Abstract
Several studies have revealed that MFG-E8 (milk fat globule–epidermal growth factor 8) is related to tumour development and progression. However, the relationship between MFG-E8 expression and metastasis in colorectal cancer patients and the role of MFG-E8 in colorectal cancer invasion and progression remain unknown. In this study, we performed immunohistochemistry and quantitative real-time polymerase chain reaction to assess MFG-E8 expression in colorectal cancer and adjacent non-cancerous tissues. Colorectal cancer RNAseq data from The Cancer Genome Atlas project were downloaded and MFG-E8 expression was analysed. Gene set enrichment analysis was performed for gene ontology and pathway analysis associated with MFG-E8 expression. For in vitro studies, we used lentivirus-mediated MFG-E8 RNA interference and commercialized recombinant human MFG-E8 to investigate its role in colorectal cancer cell growth, migration and invasion. It seems that MFG-E8 was overexpressed in advanced colorectal cancer tissues compared with early-stage colorectal cancer tissues and adjacent non-cancerous tissues. Correlation analysis revealed that MFG-E8 expression was significantly related to plasma membrane invasion, lymph node metastasis, distant metastasis and tumour–node–metastasis stage. Survival analysis revealed that high MFG-E8 expression predicted a poorer prognosis than low MFG-E8 expression group both in our colorectal cancer cohort and The Cancer Genome Atlas colorectal cancer cohort. In vitro study suggested that MFG-E8 knockdown can suppress the growth of colorectal cancer cells without affecting the expression of the proliferation-related gene Ki67. MFG-E8 knockdown also suppressed colorectal cancer cell migration and invasion, a change accompanied by MMP-2 and MMP-9 downregulation. Moreover, MFG-E8 knockdown induced a shift from mesenchymal makers to epithelial makers, while pretreatment with rhMFG-E8 had the opposite effect. The effect of MFG-E8 on colorectal cancer cell migration, invasion and epithelial-to-mesenchymal was partially dependent on the PI3K/AKT signalling pathway. These findings provide a better understanding of the molecular mechanism underlying colorectal cancer progression and suggest a predictive role for MFG-E8 in colorectal cancer metastasis and prognosis.
Keywords
Introduction
Colorectal cancer is the third most commonly diagnosed cancer in males and the second most commonly diagnosed cancer in females. An estimated 1.4 million cases were diagnosed, and 693,900 deaths occurred in 2012. 1 Approximately 20% of patients with colorectal cancer (CRC) are found to have metastases at the time of diagnosis, and up to 50% of all patients will develop metastases over the course of their disease. 2 The 5-year survival rate for patients with stage I CRC is greater than 90%, but the rate for patients with stage IV CRC with distant metastases is only slightly higher than 10%. 3
Metastasis is a major concern in solid tumours, as this phenomenon is closely associated with a poor prognosis and is responsible for the majority of cancer-related deaths. 4 Invasion and epithelial-to-mesenchymal transition (EMT) are the primary processes involved in metastasis. Matrix metalloproteinases (MMPs) play an indispensable role in this process because of their ability to degrade the extracellular matrix and basement membranes. 5 EMT activation has been proposed as the critical mechanism underlying the acquisition of malignant phenotypes by epithelial cancer cells, as this change substantially facilitates metastasis. 6 Since there are no effective strategies for preventing metastasis in cancer, uncovering the mechanism underlying CRC metastasis may facilitate the development of new therapies for and reduce the mortality of the disease.
Milk fat globule–epidermal growth factor 8 (MFG-E8) also known as lactadherin/BA46/SED1 is a secreted glycoprotein that was initially identified as a principle component of mammalian milk fat globule membrane (MFGM).7–9 It can bind exposed phosphatidylserines on apoptotic cells and αVβ3/5 integrins on macrophages to facilitate apoptotic cell engulfment.9–11 Some patients suffering from the autoimmune disease systemic lupus erythaematosus have elevated serum levels of MFG-E8. 10 Many studies have revealed that a relationship exists between MFG-E8 and tumour progression, and their findings have suggested that MFG-E8 functions as an oncogene in tumour development. MFG-E8 has been reported to be involved in the development of several malignancies, including human melanoma, bladder tumours, breast cancer, prostate cancer and glioblastoma,12–16 and participates in the processes of tumour growth, invasion, apoptosis, angiogenesis and EMT in these malignancies. Regarding CRC, Jinushi et al. 17 reported that the plasma levels of MFG-E8 in CRC patients who underwent complete resection were significantly lower than those in CRC patients with unresectable disease. Kusunoki et al. 18 showed that MFG-E8 gene knockout led to tumour growth suppression in azoxymethane injection–induced sporadic murine colon cancer models. Moreover, Kusunoki also showed that MFG-E8 overexpression in CRC tissues was positively correlated with clinical stages. Although the results of these previous studies strongly suggest that MFG-E8 functions as an oncogene in CRC progression, the correlation between MFG-E8 and metastasis in CRC patients has not been fully investigated, and the role of MFG-E8 in CRC invasion and progression remains to be elucidated.
In this study, we assessed MFG-E8 expression at the messenger RNA (mRNA) and protein levels in CRC tissues and analysed the relationship between MFG-E8 expression and metastasis in CRC patients. We also downloaded CRC RNAseq data from The Caner Genome Atlas (TCGA) database and performed gene ontology and pathway analysis through gene set enrichment analysis (GSEA). The correlation between MFG-E8 expression and clinicopathological parameters in CRC patients is suggestive of its involvement in CRC invasion and progression. In our in vitro study, we demonstrated that MFG-E8 can promote CRC cell migration, invasion and EMT in part via the phosphatidylinositol 3-kinase (PI3K)/AKT pathway.
Materials and methods
Tissue samples
A total of 78 cancerous and 58 adjacent non-cancerous specimens (5 cm distant from the tumour) were collected from 78 CRC patients who had undergone surgery at the General Surgery Department of Qilu Hospital of Shandong University from 2009 to 2011. The specimens were immediately fixed in 10% neutral formaldehyde for further histological study. Moreover, 26 paired CRC cancer and adjacent non-cancerous tissues were obtained during colonoscopy at the Digestive Endoscopy Center of Qilu Hospital of Shandong University. These samples were snap-frozen in liquid nitrogen and then stored at −80°C for RNA extraction. The diagnoses of all the patients were confirmed as CRC on routine histology by two experienced pathologist, and the adjacent normal biopsies were all confirmed to be histologically normal. None of the patients underwent chemotherapy or received any other treatments before sampling. This study was approved by the Clinical Ethics Committee of Qilu Hospital of Shandong University, and all the patients provided written informed consent before sampling. Patient clinicopathological information was collected and is provided in Table 1. Tumour staging was based on patient pathological findings, which were assessed according to the guidelines of American Joint Committee on Cancer (AJCC). 19
Correlation between MFG-E8 expression and clinicopathological parameters in CRC patients.

MFG-E8: milk fat globule–epidermal growth factor 8; TNM: tumour–node–metastasis; AJCC: American Joint Committee on Cancer; CRC: colorectal cancer.
Y: Yate’s continuity-corrected chi-square test. Other data were obtained via regular chi-square testing unless otherwise indicated.
p < 0.05, **p < 0.01 and ***p < 0.0001.
Immunohistochemistry
Patient biopsy samples were embedded in paraffin and cut into 3-µm-thick sections. Antigen retrieval was carried out in Tris-EDTA buffer, pH 9.0 (Dako Corporation, Carpinteria, CA, USA), after deparaffinization and rehydration. Then, 3% H2O2 (Dako Corporation) was applied to block endogenous peroxidase. Mouse anti-human MFG-E8 monoclonal antibody, diluted 1:1000 (Santa Cruz Biotechnology, Santa Cruz, CA, USA), was added to the samples, which were incubated at 4°C overnight. An EnVision™ System (Dako Corporation) was employed for immunohistochemical analysis the next day, and 3′3-diaminobenzidine (Dako Corporation) was used as a chromogen for visualization. Cell nuclei were counterstained with haematoxylin. Images were photographed under an Olympus BX51 microscope (Olympus Corporation, Tokyo, Japan), and five independent representative views were chosen for each slide. Staining intensity was scored as 0 point (negative staining), 1 point (weak staining), 2 points(moderate staining) and 3 points(strong staining), and staining extent was scored as 0 point (<5%), 1 point (5%–25%), 2 points (26%–50%), 3 points (51%–75%) and 4 points (76%–100%), according to the percentage of positively stained cells. The final MFG-E8 expression level was semi-quantitatively evaluated as the product of the staining density and extent scores and was graded on a scale ranging from 0 to 12 points. The specimens were categorized into the following two groups: low (0–6 points) and high (7–12 points), according to their final scores.
Quantitative real-time polymerase chain reaction
Total RNA from specimens or cultured cells was isolated with TRIzol (Invitrogen, San Diego, CA, USA) reagent according to the manufacturer’s instructions. Total RNA was first reverse-transcribed into complementary DNA (cDNA) in the presence of Moloney murine leukaemia virus (M-MLV) (Invitrogen). The primers for human MFG-E8 were designed according to the human MFG-E8 sequence (NCBI GenBank Accession Number NC_000015.10) using the Primer Premier 5.0 software (Premier Biosoft, Palo Alto, CA, USA). The sequences of all the primers used in this study are listed in Table 2. SYBY Green Reagent (TaKaRa, Japan) was employed in quantitative real-time PCR in a fluorescence thermo-cycler (LightCycler; Roche Diagnostics GmbH, Mannheim, Germany). Human β-actin was used as an internal control. The expression levels of each target gene were determined according to the 2−ΔΔCt method, as previously described. 20 Each experiment was performed in triplicate, and the data were presented as mean ± standard error of mean (SEM).
Selected gene sets enriched for genes significantly associated with MFG-E8 expression determined by GSEA.
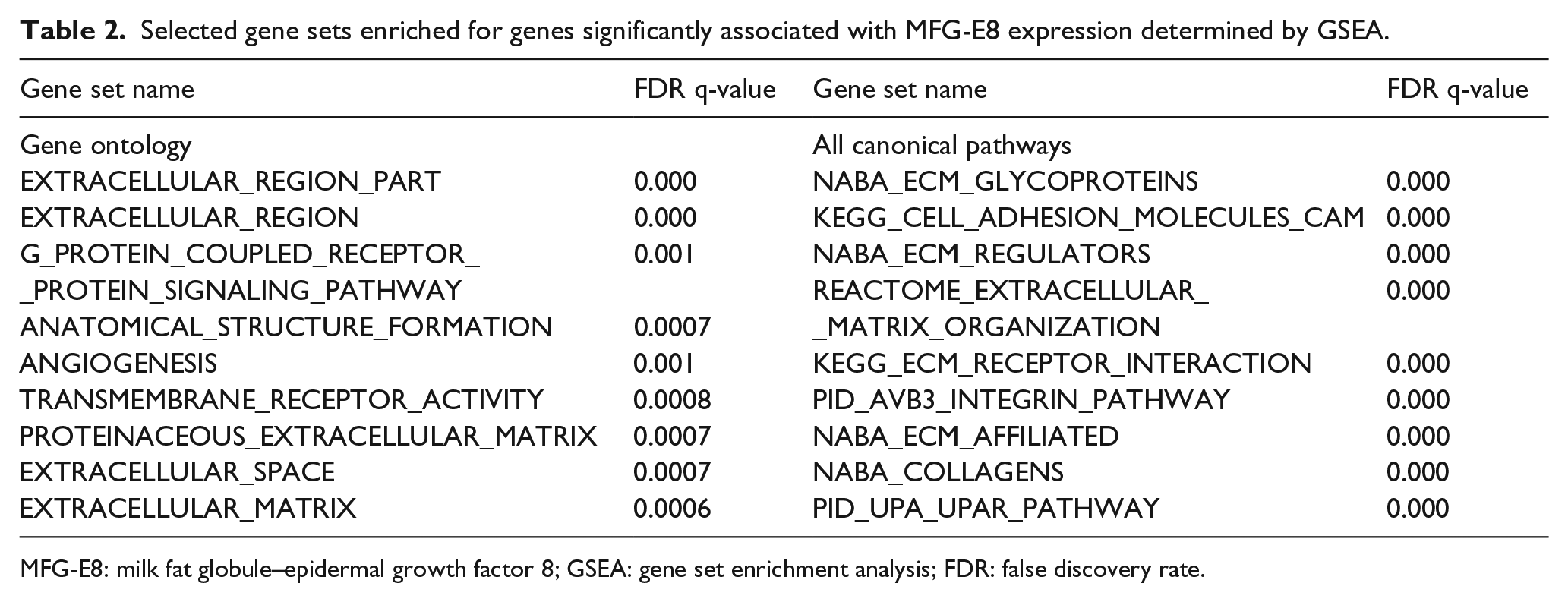
MFG-E8: milk fat globule–epidermal growth factor 8; GSEA: gene set enrichment analysis; FDR: false discovery rate.
TCGA database source and GSEA
The level-3 data of CRC RNAseq including 647 CRC specimens and 51 paratumoural normal specimens and the corresponding clinical data were downloaded from the TCGA data portal (https://portal.gdc.cancer.gov) on 10 September 2016 and processed with R. Specimens were ranked in list according to MFG-E8 mRNA levels and GSEA was performed to associate gene signature with MFG-E8 expression using the Broad Institute Tool (http://www.broadinstitute.org/gsea). A set of 6166 gene sets from the entire MSig database was used for the analysis of Gene Ontology (http://software.broadinstitute.org/gsea/msigdb/index.jsp). 21 For the analysis of pathways, all canonical pathways including 326 gene sets were applied. Gene sets with enrichment false discovery rate (FDR) q-value less than 0.05 are considered significantly enriched.
Cell culture and lentivirus-mediated MFG-E8 knockdown
The human CRC cell lines HT-29 and Caco-2 were obtained from American Type Culture Collection (Manassas, VA, USA) and cultured in Dulbecco’s modified Eagle’s medium (DMEM; Gibco by Invitrogen) supplemented with 10% (v/v) foetal calf serum (FCS; Gibco) at 37°C in a humidified incubator containing 5% CO2. Lentivirus transfection was performed, and MFG-E8 knockdown was validated in the above CRC cell lines, as described in detail in our previous study. 22 Briefly, the cells were transfected with lentiviral particles encoding short hairpin RNA (shRNA) sequences targeted to human MFG-E8 mRNA (5′-CAACCACTGTGAGAC GAAA-3′, LV-M). Lentiviral particles encoding a scrambled shRNA sequence (5′-TTCTCCGAAC GTGTC ACGT-3′, LV-C) were used as negative controls. The lentiviruses were purchased from Santa Cruz Biotechnology, and all of them carried a puromycin-resistant gene. Cell clones showing stable transfection were selected with puromycin (Santa Cruz Biotechnology) and expanded for subsequent experiments.
Western blotting
Total protein was extracted in radioimmunoprecipitation assay (RIPA) buffer (Beyotime Institute of Biotechnology, Shanghai, China). Protein concentrations were quantified by bicinchoninic acid (BCA) assay (Beyotime Institute of Biotechnology). Western blotting was performed as previously described. 22 The following primary antibodies were used: anti-MMP-2 (1:500; Bioworld Technology, Atlanta, GA, USA), anti-MMP-9 (1:500; Bioworld Technology), anti-E-cadherin (1:300; Boster, Wuhan, China), anti-Vimentin (1:200; Boster), anti-phosphorylated AKT (1:500; Cell Signaling Technology, Beverly, MA, USA), anti-AKT (1:1000; Cell Signaling Technology) and anti-β-actin (1:300; Zhongshan Golden Bridge, Beijing, China). Protein bands were quantified by densitometry using Quantity One 4.6.2 (Bio-Rad Laboratories, Hercules, CA, USA) and normalized to β-actin or total AKT.
Cell Counting Kit-8 assay
Cell growth was analysed by Cell Counting Kit-8 (CCK-8; Dojindo, Japan). Briefly, the cells were seeded into 96-well plates at a density of 1000 cells per well. A volume of 10 µL of CCK-8 reagent was added to each well, and the cells were subsequently incubated for 45 min at 37°C on days 1, 2, 3, 4, 5, 6 and 7 after plating. The absorbance was measured at 450 nm on a SpectraMax 190 Microplate Reader (Molecular Devices, Sunnyvale, CA, USA). In addition, recombinant human MFG-E8 (rhMFG-E8; R&D Systems Inc, Minneapolis, MN, USA) dissolved in phosphate-buffered saline (PBS) at different concentrations (0, 5, 10, 50,100, 200 and 500 ng/mL) was added to parental HT-29 and Caco-2 cells for 3 days of treatment, and then, CCK-8 assay was carried out. The medium was changed daily, and the final absorbance was presented as the mean value for samples analysed in triplicate.
Plate colony formation assay
HT-29 and Caco-2 cells transfected with LV-C or LV-M at log phase were plated in six-well plates at a density of 1500 cells/well. After culturing for 18 days, the cells were fixed with methyl alcohol for 10 min and stained with 0.2% crystal violet. Their colonies were then photographed and counted. All assays were independently conducted three times in triplicate.
Cell migration and invasion assay
Migration and invasion assays were performed using a Transwell system (8-µm pore size; Corning Inc., Corning, NY, USA). Briefly, 5 × 104 cells were seeded onto the apical side of the Transwell chamber (24-well insert) in serum-deprived culture (DMEM supplemented with 0.1% FCS). DMEM supplemented with 20% FCS was added to the basal compartment to serve as a chemoattractant. The cells were allowed to migrate for 24 h, and then, the cells that remained on the apical side of the chamber were gently scraped off using wetted cotton swabs. The cells that had migrated to the basal side were fixed in methanol for 10 min, stained with 0.2% crystal violet and then photographed. For invasion assays, the chamber was pre-coated with Matrigel (BD Biosciences, San Jose, CA, USA) and 2 × 105 cells were plated on the apical side of the chamber. The other procedures were carried out in the same manner as the migration assay. The migration and invasion assays were performed in triplicate and were repeated three times independently. RhMFG-E8, LY294002 (Sigma-Aldrich, St. Louis, MO, USA), PBS or dimethyl sulfoxide (DMSO; Beyotime Institute of Biotechnology) were added to both the upper and lower compartments of the Transwell chamber as needed.
Sequences of the primers used for real-time quantitative PCR analysis.
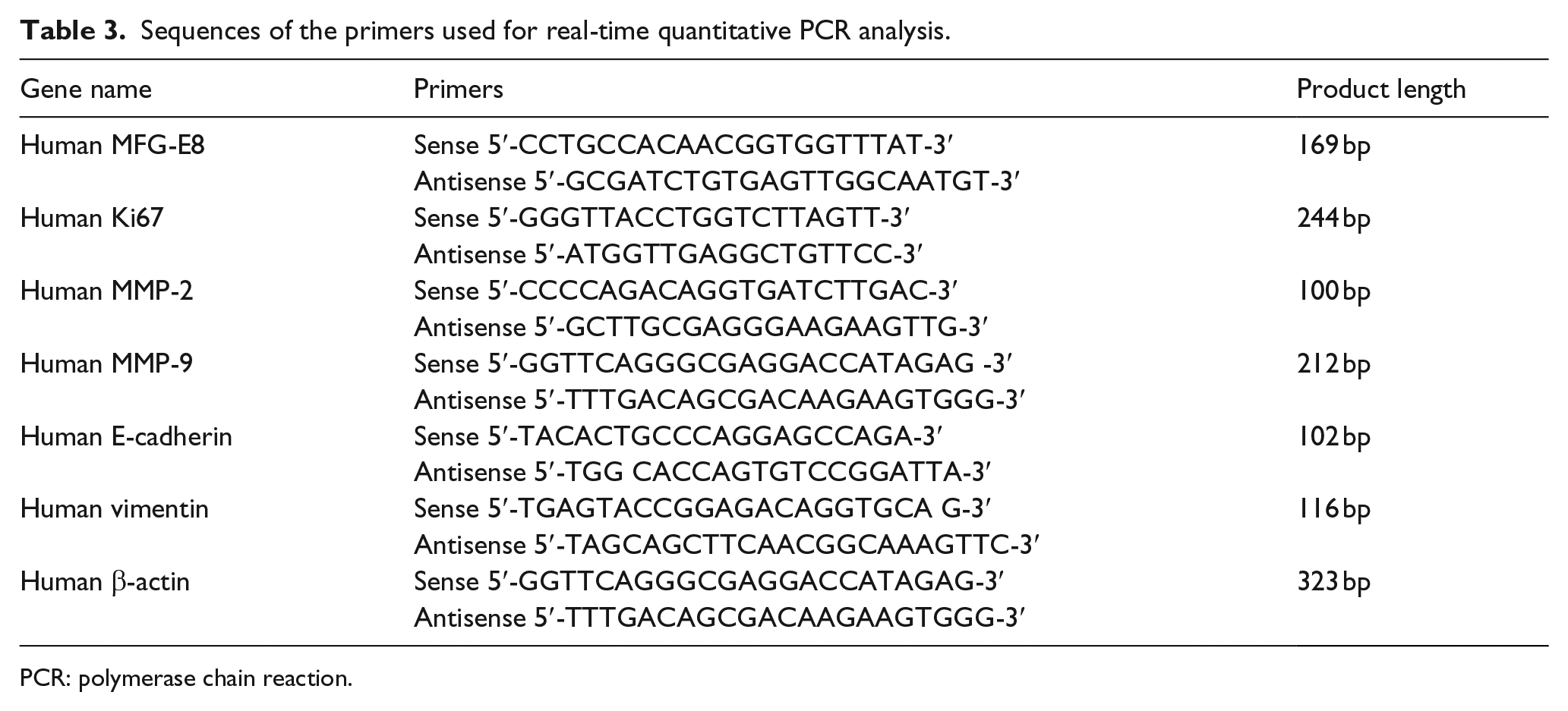

PCR: polymerase chain reaction.
Statistical analysis
All values were represented as the mean ± SEM from at least three independent experiments for cellular data. Chi-square tests or Yate’s continuity-corrected chi-square test was used for correlation studies. Paired and unpaired t-tests were used to analyse mRNA expression levels in the tissues of our CRC cohort and TCGA CRC cohort, respectively. For our in vitro studies, Student’s t-test was performed to evaluate the statistical significance of the differences between the two groups, and one-way analysis of variance (ANOVA) was performed to evaluate the significance of the differences among three or more groups. All analyses were performed using GraphPad Prism 5.01 (GraphPad Software, San Diego, CA, USA). Differences were considered statistically significant at p < 0.05 or less.
Results
MFG-E8 expression in CRC patients
As shown in Figure 1(a), MFG-E8 expression was detectable in normal colonic epithelium. In CRC tissue samples, MFG-E8-positive staining was distributed in cancer cell cytoplasms and cytomembranes. Among 78 CRC cancerous specimens, 58 specimens (74.4%) displayed staining for MFG-E8 that was significantly more intense than that displayed by adjacent non-cancerous specimens (21/58, 36.2%, p < 0.01; Figure 1(b)). Moreover, further analysis demonstrated that MFG-E8 expression was significantly higher in stage III–IV CRC specimens than stage I–II CRC specimens and non-cancerous specimens, while no significant MFG-E8 overexpression was observed in stage I–II CRC specimens compared with non-cancerous specimens (Figure 1(b)). Quantitative real-time polymerase chain reaction (qRT-PCR) analysis of 26 paired CRC and adjacent non-cancerous tissues revealed that MFG-E8 mRNA expression levels were also significantly upregulated in CRC tissues compared with adjacent non-cancerous colon tissues (p < 0.01, Figure 1(c)).

MFG-E8 expression in CRC specimens and paratumoural normal specimens and its correlation with prognosis of CRC patients. (a) Immunohistochemistry for MFG-E8 in adjacent non-cancerous tissues (A and B) and CRC tissues (C, D, E and F; 200×). Positive staining is mainly localized in the cytoplasm and cytomembrane of the epithelium and tumour cells. (b) High and low proportions of MFG-E8 expression in adjacent non-cancerous specimens (N), CRC specimens (Ca), stage I–II CRC specimens (I + II) and stage III–IV CRC specimens (III + IV). (c) qRT-PCR analysis of MFG-E8 mRNA levels in 26 paired adjacent non-cancerous specimens (N) and CRC specimens (Ca). MFG-E8 mRNA expression levels were normalized to those of β-actin. (d) Kaplan–Meier survival analysis and the log-rank test showed that patients with higher MFG-E8 expression levels had poorer cumulative survival than patients with lower MGF-E8 expression levels (p = 0.0227).
Correlation between MFG-E8 expression and CRC patient clinicopathological parameters
We also analysed the correlation between MFG-E8 staining scores and patient clinicopathological parameters in cancerous tissue samples. We found that MFG-E8 expression levels in CRC tissues were significantly correlated with T stage (p = 0.0322), lymph node metastasis (p = 0.0085), distant metastasis (p = 0.0025) and patient TNM stage (p = 0.0003) but were not related to age, gender, tumour size and differentiation (Table 1). Using the Kaplan–Meier method and log-rank test, we analysed the association between MFG-E8 levels and cancer-specific survival in CRC patients. We found that the high MFG-E8 expression group displayed significantly poorer survival than the low MFG-E8 expression group (Figure 1(d), p = 0.0227). This finding indicated that MFG-E8 seems to be a predictor of a poor prognosis and is related to metastasis in CRC patients.
MFG-E8 mRNA expression in TCGA CRC cohort and GSEA
To further confirm our results with MFG-E8 expression in CRC specimens, we downloaded level 3 CRC RNAseq data from TCGA project. No significant difference of MFG-E8 mRNA expression between 647 CRC specimens and 51 paratumoural normal specimens from TCGA project was observed (Figure 2(a)). However, in accordance with our immunohistochemistry results, MFG-E8 mRNA levels of stage III–IV CRC specimens were significantly higher than stage I–II CRC specimens and paratumoural normal specimens in TCGA CRC cohort (Figure 2(b)). No significant difference of MFG-E8 mRNA expression between stage I–II CRC specimens and paratumoural normal tissues was observed (Figure 2(b)). Survival analysis also suggested that the high MFG-E8 expression group displayed significantly poorer survival than the low MFG-E8 expression group in TCGA CRC cohort (Figure 2(c), p = 0.0148). Since human MFG-E8 binds to αVβ3/5 integrins, we further analysed the expression of integrin αV (ITGAV), β3 (ITGB3), as well as β5 (ITGB5) in CRC specimens from TCGA cohort. We found that ITGAV levels are similar, while ITGB3 levels are downregulated in CRC specimens compared with normal specimens (data not shown). Neither ITGAV nor ITGB3 expression is correlated with TNM stage of CRC specimens (data not shown). However, we found moderate but significant upregulation of ITGB5 levels in CRC specimens compared with normal specimens (Figure 2(d)). Further analysis revealed that ITGB5 mRNA levels of stage III–IV CRC specimens were higher than stage I–II CRC specimens in TCGA CRC cohort (Figure 2(e)). Gene ontology analysis and pathway analysis through GSEA revealed that high MFG-E8 expression group was enrichment of gene sets including EXTRACELLULAR_REGION_PART, G_PROTEIN_COUPLED_RECEPTOR_PROTEIN_SIGNALING_PATHWAY, ANGIOGENESIS, EXTRACELL ULAR_ MATRIX, ECM_GLYCOPROTEINS, ECM_REGULATORS and EXTRACELLULAR_MATRIX_ORGANIZATION, ECM_RECEPTOR_INTERACTION (Table 2).

MFG-E8 and ITGB5 mRNA expression in TCGA CRC cohort. (a) MFG-E8 mRNA levels are similar in 647 CRC specimens and 51 paratumoural normal specimens from TCGA cohort (p = 0.3021). (b) MFG-E8 mRNA levels in stage III–IV CRC specimens are significantly higher than stage I–II CRC and paratumoural normal specimens (p = 0.0004, stage III–IV CRC vs stage I–II CRC; p = 0.0051, stage III–IV vs normal). No significant difference of MFG-E8 mRNA levels between stage I–II CRC and paratumoural normal specimens was observed (p = 0.7970). (c) Correlation between MFG-E8 expression levels and patient cumulative survival in TCGA CRC cohort. Kaplan–Meier survival analysis and the log-rank test showed that patients with higher MFG-E8 expression levels had poorer cumulative survival than patients with lower MGF-E8 expression levels in TCGA CRC cohort (p = 0.0148). (d) ITGB5 mRNA levels are moderately but significantly upregulated in CRC specimens compared with paratumoural normal specimens from TCGA cohort. (e) ITGB5 mRNA levels in stage III–IV CRC specimens are significantly higher than stage I–II CRC and paratumoural normal specimens (p = 0.0003, stage III–IV CRC vs stage I–II CRC; p < 0.0001, stage III–IV vs normal).
Validation of MFG-E8 knockdown in CRC cells
MFG-E8 knockdown in HT-29 and Caco-2 CRC cells was validated by qRT-PCR and western blotting as described in detail in our previous study. 22
Effects of MFG-E8 on CRC cell growth
Using CCK-8, we monitored HT-29 and Caco-2 cell viability everyday for 7 consecutive days. We found that cell growth was significantly suppressed in the MFG-E8-knockdown groups compared with the LV-C-transfected groups (Figure 3(a)). Moreover, plate colony formation assay showed that CRC cells transfected with LV-M experienced a dramatic reduction in their colony numbers compared with LV-C transfected cells (Figure 3(b)). To confirm the role of MFG-E8 in CRC cell proliferation, we also detected the expression of the proliferation-related gene Ki67 by qRT-PCR. We did not observe any significant difference in Ki67 mRNA levels between LV-C- and LV-M-transfected CRC cells (Figure 3(c)). Since we previously showed that MFG-E8 knockdown can induce apoptosis in CRC cells, 22 we surmised that CRC cell growth suppression after MFG-E8 knockdown was caused by increased apoptosis rather than decreased proliferation. Pretreatment with different concentrations of rhMFG-E8 for 3 days did not seem to promote HT-29 (Figure 3(d)) or Caco-2 cell growth (similar to HT-29 cells, data not shown). Moreover, it seemed that rhMFG-E8 decreased CRC cell populations at high concentrations (200 and 500 ng/mL), although this effect was not significant (Figure 3(d)). We found this discrepancy involving MFG-E8-knockdown CRC cells confusing. We surmised that the basal levels of parental HT-29 and Caco-2 CRC cell apoptosis are very low; thus, the anti-apoptotic effects of rhMFG-E8 were not evident, and no differences in the sizes of the cell populations were observed after rhMFG-E8 treatment. Our results indicated that MFG-E8 knockdown suppressed CRC cell growth in vitro and that this effect was not induced by decreased cell proliferation, as Ki67 mRNA levels did not change after MFG-E8 knockdown in HT-29 and Caco-2 cells.

Effects of MFG-E8 on CRC cell growth. (a) CCK-8 assay of cell growth in HT 29 and Caco-2 cells transfected with LV-C and LV-M for 7 days. Cells transfected with LV-M exhibited significantly decreased cell growth. (b) Plate colony formation assay of HT-29 and Caco-2 cells transfected with LV-C and LV-M. LV-M-transfected cells exhibited dramatically reduced colony numbers. (c) qRT-PCR analysis revealed that Ki67 mRNA levels were similar in LV-C- and LV-M-transfected CRC cells (**p < 0.01, ***p < 0.0001). (d) CCK-8 cell growth assay of HT-29 cells treated with different concentrations of rhMFG-E8 for 3 days.
MFG-E8 promoted CRC cell migration and invasion accompanied by MMP-2 and MMP-9 upregulation
To explore the role of MFG-E8 in CRC cell migration and invasion, we performed Transwell and Matrigel assays of HT-29 and Caco-2 cells transfected with LV-M and LV-C. We found that LV-M-transfected cells exhibited impaired migration and invasion compared with LV-C-transfected cells (Figure 4(a)). To verify these findings, we added different concentrations of rhMFG-E8 (10 and 100 ng/mL) or vehicle (PBS) to both the upper and lower compartments of a Transwell system at the time of cell plating. We found that rhMFG-E8 can dramatically promote HT-29 and Caco-2 cell migration and invasion in a concentration-dependent manner (Figure 4(b)). Furthermore, qRT-PCR analysis revealed that MMP-2 and MMP-9 mRNA levels were downregulated in LV-M transfected HT-29 cells compared with LV-C transfected cells (Figure 5(a)). Western blotting showed similar results regarding MMP-2 and MMP-9 protein levels (Figure 5(c)). Moreover, pretreatment of HT-29 cells with rhMFG-E8 for 24 h upregulated MMP-2 and MMP-9 expression at both the mRNA and the protein levels (Figure 5(b) and (d)). This effect of rhMFG-E8 was dramatic at a concentration of 100 ng/mL.

Migration and Matrigel invasion assay of HT-29 and Caco-2 cells using a Transwell system. (a) LV-M-transfected HT-29 and Caco-2 cells exhibited impaired migration and invasion compared with LV-C-transfected cells. (b) The cells were treated with rhMFG-E8 at the time of plating. RhMFG-E8 significantly improved migration and invasion in both HT-29 and Caco-2 cells in a concentration-dependent manner (**p < 0.01).

Effect of MFG-E8 on the expression of MMP-2, MMP-9 and EMT markers in CRC cells. (a) qRT-PCR analysis of MMP-2, MMP-9, E-cadherin and vimentin expression in HT-29 cells transfected with LV-C or LV-M. MMP-2, MMP-9 and vimentin expression was downregulated while E-cadherin expression was upregulated at the mRNA level after MFG-E8 knockdown. (b) HT-29 cells were pretreated with rhMFG-E8 at different concentrations for 24 h. qRT-PCR revealed that rhMFG-E8 can upregulate MMP-2, MMP-9 and vimentin mRNA expression but suppress E-cadherin mRNA expression in HT-29 cells. The effect of rhMFG-E8 was dramatic at a concentration of 100 ng/mL. (c) Western blotting for MMP-2, MMP-9, E-cadherin and vimentin protein expression in HT-29 cells transfected with LV-C or LV-M, similar results with qRT-PCR. (d) Western blotting for MMP-2, MMP-9, E-cadherin and vimentin protein expression in HT-29 cells pretreated with rhMFG-E8 at different concentrations for 24 h. Band densities were quantified by Quantity One software and normalized to β-actin (*p < 0.05, **p < 0.01 and ***p < 0.0001). (e) Correlation analysis of MFG-E8 mRNA with MMP-2, MMP-9, vimentin and E-cadherin mRNA levels using CRC RNAseq data from TCGA. MFG-E8 is positively correlated with MMP-2, MMP-9 and vimentin, but negatively correlated with E-cadherin at mRNA level.
MFG-E8 promotes EMT in CRC cells
To investigate the role of MFG-E8 in EMT in CRC cells, we detected the expression levels of the epithelial cell maker E-cadherin and the mesenchymal cell maker vimentin in HT-29 cells transfected with LV-M and LV-C. Our real-time quantitative PCR analysis results were suggestive of an increase in E-cadherin mRNA levels and a decrease in vimentin mRNA levels in LV-M-transfected HT-29 cells compared with LV-C-transfected cells (Figure 5(a)). Western blotting showed the same results at the protein level (Figure 5(c)). Furthermore, pretreatment of HT-29 cells with rhMFG-E8 for 24 h suppressed E-cadherin expression but enhanced vimentin expression at both the mRNA and the protein level (Figure 5(b) and (d)). This effect was dramatic when a relatively high concentration of rhMFG-E8 (100 ng/mL) was used. Besides, correlation analysis with TCGA CRC RNAseq data proved that MFG-E8 mRNA levels are positively correlated with MMP-2, MMP-9 and vimentin mRNA levels but negatively correlated with E-cadherin mRNA levels (Figure 5(e)), which is in accordance with our in vitro studies.
AKT phosphorylation was responsible for MFG-E8-induced migration and invasion in CRC cells
PI3K/AKT pathway overactivation is frequently present in human malignancies and is involved in tumour invasion and progression. 23 We detected total AKT and phosphorylated AKT (p-AKT) by western blotting and found that HT-29 cells transfected with LV-M exhibited decreased p-AKT levels compared with LV-C-transfected cells (Figure 6(a)). Pretreatment of HT-29 cells with rhMFG-E8 for 1 h could upregulate p-AKT levels in a concentration-dependent manner in HT-29 cells (Figure 6(a)). To confirm the role of AKT activation in MFG-E8-induced cell migration and invasion, we employed the PI3K/AKT inhibitor LY294002. When HT-29 cells were pretreated with the PI3K inhibitor LY294002 (20 µM) for 2 h, rhMFG-E8-induced migration and invasion were suppressed compared with DMSO-pretreated cells (Figure 6(b)). Pretreatment with LY294002 also decreased rhMFG-E8-induced MMP-2 and MMP-9 upregulation in HT-29 cells (Figure 6(c)). These data indicate that PI3K/AKT pathway activation was involved in MFG-E8-induced CRC cell migration and invasion. Moreover, rhMFG-E8-induced EMT in HT-29 cells was also suppressed after pretreatment with the PI3K/AKT inhibitor LY294002 (Figure 6(c)).

Involvement of the PI3K/AKT pathway in MFG-E8-induced invasion, migration and downstream molecule expression in HT-29 cells. (a) Western blotting analysis revealed phosphorylated AKT was significantly downregulated in LV-M-transfected HT-29 cells compared with LV-C-transfected ones, while pretreatment with rhMFG-E8 can upregulate phosphorylated AKT (*p < 0.05, **p < 0.01 and ***p < 0.0001). (b) HT-29 cells were treated with LY294002 (20 nM) or DMSO and stimulated with rhMFG-E8 (100 ng/mL) or PBS at the time of plating using a Transwell system. After 24 h, the migrated and invaded cells on the basal side of the membrane were dyed and counted. (c) HT-29 cells were pretreated with LY294002 (20 µM) or DMSO for 2 h and then rhMFG-E8 (100 ng/mL) or PBS was added to the cells for 24 h of incubation. Western blotting was then performed to detect E-cadherin, vimentin, MMP-2, MMP-9, AKT and p-AKT protein levels. Band densities were quantified by Quantity One software and normalized to β-actin for E-cadherin, vimentin, MMP-2 and MMP-9 expression and to total AKT for p-AKT expression (L + M: LY294002 + rhMFG-E8; D + M: DMSO + rhMFG-E8; **p < 0.01,***p < 0.0001 vs PBS; ##p < 0.01, ###p < 0.0001 vs L + M).
Discussion
MFG-E8 has been reported to participate in the progression of various malignancies, including breast cancer, melanoma, bladder tumours and ovarian cancer.12,16,23,24 The plasma levels of MFG-E8 in CRC patients who underwent complete resection were significantly lower than those in CRC patients with unresectable tumours. 17 MFG-E8 overexpression was detected in CRC cancer tissues and was correlated with clinical disease stages, and MFG-E8-knockout mice displayed a smaller tumour burden than wild-type mice after azoxymethane injection. 18 In spite of these findings, the relationship between MFG-E8 expression and metastasis in CRC patients and the role of MFG-E8 in CRC cell invasion are still unclear.
In this study, using immunohistochemistry and qRT-PCR, we showed that MFG-E8 is overexpressed at both the mRNA and the protein levels in CRC tissues, findings consistent with those of a previous study by Kusunoki et al. 18 Moreover, further analysis revealed that MFG-E8 was dramatically upregulated in advanced CRC but not in early-stage CRC compared with paratumoural normal specimens. This suggests that maybe MFG-E8 is not involved in the process of early-stage CRC but probably participate in the process of advanced CRC. By analysing patient clinicopathological characteristics in detail, we showed that MFG-E8 levels in CRC tissues were positively correlated with T stage, lymph node metastasis, distant metastasis, and TNM stage in CRC patients. Kaplan–Meier curve analysis and the log-rank test showed that MFG-E8 was an independent factor for a poor prognosis in CRC patients. Furthermore, we got similar results by analysis of TCGA CRC RNAseq data which suggest an upregulation of MFG-E8 in stage III–IV CRC but not in stage I–II CRC. Besides, we found that the expression of ITGB5 is also upregulated in CRC specimens and correlated with TNM stage in TCGA CRC cohort. These results highly imply that MFG-E8 may be involved in CRC invasion and metastasis, findings consistent with those of previous studies regarding the role of MFG-E8 in melanoma and bladder tumours.12,13
The role of MFG-E8 in tumour progression is very complicated, and its biological functions have not been fully elucidated. In melanoma, elevated MFG-E8 expression promotes tumour invasion, EMT and angiogenesis through an AKT-dependent pathway. 12 In triple-negative breast cancer, MFG-E8 is upregulated by p63/p73 and serves as an oncogene, while in ER+ and erbB2+ breast cancers, MFG-E8 is downregulated and serves as a tumour suppressor. 16 In bladder tumours, MFG-E8 plays a protumoural role related to adaptive immune tolerance, but this role is not related to tumour angiogenesis. 13 In the study by Kusunoki et al., 18 the authors focused on the involvement of MFG-E8 in tumour cell proliferation and showed that recombinant mouse MFG-E8 exerted pro-proliferative effects in colon 26 murine colon cancer cells. In our study, we found that MFG-E8 knockdown led to decreased cell growth in HT-29 and Caco-2 cells. However, in contrast to the results of the previous study, our real-time PCR results revealed that MFG-E8 expression levels were not related to the mRNA expression levels of Ki67, a proliferation-related gene, in human CRC cells. Pretreating CRC cells with different concentrations of rhMFG-E8 did not alter cell growth. The results of our previous study indicated that MFG-E8 knockdown can induce apoptosis in HT-29 and Caco-2 cells by downregulating Bcl-2 and upregulating BAX. 22 Since cell growth is a concomitant result of cell proliferation and apoptosis, according to the above results, we concluded that MFG-E8 knockdown-induced cell growth suppression in CRC cells was caused by increased apoptosis rather than decreased proliferation. Our results are similar to those of a previous study involving a mouse pancreatic cancer model, which showed that the rate of cell proliferation in the tumour tissues of the MFG-E8-knockout group, as detected by 5-bromo-2′-deoxyuridine (BrdUrd) in situ, was indistinguishable from that of the control group, while the rate of cell apoptosis was increased compared with that of the control group. Thus, MFG-E8-knockout animals ultimately displayed dramatic reductions in their tumour burden compared with control animals. 25 However, proliferation in angiogenic islets from MFG-E8 knockout animals was decreased compared with that in control angiogenic islets. 25 The results of these reports indicate that the effect of MFG-E8 on cell proliferation varies depending on the cell type. Our study was carried out with the human colon cancer cell lines HT-29 and Caco-2, which have been shown to express human MFG-E8 at baseline, whereas Kusunoki employed a murine colon cancer cell line, colon 26, in their study. Moreover, human MFG-E8 shares only 52% sequence identity with mouse MFG-E8. 7 Therefore, we postulated that the difference between the results of our study and those of the study by Kusunoki may be due to differences in the origins of the species and the cell types used in each study.
Invasion and migration have been acknowledged as the most lethal attributes of solid tumours and account for the majority of metastases. 26 Although MFG-E8 has been associated with tumour invasion and migration in many malignancies, its role in CRC cell invasion is unclear. Here, using the Transwell Matrigel assay, we showed that MFG-E8 knockdown dramatically suppressed CRC cell migration and invasion, while the addition of rhMFG-E8 exerted the opposite effect. MMPs are a family of endopeptidases with multifaceted roles and are best known for their ability to degrade the components of the extracellular matrix. They play critical roles in tumour cell invasion and are involved in many steps of tumour metastasis.5,27 MMP-2 and MMP-9 have been proved to be upregulated and related to invasion and metastasis in CRC.28,29 In this study, we noted decreased MMP-2 and MMP-9 mRNA and protein expression after MFG-E8 knockdown in HT-29 cells. We also found that pretreatment of HT-29 cells with rhMFG-E8 can upregulate MMP-2 and MMP-9 expression. EMT is a pivotal event in tumour metastasis through which epithelial cells lose their polarity and cell–cell contact and undergo remarkable remodelling of their cytoskeleton, facilitating cell motility and invasion. 6 In this study, we found that MFG-E8 knockdown caused a reversal of the EMT process in HT-29 cells, as shown by decreases in the expression of the mesenchymal maker vimentin and enhancements of the expression of the epithelial maker E-cadherin. Meanwhile, pretreatment of HT-29 cells with rhMFG-E8 induced EMT, a finding supported by our observation of a shift from epithelial markers to mesenchymal makers. Our results are similar to those of a previously published study regarding melanoma, which showed that MFG-E8 can enhance human melanoma cell invasion and metastatic seeding, a phenomenon accompanied by EMT induction and MMP-2 and MMP-9 expression in macrophages. 12 Recently, Juan Le et al. 30 reported that exogenous human intestinal trefoil factor (ITF/TFF3) can promote transcriptional expression of MMP-2 and MMP-9 but can induce E-cadherin degradation in HT-29 cells. Interestingly, the findings of our previous study indicated that MFG-E8 knockdown can lead to decreased ITF/TFF3 expression in HT-29 cells. 22 Thus, to some extent, the findings of this study, which showed that MFG-E8 knockdown led to MMP-2 and MMP-9 suppression, as well as E-cadherin induction, in HT-29 cells, are consistent with those of the study by Juan Le et al. 30
PI3K/Akt pathway overactivation is frequently present in CRC and is associated with tumour progression processes, including cell proliferation, migration and invasion. 31 It has been shown to contribute to tumour metastasis by promoting the secretion of MMPs and the induction of EMT. 23 PI3K/AKT pathway activation can occur in response to a variety of extracellular signals, including αVβ3integrin receptor pathway activation. 32 As a ligand of αVβ3/5integrins, MFG-E8 has been recognized as a critical activator of the PI3K/AKT pathway in several cellular systems, such as melanoma and endothelial cell systems.12,33 Therefore, we focused on this pathway and found that MFG-E8 knockdown suppressed AKT phosphorylation, while rhMFG-E8 treatment greatly induced AKT phosphorylation in HT-29 cells. Moreover, pretreatment with the PI3K/AKT inhibitor LY294002 dramatically suppressed MFG-E8-induced HT-29 cell migration and invasion. Many studies have shown that MMPs and EMT can be induced by PI3K/AKT pathway activation.34–36 In this study, we also showed that MFG-E8 induced increases in MMP-2 and MMP-9 expression and that a shift from epithelial markers to mesenchymal markers was suppressed by LY294002. These results imply that PI3K/AKT signalling is involved in MFG-E8-induced cell migration, invasion and EMT. However, several other signalling molecules, including mitogen-activated protein kinase (MAPK), extracellular signal–regulated protein kinases 1 and 2 (ERK1/2) and signal transducer and activator of transcription 3 (STAT3), have also been shown to be associated with MFG-E8 downstream signalling.37–39 Their roles in MFG-E8-related invasion and EMT in CRC cells need to be explored.
In conclusion, the findings of this study indicated that MFG-E8 was overexpressed in advanced CRC tissues but not in early-stage CRC tissues. Moreover, MFG-E8 expression was associated with plasma membrane invasion, lymph node metastasis, distant metastasis and patient TNM stage. High MFG-E8 expression levels are predictive of a poor prognosis in CRC patients. GSEA revealed that high MFG-E8 expression is enrichment of gene sets mostly correlated with extracellular matrix. The results of our in vitro studies showed that MFG-E8 knockdown can slow CRC cell growth and that cell growth impairment was not caused by decreased cell proliferation. MFG-E8 knockdown also suppressed CRC cell migration, invasion and EMT, changes accompanied by MMP-2 and MMP-9 downregulation in CRC cells, while rhMFG-E8 treatment exerted the opposite effects. Finally, MFG-E8-induced migration, invasion, EMT and MMP induction were partially mediated by the PT3K/AKT pathway. The findings of our study suggest that MFG-E8 is a risk factor for CRC metastasis that its expression is indicative of a poor prognosis. Moreover, our findings have provided new insight into the mechanisms underlying the role of MFG-E8 in CRC progression.
Footnotes
Compliance with ethical standards
This study was approved by the Clinical Ethics Committee of Qilu Hospital of Shandong University, and all the patients provided written informed consent before sampling.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
