Abstract
Glioma accounts for the majority of primary malignant brain tumors in adults and is highly aggressive. Although various therapeutic approaches have been applied, outcomes of glioma treatment remain poor. MicroRNAs are a class of small noncoding RNAs that function as regulators of gene expression. Accumulating evidence shows that microRNAs are associated with tumorigenesis and tumor progression. In this study, we found that miR-105 is significantly downregulated in glioma tissues and glioma cell lines. We identified suppressor of Zeste 12 homolog as a novel direct target of miR-105 and showed that suppressor of Zeste 12 homolog protein levels were inversely correlated with the levels of miR-105 expression in clinical specimens. Overexpression of miR-105 inhibited cell proliferation, tumorigenesis, migration, invasion, and drug sensitivity, whereas overexpression of suppressor of Zeste 12 homolog antagonized the tumor-suppressive functions of miR-105. Taken together, our results indicate that miR-105 plays a significant role in tumor behavior and malignant progression, which may provide a novel therapeutic strategy for the treatment of glioma and other cancers.
Introduction
Glioma is the most prevalent type of primary brain tumor in adults. 1 Despite recent advances in standard therapy, including surgical resection followed by radiation and chemotherapy, the prognosis for patients with malignant gliomas remains dismal.2,3 Thus, there is an urgent need to identify new molecular targets and treatment strategies for this disease.
MicroRNAs (miRNAs) are short single-stranded RNA molecules, 20–25 nucleotides in length, which post-transcriptionally regulate gene expression by binding to complementary sequences in the 3′-untranslated region (UTR) of target mRNAs. 4 MiRNAs have crucial roles in regulating diverse cellular processes, such as proliferation, invasion, apoptosis, stress responses, differentiation, and development.5,6 Dysregulation or aberrant expression of miRNAs is associated with a variety of disorders, particularly cancers such as gliomas.7–13 Moreover, miRNAs can be useful diagnostic and prognostic indicators of disease type and severity.14–17 Nevertheless, our understanding of how glioma-associated miRNAs can affect the molecular mechanisms of glioma occurrence and progression remains limited.
Suppressor of Zeste 12 homolog (SUZ12) is an essential component of polycomb repressive complex 2 (PRC2), which modifies transcription by affecting histone and DNA methylation. 18 SUZ12 is rarely expressed in normal tissues, but is amplified and overexpressed in several human cancers, such as ovarian cancer, mantle cell lymphoma, and breast cancer.19–21 Accumulating evidence indicates that SUZ12 plays a significant role in carcinogenesis by acting as an oncogene, stimulating cell proliferation, blocking apoptosis, and promoting cell invasion and metastasis.19–23 However, the status and function of SUZ12 in glioma are still unknown.
The aim of this study was to identify miR-105 as a novel tumor-suppressive miRNA and to further investigate its role and function in human glioma. Moreover, we confirmed that SUZ12 is a direct target for miR-105 and showed that overexpression of SUZ12 in glioma tissues and glioma cell lines could result from miR-105 downregulation. Finally, our results revealed that the expression level of miR-105 correlated with the grade of glioma, and overexpression of miR-105 reduced cell proliferation, induced G1-phase arrest, attenuated tumor growth in vivo, decreased cell invasion and migration capacities, and improved drug sensitivity. SUZ12 overexpression reversed these effects. These findings provide novel insights into the role and mechanism of action of miR-105 in glioma and indicate a potential therapeutic strategy for future glioma treatment.
Materials and methods
Human tissue samples
In all, 32 human glioma tissue samples, including 8 grade II tumors, 8 grade III tumors, and 16 grade IV (glioblastoma multiforme) tumors, graded by a clinical pathologist in accordance with the World Health Organization criteria, were obtained from the Department of Neurosurgery of the First Affiliated Hospital of Nanjing Medical University, China. A total of 12 normal brain tissues were collected as negative controls (NCs) from patients undergoing decompressive craniectomy for traumatic brain injury. This study was approved by the institutional review board and the ethics committee of Nanjing Medical University, China, and written informed consent was obtained from all patients. All samples were collected and immediately frozen and stored in liquid nitrogen until analysis.
Cell culture and antibodies
Human glioma cell lines, U87, U251, T98, LN229, A172, and U118, were purchased from American Type Culture Collection. All cell lines were maintained at 37°C in a 5% CO2 atmosphere in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin. Normal human astrocytes (NHAs) were obtained from Lonza and were cultured in accordance with the manufacturer’s instructions. Antibodies against SUZ12, matrix metalloproteinase (MMP)-2, MMP-7, MMP-9, methylguanine DNA-methyltransferase (MGMT), and Cleaved Caspase-3 were purchased from Cell Signaling Technology. Antibodies against glyceraldehyde 3-phosphate dehydrogenase (GAPDH) were obtained from Beyotime Biotechnology. Antibodies against cyclin-dependent kinase 6 (CDK6), Cyclin D1, and Cyclin E1 were from Open Biosystems, and temozolomide (TMZ) was obtained from Sigma.
RNA extraction and quantitative real-time polymerase chain reaction
Total RNA was isolated from human glioma specimens, normal brain tissues, or cultured cells using TRIzol reagent (Invitrogen), in accordance with the manufacturer’s instructions. To detect the relative levels of miR-105 in tissues and cells, the ABI 7300 HT sequence detection system (Applied Biosystems) was used for TaqMan-based real-time reverse transcription polymerase chain reaction (RT-PCR) assays. The primers and probes for the miR-105 TaqMan miRNA assays were purchased from Applied Biosystems. Each reaction was performed in triplicate and analysis was performed using the 2−ΔΔCt method.
Lentiviral packaging and stable cell line establishment
A lentiviral packaging kit was purchased from GenePharma. Lentiviruses carrying hsa-miR-105 or hsa-miR-NC were packaged in human embryonic kidney 293T cells and collected from the supernatant, in accordance with the manufacturer’s instructions. Stable cell lines were established by infecting U87 and U251 cells with lentivirus, followed by puromycin selection.
Luciferase reporter assay
Wild-type and mutated putative miR-105 binding sites in the 3′-UTR region of SUZ12 were amplified and cloned into the pGL3-control luciferase reporter plasmid (Invitrogen). The pRL vector constitutively expressing Renilla luciferase was used to normalize transfection efficiency. Luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega).
Cell proliferation
A total of 2000 cells were seeded per well in 96-well plates and cultured 72 h after transfection. The absorption of cells was measured using a Cell Counting Kit-8 (CCK-8; Dojindo Laboratories) in accordance with the manufacturer’s instructions at the indicated time points.
Subsequently, U87 cells were labeled with 5-ethynyl-2′-deoxyuridine (EdU) for 12 h. The cell proliferation rate was detected using a Cell-Light™ EdU In Vitro Imaging Detection Kit (cat no. C10310-1; Guangzhou RiboBio Co., Ltd.) in accordance with the manufacturer’s instructions. Data were recorded from three separate experiments, each with four replicates.
Colony formation assay
For colony formation assays, 5 × 102 cells were plated onto 60-mm tissue culture plates. After 10–14 days, visible colonies were fixed with 100% methanol and stained with 0.1% crystal violet in 20% methanol for 15 min. Colony-forming efficiency was calculated as the number of colonies/number of plated cells × 100%.
Cell-cycle assay
Cells in the logarithmic phase of growth were harvested by trypsinization 48 h after transfection, washed with phosphate-buffered saline (PBS), and fixed with 75% ethanol at −20°C for at least 1 h. After extensive washing, the cells were resuspended in PBS containing 50 mg/mL propidium iodide (PI; Sigma-Aldrich), 100 mg/mL RNase A (Sigma-Aldrich), and 10 µL/mL 1% Triton X-100 and incubated for 1 h at room temperature in the dark, before being analyzed by flow cytometry. Data are presented as the proportion of cells in a particular phase. The experiments were performed in triplicate.
Apoptosis assay
The apoptotic rates were determined by flow cytometric (FCM) analysis. Briefly, U87 and U251 cells (2 × 105) were plated into six-well plates. Double staining with Annexin V-fluorescein isothiocyanate (FITC) and PI was used to detect and quantify cellular apoptosis 48 h after transfection. Cells without Annexin V and PI (Annexin V− and PI− cells) were used as controls. Annexin V+ and PI− cells were designated as apoptotic, while Annexin V+ and PI+ cells were necrotic. Tests were performed in triplicate.
Wound-healing assay
Wound-healing assays were used to assess cell migration. Cells were cultured to 90% confluency in six-well plates. Some of them were then scratched away using a 20-µL tip to form wound gaps, washed with PBS twice, and cultured. The wound gaps were photographed at 0, 12, and 24 h and the width of each open wound was measured.
Cell invasion assay
Invasion assays were performed using 24-well BD Matrigel invasion chambers (BD Biosciences), in accordance with the manufacturer’s instructions. A total of 2 × 104 cells were seeded in the upper well of each invasion chamber in DMEM without serum. The lower chamber well contained DMEM supplemented with 10% FBS to stimulate cell invasion. After incubation for 24 h, noninvading cells were removed from the top well with a cotton swab, while the cells that had migrated to the lower surface of the membrane were fixed with 4% paraformaldehyde and stained with 0.1% crystal violet. The total number of cells adhering to the lower surface of the membrane was recorded in six representative fields. Experiments were performed in triplicate.
Chemosensitivity assay
Cancer cells were seeded at a density of 3000 cells per well in a 96-well plate overnight. Freshly prepared TMZ solution was added at different concentrations as indicated, and cell survival was determined by CCK-8 assays 48 h later. The proportions of live cells were calculated after normalization to the controls (cells without prior drug treatment).
We also assessed drug sensitivity through cell proliferation by seeding 1 × 105 cells in medium containing 100 µM TMZ. The cells were treated with 100 µM TMZ for 24 h and then either treatment continued or the cells were cultured without TMZ (medium change (MC)) as indicated. The cell number was subsequently determined in the parallel cultures by counting trypsinized cells on each day for 7 days.
Western blot analysis
Human glioma cells were lysed in radioimmunoprecipitation assay buffer, and equal amounts of protein were separated by 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), followed by electrotransfer onto a polyvinylidene difluoride membrane (Thermo Fisher Scientific). Membranes were blocked for 1 h with 5% nonfat milk and then incubated at room temperature with diluted primary antibodies against SUZ12, MMP-2, MMP-7, MMP-9, MGMT, Cleaved Caspase-3 (all 1:1000; Cell Signaling Technology), CDK6, Cyclin D1, or Cyclin E1 (all 1:800; Open Biosystems), followed by incubation with a horseradish peroxidase–conjugated secondary antibody (1:3000; Santa Cruz Biotechnology). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as a control (Beyotime Biotechnology).
Subcutaneous xenograft model
Male BALB/c nude mice 4 weeks old were purchased from Beijing Weitonglihua Laboratory Animal Company and maintained under specific pathogen-free (SPF) conditions for 1 week. Animal handling and experimental procedures were performed in accordance with the Guide for the Care and Use of Laboratory Animals and were approved by the Animal Experimental Ethics Committee of Nanjing Medical University. A total of 12 nude mice were randomly divided into two groups. U87 cells (5 × 106 cells in 100 mL) infected with lentiviruses harboring miR-NC or miR-105 were subcutaneously injected into the posterior flank of the BALB/c nude mice. Tumor sizes were measured using a vernier caliper every 4 days after injection, and tumor volumes were calculated using the following formula: volume = 0.5 × length × width 2 . The mice were sacrificed and the tumors were weighed 36 days after injection. Total proteins were extracted for western blot assays to test for specific protein levels.
Immunohistochemical assay
Immunohistochemical assays to examine SUZ12 levels were conducted on paraffin-embedded tissue sections (nude mouse tumors). The tissue sections were incubated with 1:200-diluted primary antibodies overnight at 4°C, followed by incubation with biotin-labeled immunoglobulin G (1:100 dilution; GeneTech) for 1 h at room temperature. As a NC, the primary antibody was replaced with PBS. The sections were then incubated with avidin–biotin–peroxidase complex followed by diaminobenzidine, counterstained with hematoxylin (GeneTech), and visualized under a light microscope. The proportion of positive tumor cells and the staining intensity were assessed at 200× magnification.
Statistical analysis
All experiments were performed three times and data were analyzed using GraphPad Prism 5 software. Correlations between miR-105 and SUZ12 levels in glioma tissues were analyzed using Spearman’s rank test. Student’s t-test was used to statistically evaluate the data. Differences were considered statistically significant at p < 0.05.
Results
Expression of miR-105 is downregulated in glioma tissues and glioma cell lines and correlates with glioma grade
To assess the significance of miR-105 in glioma, total RNAs were extracted from 12 normal brain tissues and 32 glioma tissues (grade II 8, grade III 8, and grade IV 16), and the expression levels of miR-105 were analyzed using quantitative RT-PCR (qRT-PCR). As shown in Figure 1(a), the levels of miR-105 were consistently lower in glioma tissues than in normal brain tissues; furthermore, miR-105 levels negatively correlated with tumor grade. We also explored the expression of miR-105 in six glioma cell lines (U87, U251, T98, LN229, A172, and U118) and NHAs; miR-105 levels in all six glioma cell lines were clearly lower than in the NHAs (Figure 1(b)).
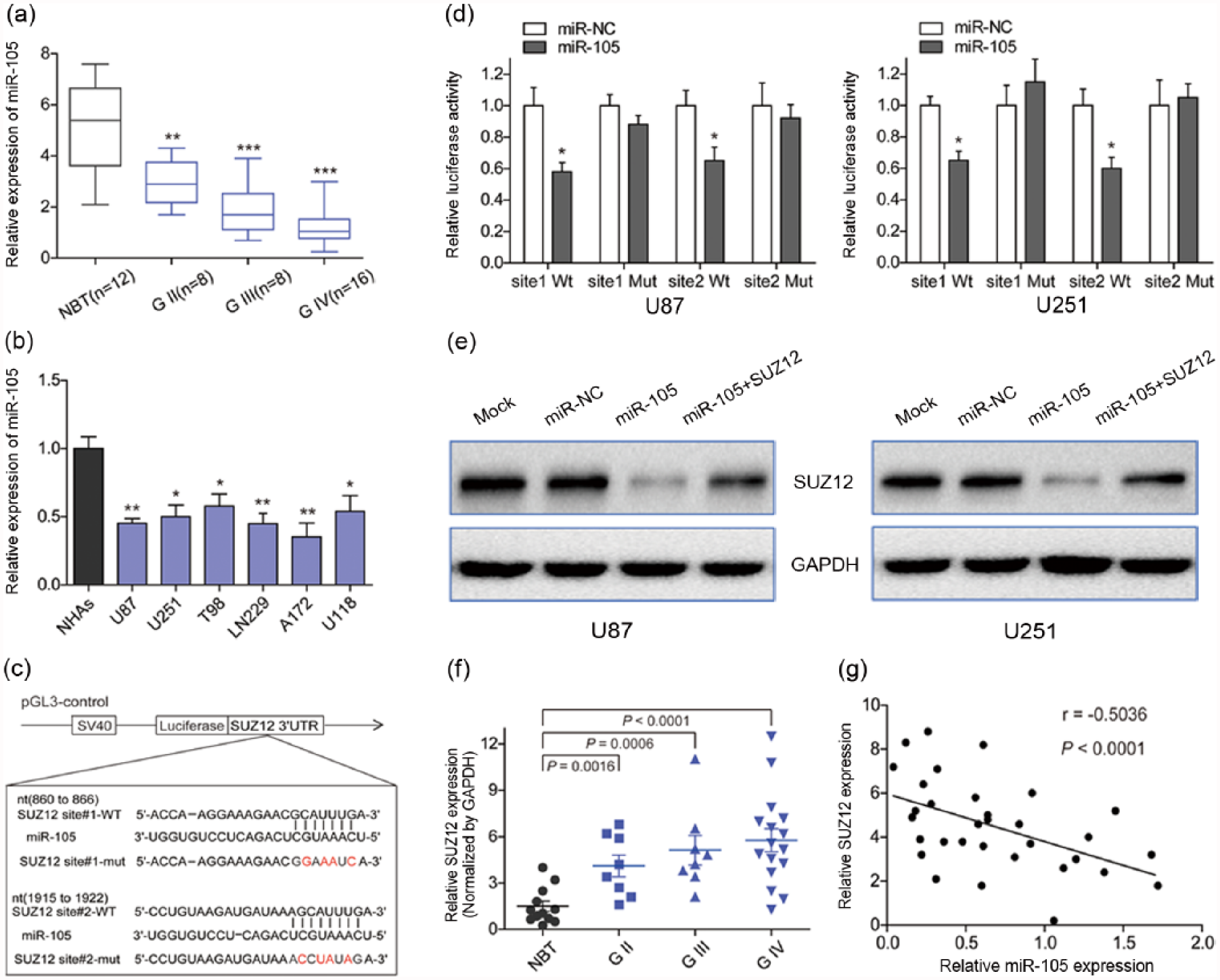
Expression of miR-105 in glioma tissues and glioma cell lines is decreased and this miRNA targets SUZ12. (a) Comparison of miR-105 levels among normal brain tissue (NBT) and different pathological grades of glioma using qRT-PCR analysis. Mean miR-105 level of normal brain tissue was higher than that of glioma tissue (**p < 0.01 and ***p < 0.001). (b) qRT-PCR analysis of miR-105 in normal human astrocytes (NHAs) and six glioma cell lines (U87, U251, T98, LN229, A172, and U118). Relative expression level of miR-105 in NHAs was higher than in the six glioma cell lines (*p < 0.05 and **p < 0.01). (c) The sequence of miR-105 binding sites within SUZ12. The reporter constructs of the SUZ12 3′-UTR sequences and the mutated nucleotides of the SUZ12 3′-UTR are labeled in red. (d) Luciferase reporter assay showed lower luciferase activity when cells were transfected with wild-type reporter, while there was no significant change of reporter activity when cells were transfected with mutated reporter (*p < 0.05 and **p < 0.01). (e) Western blot analysis showed that overexpression of miR-105 markedly suppressed SUZ12 protein expression in the two cell lines. GAPDH served as a loading control. (f) The expression levels of SUZ12 in normal brain tissues were lower than in specimens from glioma of different pathological grades, as determined by western blot analysis, and fold changes were normalized by GAPDH. (g) Spearman’s correlation analysis revealed an inverse correlation between the expression levels of SUZ12 and miR-105 in human glioma specimens (Spearman’s correlation analysis, r = −0.5036, p < 0.0001).
SUZ12 is a direct target of miR-105 and its expression inversely correlates with that of miR-105 in glioma
To predict target genes of miR-105, we applied three bioinformatic algorithms, TargetScan, miRanda, and miRBase, and identified SUZ12 as a potential target gene of miR-105 (Figure 1(c)). To confirm this, a luciferase reporter assay was carried out. As shown in Figure 1(d), luciferase activities were significantly reduced in cells transfected with wild-type SUZ12 3′-UTR and miR-105, but not in cells containing mutant SUZ12 3′-UTR and miR-105. Meanwhile, western blot analysis revealed that SUZ12 levels were downregulated in U87 and U251 cells by the overexpression of miR-105 compared with those in mock or miR-NC group cells, but the level of SUZ12 was effectively rescued by the overexpression of SUZ12 in miR-105-overexpressing cells (Figure 1(e)). Similarly, by performing immunofluorescence staining, we found that the expression levels of SUZ12 in miR-105-overexpressing cells were lower than in the miR-NC group and in the SUZ12-overexpressing group in the nucleus (Supplementary Figure A). These results, taken together, indicated that SUZ12 3′-UTR serves as a target of miR-105.
We also measured the levels of SUZ12 protein in glioma specimens and normal brain tissues. The level of SUZ12 was significantly higher in tumor tissues than in normal brain tissues, and the levels correlated with the World Health Organization glioma grade (Figure 1(f)). We then determined the correlation between SUZ12 and miR-105 levels. As shown in Figure 1(g), Spearman’s rank correlation analysis showed that SUZ12 in human glioma specimens were inversely correlated with miR-105 expression levels (r = −0.5036, p < 0.0001).
MiR-105 targets SUZ12 to inhibit cell proliferation and induce G1-phase arrest in glioma cells
Having demonstrated SUZ12 to be a target of miR-105, we further specifically investigated whether it is a functional target of miR-105. To do this, we first evaluated the effect of miR-105 on cell proliferation using CCK-8 assays, EdU assays, and colony formation assays. The CCK-8 assays showed that miR-105-transduced U87 and U251 cells exhibited significantly lower growth rates than mock or NC cells; however, cells transfected with miR-105 followed by SUZ12 did not show significant growth inhibition (Figure 2(a)). The same results were also observed in the colony formation assays (Figure 2(b)) and EdU assays (Supplementary Figure B).

SUZ12 is a functional target of miR-105 to inhibit cell proliferation and induce G1-phase arrest in glioma cells. (a) CCK-8 assay revealed that miR-105 suppressed cell growth, while SUZ12 overexpression promoted it (*p < 0.05 and **p < 0.01). (b) MiR-105 decreased the colony number of glioma cells, while SUZ12 overexpression increased it (*p < 0.05). (c and d) The cell-cycle distribution was analyzed by flow cytometry. Data reveal that miR-105 overexpression increased the proportion of cells in the G1 phase, while SUZ12 overexpression decreased the proportion. (e) The effects of miR-105 and its target SUZ12 on cell-cycle proteins (Cyclin D1, Cyclin E1, and CDK6) in the G1/S transition were determined by western blot analysis. GAPDH was used as a loading control.
To examine the mechanism by which cell growth was inhibited, we used FCM to analyze the cell-cycle distribution and apoptosis of cells. Compared with mock and miR-NC groups, U87 and U251 cells overexpressing miR-105 showed a marked increase in the number of cells in G1 phase, while the proportion of cells transfected with miR-105 followed by SUZ12 in G1 phase was reduced (Figure 2(c) and (d)). Furthermore, a greater level of apoptosis was exhibited by cells treated with miR-105 than with the mock treatment in the miR-NC control group, while the overexpression of SUZ12 decreased apoptosis (Supplementary Figure C). To explore the molecules involved in determining the cell-cycle distribution, we measured the expression of the G1-phase regulators, Cyclin D1, Cyclin E1, and CDK6,24–26 in transfected cells. As shown in Figure 2(e), the expression level of Cyclin D1 was suppressed in cells that were transfected with miR-105; however, the expression levels of Cyclin E1 and CDK6 were not significantly changed. Meanwhile, the overexpression of SUZ12 restored the Cyclin D1 expression. Collectively, these results suggest that miR-105 inhibits glioma cell proliferation and induces G0/G1 cell-cycle arrest by downregulating SUZ12.
MiR-105 attenuated tumor growth in nude mice and decreased SUZ12 protein levels in subcutaneous tumors
Several tumor-suppressive miRNAs that inhibit tumor growth in mouse models have been identified;27–29 therefore, we investigated whether miR-105 could also attenuate the progression of glioma in vivo. As shown in Figure 3(a)–(c), compared with the miR-NC group, miR-105-expressing cells generated tumors that were significantly smaller in size and lighter in weight. In addition, immunohistochemistry and immunofluorescence showed that the level of SUZ12 in the subcutaneous tumor tissues of the miR-105-expressing group was lower than that of the miR-NC group (Figure 3(d) and Supplementary Figure D). Consistent with our previous studies, western blotting also showed that the levels of SUZ12 and Cyclin D1 in the tumor tissues of the miR-105-overexpressing group were lower than those of the miR-NC group (Figure 3(e) and (f)). Together, these results indicate that miR-105 inhibits cell proliferation and tumor growth and might have therapeutic potential for the treatment of glioma.

MiR-105 attenuated tumor growth in nude mice and decreased SUZ12 protein level in subcutaneous tumors. (a–c) Male BALB/c nude mice were subcutaneously injected with U87 cells and tumor volumes were measured every 4 days, and the tumors were harvested and photographed after 36 days. Representative images from each group are shown. The tumors from the miR-105 group were smaller and lighter than those from the miR-NC group (*p < 0.05, **p < 0.01, and ***p < 0.001). (d) SUZ12 expression was measured by immunohistochemistry in mouse tumor tissues; compared with that in the miR-NC group, the level of SUZ12 in the miR-105 group was lower (magnification 200×). (e and f) The levels of SUZ12, Cyclin D1, Cyclin E1, and CDK6 from the tumor tissues in the two groups were analyzed by western blotting. GAPDH was used as an internal control.
SUZ12 is involved in miR-105-induced inhibition of glioma cell migration and invasion
Migration and invasion are cardinal features of malignancies, including glioma. Therefore, the wound-healing assay and Transwell assay were conducted to investigate the effects of miR-105 and its target gene, SUZ12, on cell invasion and migration capacity. We found that the overexpression of miR-105 decreased the migration capacity of glioma U87 and U251 cells, and the relative widths of the open cell wounds were greater than those of the mock and NC groups at 12 and 24 h. However, SUZ12 rescued the capacity for cell migration in U87/miR-105 and U251/miR-105 cells (Figure 4(a) and (b).

SUZ12 is involved in miR-105-induced inhibition of glioma cell migration and invasion. (a and b) Wound-healing assay showed that miR-105 significantly suppressed wound healing, which was rescued by the overexpression of SUZ12 in miR-105-overexpressing cells (*p < 0.05 and **p < 0.01). (c and d) Transwell assay indicated that the number of invasive cells in the miR-105 group was lower than that in the control, while this number in the SUZ12-overexpressing group did not show a significant change compared with the negative control group. Columns are the average of three independent experiments (*p < 0.05). (e) The effects of miR-105 and its target SUZ12 on metalloproteinases (MMP-2, MMP-7, and MMP-9) were determined by western blot analysis. MiR-105 suppressed the levels of MMP-2 and MMP-9, but these levels were restored by SUZ12 overexpression. GAPDH was used as a loading control.
The same effect was also observed in the cell invasion assay; specifically, miR-105 dramatically reduced the invasion of U87 and U251 cells in the Matrigel Transwell assays, while there was no significant change in invasion capacity after transfection with miR-105 followed by SUZ12, compared with that in the mock and NC groups (Figure 4(c) and (d)). Furthermore, miR-105 overexpression led to decreased levels of MMP-2 and MMP-9, which play important roles in tumor invasion and metastasis. 30 The reduced levels of MMP-2 and MMP-9 were rescued by the overexpression of SUZ12 in miR-105-overexpressing cells (Figure 4(e)).
MiR-105 increases chemosensitivity of glioma cells to TMZ treatment
To explore the potential role of miR-105 in chemotherapy, we chose TMZ as our experimental chemotherapy drug. Compared with mock or miR-NC group cells, the overexpression of miR-105 in U87 and U251 cells significantly increased the chemosensitivity to TMZ treatment (Figure 5(a)), and cell viability was substantially suppressed by TMZ treatment. Furthermore, we assessed drug sensitivity through cell proliferation in the presence of TMZ (100 µM) using the CCK-8 assay. The results showed that the capacity for proliferation was inhibited and that this inhibition was more pronounced in cells overexpressing miR-105 (Figure 5(b), red line). In contrast, when TMZ was removed after 24 h, miR-105-overexpressing cells showed only the partial restoration of proliferation, while other group cells were able to proliferate again (Figure 5(b), blue line). Moreover, we found that the expression of cleaved Caspase-3, 31 a key executor of apoptosis, was significantly upregulated upon treatment with miR-105 + TMZ compared with that in mock + TMZ or miR-NC + TMZ. In addition, the expression of MGMT, 32 an enzyme that repairs damaged DNA, was downregulated (Figure 5(c)).

MiR-105 increases the chemosensitivity of glioma cells to TMZ treatment. (a) Cell viability was evaluated in U87 and U251 cells with or without TMZ treatment at different doses; miR-105 overexpression in cells significantly increased the chemosensitivity to treatment of TMZ (*p < 0.05 and **p < 0.01). (b) Cell proliferation in the presence of TMZ (100 µM) was tested by the CCK-8 assay. The capacity for proliferation was inhibited in cells overexpressing miR-105, whether the cells had been treated with TMZ or not (MC, medium change only), and SUZ12 overexpression restored the inhibitory effects of miR-105 (*p < 0.05 and **p < 0.01). (c) MiR-105 overexpression in U251 cells decreased MGMT and increased cleaved caspase-3 at the protein level, while SUZ12 showed the opposite effects. GAPDH was used as a loading control.
Discussion
MiRNAs, a class of small regulatory RNAs, can function as tumor suppressor genes or oncogenes, regulating many cellular events, such as proliferation, invasion, angiogenesis, apoptosis, stress responses, differentiation, and development.5,6,33 Dysregulated miRNAs have been observed in various kinds of tumor, including gliomas. Recent studies have also shown that miR-105 is potentially a tumor suppressor in many kinds of cancer.34,35 In our study, we found that miR-105 was downregulated in human glioma tissues and glioma cell lines compared with its level in normal brain tissues and NHAs. Moreover, we demonstrated that the overexpression of miR-105 in glioma cells can inhibit cell proliferation in vitro, induce G1-phase arrest, attenuate subcutaneous tumor growth in vivo, suppress cell invasion and migration, and increase chemosensitivity. In particular, analysis of in vivo subcutaneous tumor growth revealed a significant decrease when using miR-105-transfected cells, indicating its therapeutic potential for patients with glioma.
SUZ12, together with Zeste 2 enhancer and embryonic ectoderm development, is a component of the PcG complex, which inactivates target gene promoters to regulate cell proliferation, differentiation, and aging. 36 SUZ12 is critical for tumor pathogenesis and development 20 and may be involved in the regulation of tumor stem cells. 37 Previous studies demonstrated high SUZ12 levels in aggressive tumors, for example, in ovarian cancer, mantle cell lymphoma, and breast cancer,19–21 and demonstrated a marked correlation between SUZ12 levels and tumor malignancy. However, to the best of our knowledge, no studies have analyzed the role or mechanism of action of SUZ12 in glioma. Our results showed that SUZ12 is a direct target of miR-105 in glioma cells not only in vitro but also in vivo. First, luciferase reporter assays confirmed that miR-105 directly recognizes the 3′-UTR of SUZ12 transcripts. Second, SUZ12 expression was significantly decreased in glioma cells stably expressing miR-105. Third, an inverse correlation between SUZ12 protein and miR-105 levels in clinical samples was found. Finally, the inhibition of SUZ12 protein synthesis by miR-105 inhibited subcutaneous tumor growth, according to the results of immunohistochemistry and western blotting. As a direct target of miR-105, SUZ12 displayed an important functional role affecting the proliferation, cell cycle, migration, and invasion of U87 and U251 glioma cell lines.
In summary, we found that miR-105 is significantly downregulated in glioma tissues and cell lines, and we identified a link between miR-105 and SUZ12, which is a novel factor in glioma tumorigenesis. Moreover, we demonstrated for the first time the role of miR-105/SUZ12 in regulating glioma cell proliferation, cell cycle, subcutaneous tumor growth, invasion, migration, and drug sensitivity. This newly identified miR-105/SUZ12 regulation provides a new insight into the mechanisms underlying glioma development, and targeting miR-105/SUZ12 may represent a promising therapeutic strategy for glioma.
Footnotes
Acknowledgements
J.Z., W.W., and S.X. contributed equally to this work.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the Research Special Fund for Public Welfare Industry of Health (201402008), National Natural Science Foundation of China (81472362, 81372709, and 81302185), Jiangsu Province’s Natural Science Foundation (BK20131019 and BK20151585), the Program for Advanced Talents within Six Industries of Jiangsu Province (2015-WSN-036 and 2016-WSW-023), and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
