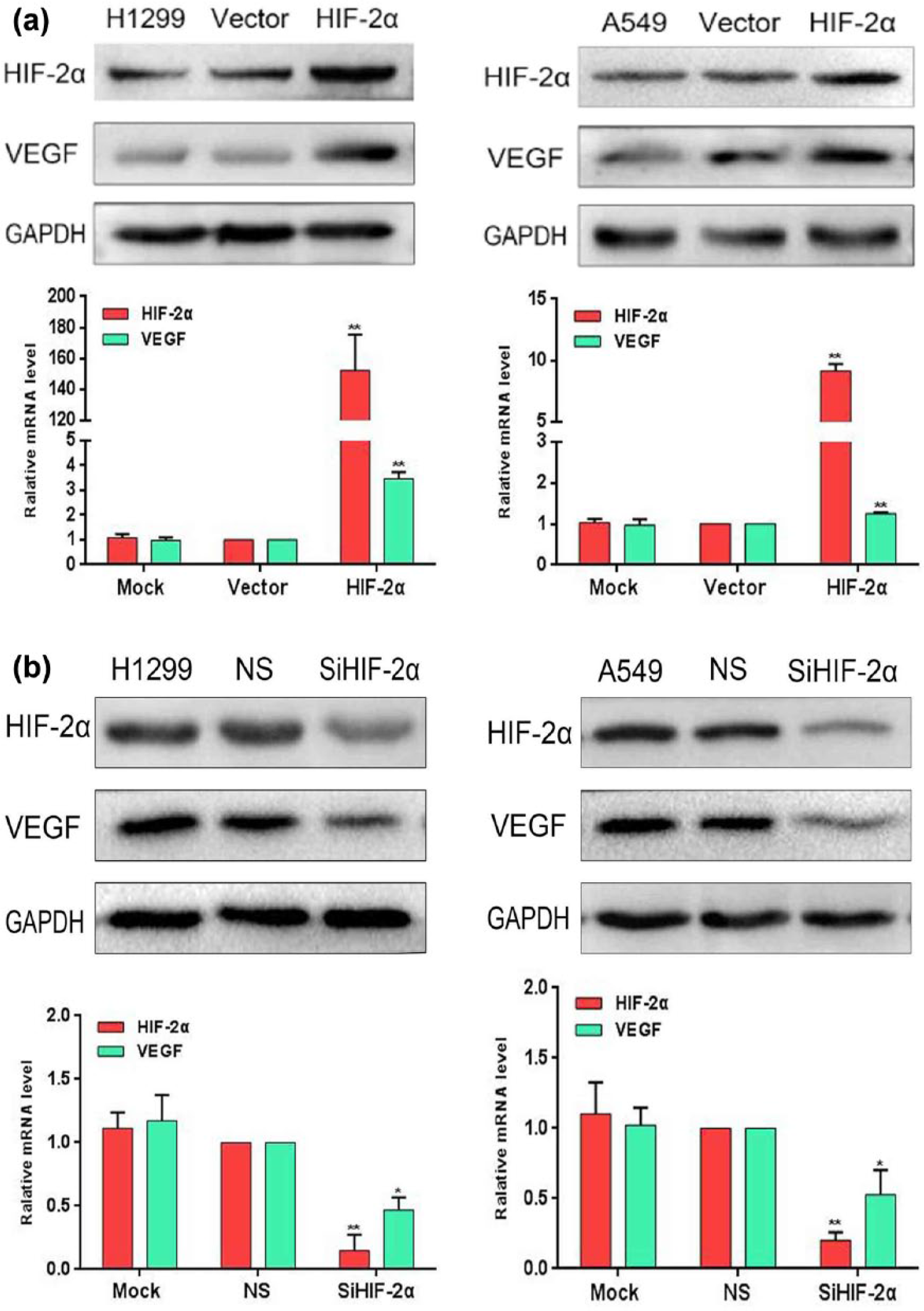
(a) Overexpression of HIF-2α, but low expression of VEGF in H1299 and NCI-H460 cells. Detection of the expression of protein, mRNA, HIF-2α, and VEGF in lung cancer cells as well as in control cells by western blot and qRT-PCR (mock: mock transfection; vector: empty vector; ns: no significance (*p < 0.05; **p < 0.01)). GAPDH served as an internal control. (b) Low expression of HIF-2α, but overexpression of VEGF in H1299 and NCI-H460 cells. Detection of the expression of protein and mRNA in lung cancer cells as well as in control cells by western blot and qRT-PCR (mock: mock-specific siRNA, NS: nonspecific siRNA; SiHIF-2α: HIF-2α-specific siRNA (*p < 0.05; **p < 0.01)). GAPDH served as an internal control.
Knocked HIF-2α downregulated the expression of VEGF
Using HIF-2α-specific siRNA techniques, we knocked down the expression of HIF-2α in both H1299 and A549 cell lines, which result in downregulation of VEGF expression at both protein and mRNA levels. The results are presented in Figure 4(b).
Discussion
In this study, we provide evidence that the overexpression of E6 and E7 in lung cancer cells downregulates the expression of LKB1 at both protein and mRNA levels, and the LKB1 knockdown upregulates HIF-2α protein expression, with subsequent upregulation of VEGF protein and mRNA (Figures 1 and 2). Recent studies had showed that high-risk HPV16 infection is closely related to the inactivation of tumor-suppressor gene LKB1 in the pathogenesis of cervical cancer,21,22 others had found that the inactivation of tumor-suppressor gene LKB1 is also closely related to the occurrence of lung cancer; 23 however, it remains unclear whether high-risk HPV16 persistent infection is related to the inactivation of LKB1 in the occurrence of lung cancer. In this study, we have demonstrated for the first time that HPV16 is a persistent infection and the cancer proteins E6 and E7 in HPV16 have obvious inhibitory effects on LKB1. Furthermore, we demonstrated that HIF-2α and VEGF are downstream effectors of LKB1, revealing new molecular mechanisms of HPV16 E6/E7-induced tumorigenesis in NSCLC.
Genetic manipulation of various genes provides a powerful tool for dissection molecular mechanisms of tumorigenesis. It has been reported that LKB1 can inhibit the proliferation, invasion, and migration of lung cancer cells.24,25 Faubert et al. 26 showed that LKB1-dependent reprogramming of cell metabolism was dependent on HIF-1α; LKB1 loss increased HIF-1α protein expression in the lung cancer cells. Like HIF-1α, HIF-2α is a transcription factor, and both factors belong to HIF family and share 48% homology. 17 The activation of both factors produces overlapping yet distinct gene expression profiles. In comparison to HIF-1α, HIF-2α also involves in regulation of genes important for tumor growth, cell-cycle progression, and maintaining stem cell pluripotency. However, the regulatory role of LKB1 in HIF-2α protein expression has not been previously reported. In four well-established lung cancer cell lines of different E6 and E7 expression levels, we first proved HPV16 E6 and E7 downregulated LKB1. Then, we bi-directionally manipulated the expression of LKB1 gene by gene transfection or siRNA interference in these cell lines. Our results convincingly showed that LKB1 is a downregulator of HIF-2α (Figure 3(a) and (b)). Our results are the first to illustrate that HIF-2α is a downstream effector of LKB1 and propose a HPV-LKB1-HIF-2α-VEGF axis for the tumorigenesis of lung cancer. It is noticed that in our study, knockdown of LKB1 only affected HIF-2α expression at protein level only but not on mRNA level, suggesting the regulation was probably through translational, post-translational, or even protein degradation mechanisms. Faubert et al. proposed that LKB1 regulated HIF1α by AMP-activated protein kinase (AMPK)-dependent and -independent mechanisms. In the independent mechanism, the LKB1 regulated HIF-1α at both transcriptional and translational levels, while in the dependent mechanism at the translational level only. In the AMPK-dependent pathway, loss of LKB1 disrupted the activity of AMPK, which upregulated mechanistic target of rapamycin (mTOR) activity and subsequently promoted the translation of HIF-1α.27,28 The mechanisms of LKB1 knockdown–induced HIF-2α expression are currently under investigation.
The functional activity of HIF-2α in the four lung cancer cell lines of our study was demonstrated by its capability to enhance VEGF expression in cancer cells. As an important angiogenesis factor, VEGF is critical in the development of tumor vasculature. It is well known that cancer cells have a different metabolism pattern from the normal cells. Even under aerobic conditions, the cancer cells consume more glucose to obtain energy by glycolysis, and this unique metabolic pattern is known as the Warburg effect. 29 The Warburg effect is achieved by regulating the expression of GLUT1, VEGF, 30 erythropoietin, 31 and other survival genes. Thus, VEGF, the major factor in the Warburg effect, plays a central role in response to hypoxia. 32 Jackson et al. 33 believe that VEGF is the key downstream target of the HIF pathway and plays an important role in the initiation and progression of NSCLC. Zhang et al. 34 reported that E6 and E7 upregulated the expression of VEGF at both protein and mRNA levels mediated by HIF-1α in NSCLC. Recently, Zhu et al. 35 found that the expression of VEGF was positively correlated with the expression of HIF-2α in oral squamous cell carcinoma. Both HIF-1α and HIF-2α upregulate VEGF expression. However, only HIF2α upregulated the expression of another angiogenesis factor IL-8. 36 In this study, we bi-directionally manipulated the expression of HIF-2α gene by transfection or siRNA interference in lung cancer cells. As expected, our results showed that HIF-2α upregulated the expression of VEGF at both protein and mRNA levels (Figure 4(a) and (b)). Beside angiogenesis, other functions of HIF-2α, such as cell proliferation, resistant to apoptosis, and metabolic reprogramming, are being further investigated.
In conclusion, we demonstrated that HPV16 E6/E7 inhibited the expression of LKB1 in several lung cancer cell lines and the downregulation of LKB1 increased the expression of HIF-2α and subsequent VEGF. Our results propose a HPV-LKB1-HIF-2α-VEGF axis for the tumorigenesis of lung cancer. These findings suggest new therapeutic targets for HPV-related lung cancers.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the National Natural Science Foundation of China to Guang-Ping Wu (Grant Nos 81171650 and 81672082).
