Abstract
The mechanism of dsRNA-induced gene activation (RNAa) is being gradually unveiled. The plentiful evidence that it existed in mammalian species other than human demonstrated that dsRNA-mediated RNAa is a conservative phenomenon. Simultaneously, accumulating evidence suggested that microRNAs could activate gene expression by targeting promoter. Nevertheless, it is ambiguous whether microRNA-induced gene activation in different human cells is a common phenomenon. The study we performed verified that miR-1236-3p (miR-1236) and miR-370-5p can activate p21 expression in bladder cancer (BCa) T24, EJ cells, and non-small-cell lung carcinoma A549 cells, while in hepatocellular HepG2 cells both microRNAs cannot effectively induce the expression of P21WAF1/CIP1 (p21). In pancreatic cancer PANC-1 cells, only miR-370-5p had the potent abilities to induce p21 expression rather than miR-1236-3p. Unlike microRNA-mediated RNA activation, we can observe that dsP21-322 significantly activated p21 in above cells. Besides, we demonstrated that miR-1236 and miR-370 inhibited cyclin D1-CDK4/CDK6 pathway while upregulated E-cadherin expression by upregulation of p21. Overexpression of these two microRNAs in A549 induced cell-cycle arrest and cell senescence, delayed cell proliferation and colony formation, and inhibited migration and invasion. In conclusion, microRNA-mediated RNAa depends on the cell context, and miR-1236 and miR-370 can inhibit non-small-cell lung carcinoma cell growth by upregulating p21 expression in vitro.
Introduction
RNA activation (RNAa) is a phenomenon that small activating RNA (saRNA) stimulated gene expression by targeting promoter at the transcriptional level.1–3 The inchoate mechanism study suggested that Argonatue2 (Ago2) protein and RNA polymerase II (RNAP II) were vital for RNAa action. 4 Recently, researches suggested that saRNA guided Ago2 targeting specific promoter region to facilitate the assembly of RNA-induced transcriptional activation complex and then binding to the specific promoter regions in a seed-region-dependent manner to promote gene expression.5,6 With the development of the RNAa mechanism, more and more people are engaged in RNAa research. Increasing evidence demonstrated that the phenomenon of RNAa was not only present in human but also in other mammals, even in Caenorhabditis elegans.7–9 Meanwhile, similar findings on RNAa for more genes were reported.10–13 With the in-depth study of RNAa, a consensus has been reached that RNAa is a general mechanism for gene regulation.
MicroRNAs (miRNAs) are an abundant group of small endogenous non-coding RNAs composed of approximately 22 nucleotides. 14 Evidence suggested that miRNAs implicate in a series of cellular processes, including cell proliferation, differentiation, and metastasis. 15 Our initial awareness is that miRNAs inhibit gene expression by binding to messenger RNA (mRNA) and thus lead to the degradation of mRNA or inhibition of translation.16–18 However, accumulating evidence suggested that miRNAs hold the ability to activate gene expression by targeting promoters.2,19–21 Place et al. reported that miR-373 can induce E-cadherin expression with complementary promoter sequence. Recent studies have also reported that miR-6734 and miR-3619 had the capacity to activate p21 gene expression in breast cancer (BC) and colon cancer, respectively. However, little is known about the mechanism of miRNA-mediated gene activation.
Lung cancer is the most common cause of cancer mortality worldwide. Non-small-cell lung carcinoma (NSCLC) is the predominant form of lung cancer. In China, lung cancer causes approximately 23.03% of all new cases of cancer death in men. Moreover, with the development of environmental problems, lung cancer has become an increasingly serious threat to public health. 22 P21WAF1/CIP1 (p21), a well-known cell-cycle inhibitor, can arrest cell-cycle progression through binding and inhibiting cyclin–cyclin-dependent kinase (CDK) complexes. 23 Recently, studies also showed that p21 prevented tumor invasion and migration by inhibiting epithelial–mesenchymal transition (EMT). 24 p21 was demonstrated to be activated by dsP21-322 in an RNAa manner and thus plays an anti-tumor role in amount of human tumors.25–27 In previous study, we found that three miRNAs (miR-1236, miR-370, and miR-1180) can upregulate p21 gene expression in bladder cancer through binding p21 promoter. 19 However, whether miRNAs can upregulate p21 expression in diverse human cancer cells is unclear. Besides, whether miR-1236 and miR-370 can inhibit lung cancer and by regulating which genes also need further exploration.
Materials and methods
MiRNAs and dsRNAs
The miRNAs targeting the p21 gene promoter used in this experiment were predicted by our previous study. 19 MiR-1236-3p mimics, miR-370-5p mimics, and dsRNAs were purchased from RiboBio (China). dsP21-322 was used to activate p21 expression (positive control), and dsControl without significant homology to all known human sequences was used as a nonspecific negative control. siP21 was used to silence p21 expression. 2 All the custom saRNAs sequences are listed in Supplementary Table S1.
Cell culture and transfection
All the cell lines were from American Type Culture Collection (ATCC). EJ, A549, BEAS-2B, and L-02 were maintained in RPMI 1640 medium (HyClone, USA) supplemented with 10% fetal bovine serum (FBS) and 2 mmol/L
Real-time quantitative polymerase chain reaction
Total cellular RNA in cells was extracted using TRIzol reagent (Invitrogen). Then, the reverse transcription reaction was executed with 500 ng of RNA using a complementary DNA (cDNA) Synthesis Kit (TaKaRa, China). Real-time quantitative polymerase chain reaction (RT-qPCR) was conducted using SYBR Premix Ex TaqTM II (TaKaRa) according to the manufacturer’s instructions on the ABI Prism 7300 system (Applied Biosystems). To quantify mature miR-1236 and miR-370 expression, miRNAs were reverse transcribed using the stem-loop primers and then quantitated. Amplification of glyceraldehyde 3-phosphate dehydrogenase (GAPDH) served as internal control for p21 and related gene and U6 were used as an endogenous control for miRNAs. Each measure was done in triple replication and expression change of targeted genes was quantified using 2−ΔΔCt method. All primers used in this study were synthesized by Invitrogen (China). The sequences are available in Supplementary Table S2.
Protein extraction and western blot analysis
Proteins were extracted using NP40 buffer (Beyotime Institute of Biotechnology, China) supplemented with protease inhibitor phenylmethanesulfonyl fluoride (PMSF; Roche, Switzerland). The protein concentrations were calculated using the bicinchoninic acid (BCA) protein assay kit (Beyotime Institute of Biotechnology, China). Equivalent amounts of 30 µg protein samples were isolated using 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). Then, western blot analysis was performed following a standard procedure with the primary antibodies: anti-p21 (1:1000; Cell Signaling Technology, USA), anti-cyclin D1 (1:500; Boster, China), anti-CDK4 (1:500; Boster), anti-CDK6 (1:500; Boster), and anti-E-cadherin (1:1000; Cell Signaling Technology). The anti-β-Actin (Boster) was used as an internal control at 1:500 dilution. And then, protein bands were incubated with secondary antibodies (1:5000) conjugated with horseradish peroxidase (HRP) and visualized using the enhanced chemiluminescence (ECL) assay kit.
Cell-cycle analysis by flow cytometric
Flow cytometric analysis was performed to detect cell-cycle distribution. After 72 h of transfection of the corresponding miRNA or dsRNA, 1 × 106 cells were harvested and fixed using 70% cold ethanol at 4°C overnight. Then, cells were washed twice with cold phosphate-buffered saline (PBS) and incubated with RNaseA (Sigma, USA) at 37°C. After 30 min incubation, cells were resuspended with propidium iodide (PI; KeyGen Biotech, China) at 4°C for 30 min. Then, samples were analyzed on a FACSort flow cytometer (BD Biosciences, USA) for relative cellular DNA content, and data were interpreted using CellQuest software (BD Biosciences).
Senescence-associated β-galactosidase staining
When cells are in the aging state, their aging-related galactosidase activity levels are elevated. Thus, we detected senescence using a senescence-associated β-galactosidase assay (Sigma). Cells were transfected with the corresponding miRNA or dsRNA in a six-well plate for 72 h. After washing with cold PBS buffer twice, cells were fixed in the room temperature for 15 min. Then, cells were stained at 37°C overnight and photomicrographs were captured using an ordinary inverted phase-contrast microscope.
Cell proliferation assay
Cell proliferation was detected by Cell Counting Kit-8 (CCK-8; Beyotime Institute of Biotechnology, China). Briefly, 2000 cells were seeded into a 96-well plate and cell growth was measured after transfection at 5 time points from day 1 to day 5. At each time point, culture medium was replaced by 100 µL of the liquid containing 10 µL CCK-8 premixed with 90 µL fresh medium and incubated at an atmosphere of 5% CO2 at 37°C for 2 h. Determination of absorbance was measured with enzyme-labeled instrument (Thermo Fisher Scientific, USA) at 450 nm.
Colony formation assay
After transfection for 24 h, 1000 A549 cells were reseeded into new six-well plates with growth medium. After 10 days of continuous culture, the colonies were fixed and stained with 0.5% crystal (Sigma) at room temperature. After 30 min, each group of cell colonies was counted. The colony formation rate was calculated by the following equation: colony formation rate = number of colonies/number of seeded cells.
Transwell assay
Cell migration and invasion potential were evaluated by a Transwell system (24 wells, 8-µm pore size). For Transwell invasion assays, Matrigel (BD Biosciences) was diluted to 0.33 mg/mL concentration with serum-free medium, and 100 µL Matrigel was coated onto upper Transwell inserts and incubated at 37°C for 2 h to make it ready for seeding. After transfection for 72 h, a total of 5 × 105 cells were plated onto the top cell culture insert with serum-free medium, and the lower chamber was filled with 500 µL RPMI 1640 with 10% FBS, used as a chemoattractant. After incubation for 48 h, the cells were fixed with formaldehyde for 10 min. Then, cells were stained with crystal violet, and non-migration cells in the chamber were removed. The invaded cells of lower surface of the membrane were taken pictures by inverted microscope (Olympus, Japan) under 200 times magnification and the number of cells was counted in four randomized visual fields. Except Matrigel-free upper chambers, the procedure of cell migration assays was the same as invasion assays.
Statistical analysis
All data are presented in the form of mean ± standard deviation (SD). Statistical analyses were performed by SPSS version 13.0 software (SPSS Inc., USA) and Graphpad Prism 5.0 software (GraphPad Software, Inc.). The difference between two or more groups was analyzed using a two-tailed student’s t test or one-way analysis of variance (ANOVA) test; p value <0.05 was considered statistically significant.
Results
Unlike dsRNA-mediated RNAa, miRNA-mediated activation of p21 by targeting promoter is depending on the cell context
We previously demonstrated that the expression of miR-1236, miR-370, and p21 was reduced in BCa; furthermore, miR-1236-3p and miR-370-5p may function to induce p21 gene expression by targeting promoter region relative to transcription starting site at -243/-226 and -552/-537 (Figure 1(a)). 19 To confirm whether miR-1236-3p and miR-370-5p can activate p21 gene expression universally in diverse human carcinoma cells, we first compared the relative expression of miR-1236, miR-370, and p21 mRNA in three cancer cells (HepG2 and PANC-1, A549) with their normal tissue cells (L-02 and HPDE, BEAS-2B) using RT-qPCR. As is shown, the expression of miR-1236, miR-370, and p21 mRNA was lower in three cancer cell lines. It should be noted that miR-1236 and p21 mRNA were reduced in PANC-1 cells compared with HPDE cells without statistical significance (Figure 1(b) and (c)).
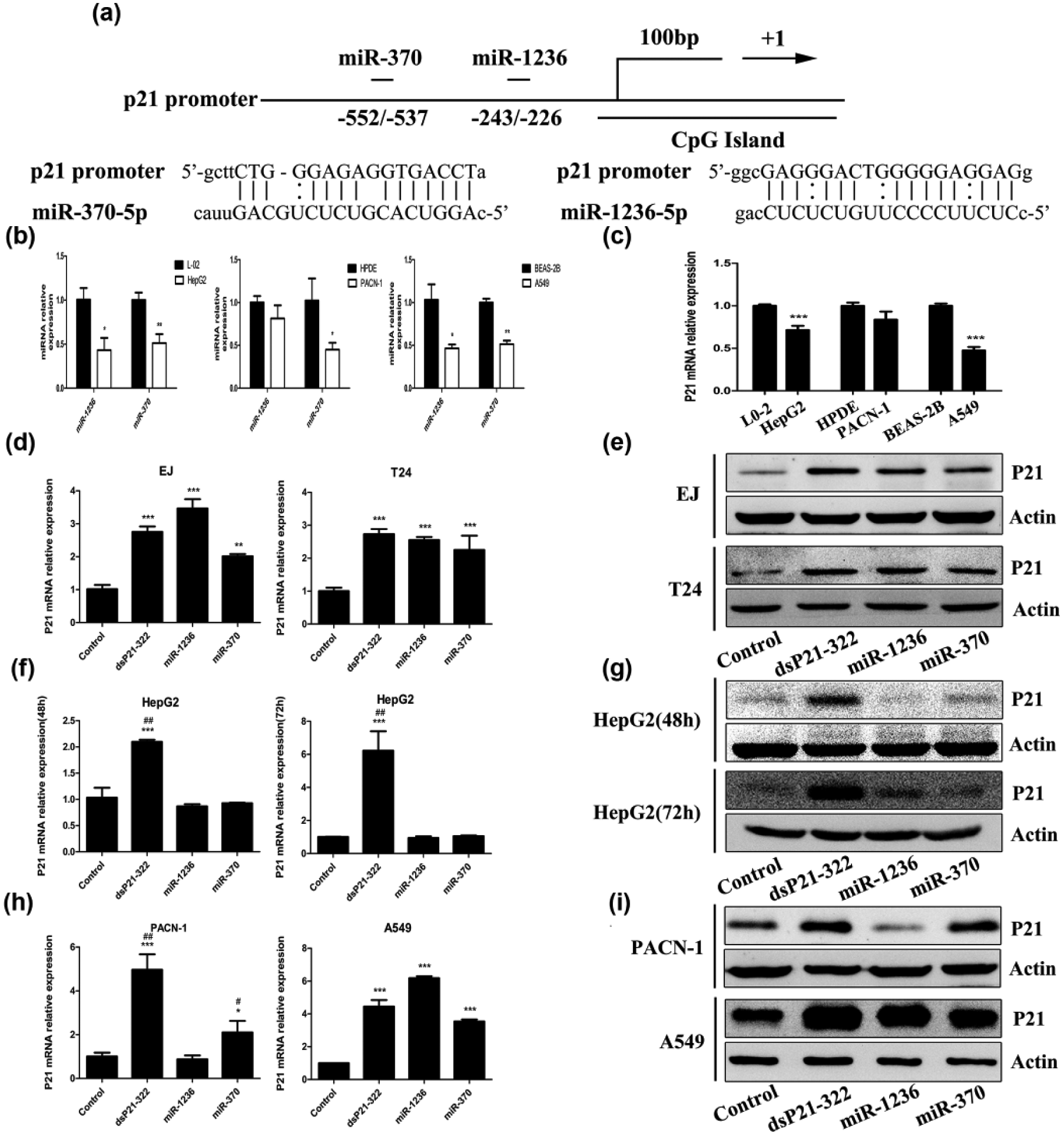
dsRNA and miRNAs showed different effects of p21 activation by targeting promoter in human tumor cells. (a) Schematic representation of the p21 promoter and corresponding miR-1236-3p, miR-370-5p target sites, and the complementary seed sequences. (b) MiR-1236 and miR-370 and (c) p21 mRNA expression levels assessed by RT-qPCR in three human cancer cells (Hepg2, PACN-1, and A549; *p < 0.05, **p < 0.01, and ***p < 0.001 compared to their primary normal tissue cells (L0-2, HPDE, and BEAS-2B), respectively. (d, f, and h) RT-qPCR evaluated p21 and GAPDH mRNA expression. Cells were transfected with 50 nM of indicated miRNAs or dsRNAs for 48 or 72 h. Values of p21 were normalized to GAPDH. (d) p21 mRNA expression of T24 and EJ cells (**p < 0.01 and ***p < 0.001 compared to dsControl). (f) p21 mRNA expression of HepG2 cells for 48 and 72 h (***p < 0.001 compared to dsControl; ##p < 0.01 compared to miR-1236 miR-370). (h) p21 mRNA expression of PACN-1 and A549 cells (*p < 0.05 and ***p < 0.001 compared to dsControl; #p < 0.05 and ##p < 0.01 compared to miR-1236). (e, g, and i) Immunoblotting analysis detected p21 protein expression in cells. β-Actin served as an internal control.
Then, corresponding miRNAs or dsRNAs were transfected into three carcinoma cells for 48 or 72 h. RT-qPCR and immunoblotting were performed to evaluate the expression levels of p21 gene. Compared with dsControl transfections, dsP21-322 effectively upregulated both mRNA and protein expression of p21 gene (Figure 1(d)–(i)). p21 mRNA and protein levels were significantly increased by miR-1236 and miR-370 compared to dsControl in BCa cells (T24 and EJ; Figure 1(d) and (e)), respectively. Surprisingly, both miRNAs failed to increase p21 expression at 48 or 72 h time points in HepG2, including mRNA and protein levels (Figure 1(f) and (g)). The results in Figure 1(h) showed that transfection of miR-1236 and miR-370 caused a 6.18- and 3.54-fold induction of p21 mRNA levels, respectively, in A549 compared to dsControl treatment, while only miR-370 induced a 2.10-fold higher expression of p21 compared to dsControl treatment in PACN-1. Moreover, the induction change of p21 protein was further confirmed by western blot analysis (Figure 1(h)). These data suggest that dsRNA-mediated activation of p21 expression is a common phenomenon in human tumor cells, while miRNA-mediated RNAa of p21 depended on cell context.
MiR-1236-3p and miR-370-5p reversed cyclin D1-CDK4/6 and E-cadherin expression of NSCLC cells
To further investigate the role of miR-1236 and miR-370 in the development of NSCLC by mediating p21 activation, the expression of genes that manipulate cell cycle was examined. In addition, in order to identity whether these miRNAs involve in migration and invasion, we assessed the expression of E-cadherin, an epithelial marker. As is shown, compared to dsControl treatment, positive control (dsP21-322), miR-1236-3p, and miR-370-5p significantly downregulated expression of Cyclin D1 and CDK4/CDK6 mRNA, while E-cadherin mRNA expression level was positively upregulated in A549 (Figure 2(a)). Western blot analysis further confirmed that miR-1236, miR-370, and dsP21-322 inhibited the expression of CyclinD1 and CDK4/CDK6 while promoted the expression of E-cadherin (Figure 2(b)).

MiR-1236-3p and miR-370-5p reversed cyclin D1, CDK4/CDK6, and E-cadherin expression. (a) RT-qPCR detected expression of cyclin D1, CDK4/CDK6, and E-cadherin mRNA after selected miRNA or dsRNAs transfection into A549 for 72 h. GAPDH served as a loading control (*p < 0.05, **p < 0.01, and ***p < 0.0 01 compared to dsControl group). (b) Western blot analysis measured expression of cyclin D1, CDK4/CDK6, and E-cadherin protein after selected miRNA or dsRNAs transfection into A549 for 72 h. β-Actin was used as an internal control.
MiR-1236-3p and miR-370-5p not only induced NSCLC cell-cycle arrest and cell senescence but also retarded cell proliferation and colony formation of NSCLC
Cell-cycle distribution was assessed by performing flow cytometry analysis after transfected with miRNAs or dsRNAs. The results indicated that compared to dsControl group, cells transfected with selected miRNAs presented a prominent accumulation in the GO/G1 population and a consistent decrease in S and G2/M population (Figure 3(a) and (b)). Then, the effect of miRNAs on cell senescence was determined. As is shown in Figure 3(c), positive β-galactosidase cells were observed to increase prominently following transfection of the dsP21-322, miR-1236, and miR-370. Subsequently, we performed a cell proliferation assay to examine the inhibitory effect of miRNAs on cell proliferation. Indeed, we measured a retarded growth from day 2 to day 5 following transfection (Figure 3(d)). For the purpose of better observing the growth inhibitory effect of both miRNAs, colony formation assay was carried out. Apparently, two miRNAs and dsP21-322 were obviously suppressed the number of colonies (Figure 3(e)). Besides, the colony formation rates of cells transfected with miRNAs and dsP21-322 were markedly reduced compared to the dsControl group (Figure 3(f)). These results suggested that miR-1236 and miR-370 caused cell-cycle arrest and inhibited lung cancer cell growth. This blocking function is consistent with previous report. 28

Overexpression of miRNAs not only induced NSCLC cell-cycle arrest and cell senescence but also retarded cell proliferation and colony formation of NSCLC. A549 cells were transfected with the indicated miRNA or dsRNAs for 72 h. (a) Representative flowcytometry images of cell-cycle distribution. (b) Percentage of cells in G0/G1-, G2/M- and S-phase was shown by Percentage Stacked Column Chart (*p < 0.05, **p < 0.01, and ***p < 0.001 compared to dsControl group at the same cell cycle phase). (c) β-galactosidase assay estimated cell senescence (400×). (d) Cell proliferation was assessed by CCK-8 assay. Results were plotted as OD values (*p < 0.05, **p < 0.01, and ***p < 0.001 compared to dsControl group at the same time point). (e) Representative photographs of colony formation assay. (f) Quantification of the cell colony formation (**p < 0.01 compared to dsControl group).
MiR-1236-3p and miR-370-5p inhibited migration and invasion of NSCLC cells
To evaluate whether the selected miRNAs have the potential role in inhibiting migration and invasion by inducing p21 and E-cadherin expression in NSCLC cells, migration and invasion assays were conducted after artificial expression of miRNAs in A549; 72 h after transfection, cells were reseeded in the upper Transwell inserts and it contained pores through which cells were allowed to migrate into complete media. As expected, the number of cells of over-expression of dsP21-322, miR-1236, and miR-370 passing through the pores were significantly reduced (Figure 4(a)). Similarly, for miR-1236, miR-370, and dsP21-322 transfected cells, relative migration and invasion rate are lower compared with the dsControl group (Figure 4(b) and (c)).

MiRNAs inhibited NSCLC cell migration and invasion. The effects of miR-1236 and miR-370 overexpression on migration and invasion of A549 cells were evaluated by Transwell assays. (a) Representative pictures of Transwell assay (200×). (b and c) Numbers of invading cells through the Matrigel and membrane were quantified in four random fields of each group. Results are presented as a percent (%) compared to dsControl group (**p < 0.01 compared to dsControl group).
MiR-1236-3p and miR-370-5p manipulated cyclin D1-CDK4/6 and E-cadherin expression by activating p21
Recent studies have reported that miR-1236 and miR-370 play an anti-tumor effect in lung cancer by targeting HDAC3 and TRAF4, respectively.28,29 To validate that miR-1236 and miR-370 regulate cyclin D1, CDK4/CKD6, and E-cadherin expression mainly by activating p21, a small interfering RNA (siP21) was transfected into A549 to knockdown the expression of p21. 30 The results suggested that co-transfection of miR-1236 or miR-370 and siP21 significantly inhibited the capability of activating p21 mRNA and protein expression triggered by miR-1236 or miR-370 (Figure 5(a) and (b)). Furthermore, the expression of downstream genes was detected in A549. As is shown in Figure 5(a), miR-1236 or miR-370 lost the function of downregulating cyclin D1 and CDK4/CDK6 but upregulated E-cadherin. This was further demonstrated by western blot analysis (Figure 5(b)).

MiRNAs manipulated Cyclin D1-CDK4/6 and E-cadherin expression by activating p21. (a) RT-qPCR measured expression of p21 mRNA in A549 cells co-transfected indicated miRNAs and siP21. Values of p21 were normalized to GAPDH. (b) Expression of p21 protein in A549 cells co-transfected indicated miRNA with siP21was detected by western blot analysis. β-actin served as an internal control. (c) RT-qPCR and (d) western blot analysis detected expression of cyclin D1, CDK4/CDK6, and E-cadherin in A549 cells co-transfected indicated miRNA with siP21 (*p < 0.05, **p < 0.01, and ***p < 0.001 compared to miR-1236-3p group; #p < 0.05 and ###p < 0.001 compared to miR-370-5p group).
MiR-1236-3p and miR-370-5p inhibited NSCLC cells by upregulating p21 expression
Next, we evaluated whether the inhibitory function of miR-1236 and miR370 in A549 could be influenced by silencing p21. The results of flow cytometry analysis suggested that co-transfection of miR-1236 or miR-370 with siP21 remarkably attenuated Go/G1 cycle arrest (Figure 6(a) and (b)). As expected, silencing p21 dramatically inhibited miR-1236 and miR-370 induced cell senescence (Figure 6(c)). Additionally, the ability of proliferation and colony formation were restored by p21 silencing (Figure 6(d)–(f)). Moreover, the abilities of miR-1236-3p and miR-370-5p to inhibit cell migration and invasion were evidently alleviated by depleting p21 (Figure 7(a)–(c)). Taken together, miR-1236 and miR-370 play an anti-tumor effect in NSCLC cells mainly through modulating p21 expression
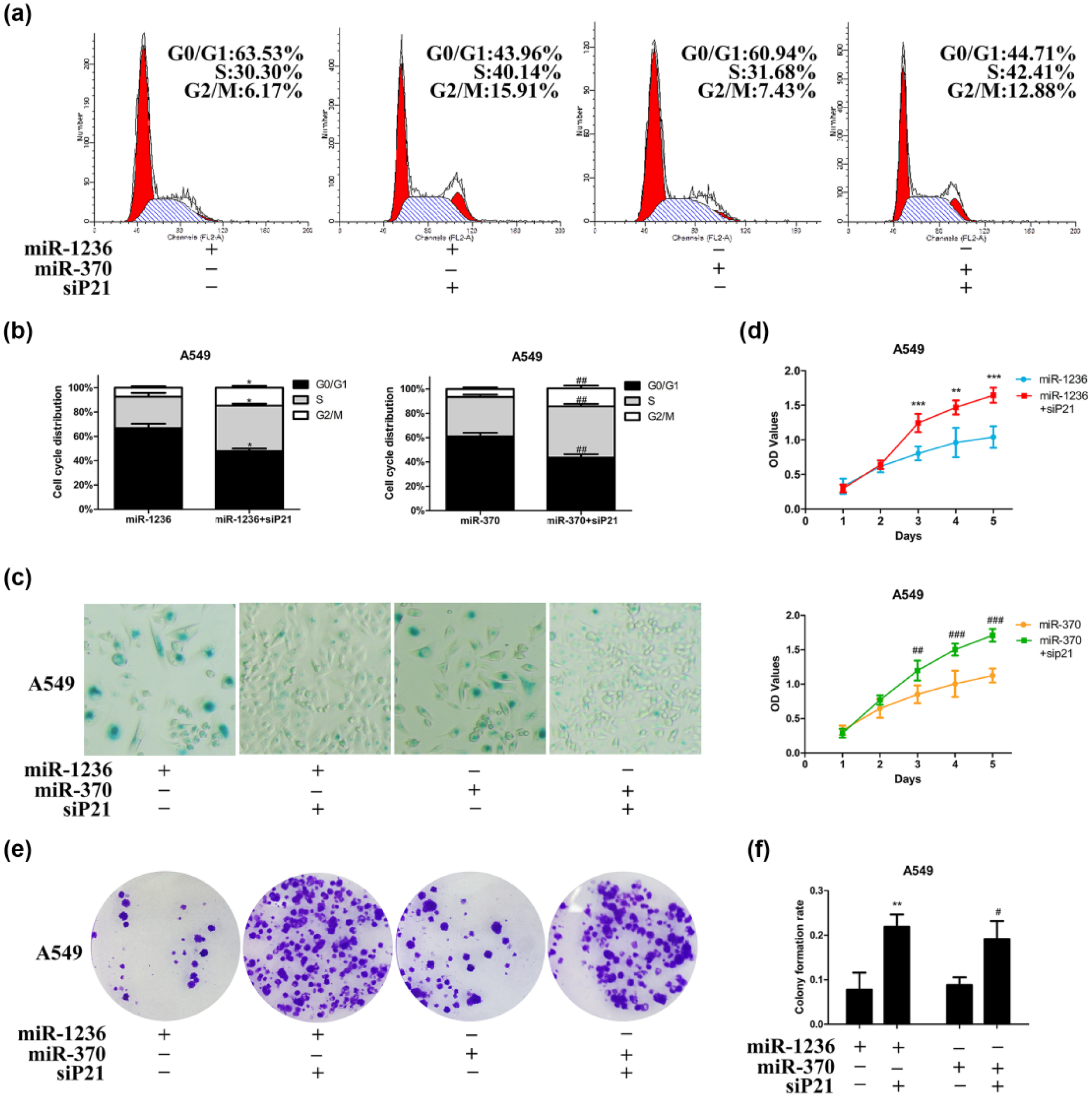
Overexpression of miRNAs inhibited NSCLC cells’ growth mainly by upregulating p21. A549 cells were co-transfected indicated miRNAs and siP21 for 72 h. (a) Representative flowcytometry images of cell-cycle distribution. (b) Percentage of cells in G0/G1-, G2/M- and S-phase was shown by Percentage Stacked Column Chart (*p < 0.05 compared to miR-1236-3p group and ##p < 0.01 compared to miR-370-5p group at the same cell-cycle phase). (c) Cell senescence was estimated by β-galactosidase assay (400×). (d) Cell proliferation was assessed by CCK-8 assay. Results were plotted as OD values (**p < 0.01 and ***p < 0.001 compared to miR-1236-3p group and ##p < 0.01 and ###p < 0.001 compared to miR-370-5p group at the same time point). (e) Representative pictures of colony formation assay. (f) Quantification of the cell colony formation (**p < 0.01 compared to miR-1236-3p group and #p < 0.0 compared to miR-370-5p).

MiRNAs inhibited NSCLC cell migration and invasion primarily via enhancing p21. Transwell assays evaluated migration and invasion of A549 cells co-transfection indicating miRNAs with siP21. (a) Representative pictures of Transwell assays (200×). (b and c) Numbers of invading cells through the Matrigel and membrane were quantified in four random fields of each group. Results are presented as a percent (%) compared to miR-370-5p or miR-1236 group (**p < 0.01 compared to miR-1236-3p; ##p < 0.01 compared to miR-370-5p).
Discussion
Since the discovery of the first miRNA for over 20 years, miRNA has been extensively studied. MiRNA therapy offers a promising new weapon for cancer and other diseases. 31 The discovery of miR-373 by targeting its promoter to increase the expression of E-cadherin provides us with a new insight into miRNA-targeted tumor therapy. 2 MiRNA-induced RNAa has shown a new promise in inhabiting tumor development. However, whether miRNAs can stably induce RNAa in different cells is unknown. In the present research, we detected that both expression of miR-1236-3p and miR-370-5p decreased in three cancer cells (HepG2, PACN-1, and A549) compared with their normal cell lines.28,32,33 Similarly, the expression of p21 mRNA was also reduced in three tumor cells. In addition, we demonstrated that overexpression of miR-1236-3p and miR-370-5p can upregulate p21 expression in BCa cells and NSCLC cells. It should be indicated that both selected miRNAs cannot activate p21 expression in hepatocellular carcinoma (HCC) cells and only miR-370-5p possessed the ability to induce p21 expression in pancreatic carcinoma cells. Nevertheless, we can observe that dsP21-322 significantly activates p21 in the above cells. We can boldly conclude that unlike the common phenomenon of dsRNA-mediated RNAa in diverse human cells, miRNA-mediated gene activation depends on the cell context.
Earlier study showed that positive p21 expression was a standalone prognostic factor and upregulation of p21 expression can improve the sensitivity of NSCLC cells to chemotherapeutic drugs.26,34 Malignant tumors are characterized by infinite reproduction, invasion, and metastasis. As is well-known, p21 protein can induce cell-cycle arrest in G0/G1-to-S phase through restraining cyclin–CDK complex activity. 30 In this study, we identified that miR-1236 and miR-370 inhibited cyclinD1-CDK4/CDK6 pathway by activating p21, resulting in cell accumulation G0/G1 phase in NSCLC A549 cells. Furthermore, we demonstrated that miR-1236 and miR-370 can promote cell senescence and inhibit cell cloning and proliferation in A549. Invasion and metastasis are often associated with poor prognosis. EMT is a process, whereby expression levels of the epithelial markers are reduced, and mesenchymal markers are increased to increase cell migration and invasion activity. 29 After transfection of miR-1236 and miR-370 in A549, the expression of E-cadherin—the EMT epithelial cell marker—was up-regulated, and retardation of migration and invasion were observed. Obviously, the anti-tumor effect of these two miRNAs was primarily realized by modulating p21 expression.
As everyone knows, Ago2 is vital for dsRNA-mediated RNAa. Ago2 can associate with either promoter DNA or promoter transcripts after being recruited and binding to target region under the guidance of the antisense chain. Lines of dependent researches have suggested that the knockdown of Ago2 could abrogate RNAa.35–37 However, Ago1 took an important role in miRNA-mediated induction of Ccnb1 expression than Ago2. ChIP analysis showed only a significantly enrichment of Ago1 protein in the CCB1 promoter region. 38 Mescalchin et al. reported that Ago2 was a key protein on small-interfering RNA (siRNA)-mediated gene silencing, while the primary participant of miRNA-mediated gene regulation was Ago1 and other family members. 39 So, we have reason to guess that mechanism of RNAa mediated by endogenic miRNA was distinct from synthetic dsRNA duplexes. This may be an important factor in miRNA-mediated activation which is different from dsRNA-mediated activation. More researches on the mechanisms of miRNA-mediated RNAa are required.
MiRNA acts as tumor suppressor or oncogene, depending on cell context. MiR-431 played a carcinogenic role in colorectal cancer, while miR-431 acted as a tumor suppressor by modulating (EMT) signaling pathway in HCC. 40 Plentiful of studies documented that miR-373 restrained target mRNA and/or degraded target mRNA. In contrast, indisputable evidence verified it could also upregulate gene expression. 41 In this study, our results showed that miR-1236 and miR-370 stimulate the expression of p21 in lung cancer cells and played an anti-tumor effect, but in hepatocarcinoma cells they had no ability to activate p21. It suggested that miR-1236 and miR-370 play a different role in different human tumors. This is consistent with recent studies that miR-370 motivated p21 expression to suppress BCa cells metastasis. In contrast, it through mediating p21 downregulation contributed to the progression of gastric carcinoma.25,42 The tumor-specificity of miRNAs warns us that we should undergo relevant in vitro/vivo experiments before carrying out miRNA therapy.
To sum up, we demonstrate that unlike the universality of dsRNA-mediated RNAa, miRNA-mediated RNAa depends on the cell context. In addition, miR-1236 miR-370 inhibited NSCLC cell proliferation, migration, and invasion via regulating p21 expression. Whereas further research is needed to identify other examples and to clarify the mechanisms by which miRNAs activate gene expression.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (grant number 81372759, China). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
