Abstract
Promoter methylation reflects in the inactivation of different genes like O6-methylguanine-DNA methyltransferase DNA repair gene and runt-related transcription factor 3, a known tumor suppressor gene in various cancers such as esophageal cancer. The promoter methylation was evaluated for O6-methylguanine-DNA methyltransferase and runt-related transcription factor 3 in CpG, CHH, and CHG context (where H is A, T, or C) by next-generation sequencing. The methylation status was correlated with quantitative messenger RNA expression. In addition, messenger RNA expression was correlated with different risk factors like tobacco, alcohol, betel nut consumption, and smoking habit. CpG methylation of O6-methylguanine-DNA methyltransferase promoter had a positive association in the development of esophageal cancer (p < 0.05), whereas runt-related transcription factor 3 promoter methylation showed no significant association (p = 1.0) to develop esophageal cancer. However, the non-CpG methylation, CHH, and CHG were significantly correlated with O6-methylguanine-DNA methyltransferase (p < 0.05) and runt-related transcription factor 3 (p < 0.05) promoters in the development of esophageal cancer. The number of cytosine converted to thymine (C→T) in O6-methylguanine-DNA methyltransferase promoter showed a significant correlation between cases and controls (p < 0.05), but in runt-related transcription factor 3 no such significant correlation was observed. Besides, messenger RNA expression was found to be significantly correlated with promoter hypermethylation of O6-methylguanine-DNA methyltransferase and runt-related transcription factor 3 in the context of CHG and CHH (p < 0.05). The CpG hypermethylation in O6-methylguanine-DNA methyltransferase showed positive (p < 0.05) association, whereas in runt-related transcription factor 3, it showed contrasting negative association (p = 0.23) with their messenger RNA expression. Tobacco, betel nut consumption, and smoking habits were associated with altered messenger RNA expression of O6-methylguanine-DNA methyltransferase (p < 0.05) and betel nut consumption and smoking habits were associated with runt-related transcription factor 3 (p < 0.05). There was no significant association between messenger RNA expression of O6-methylguanine-DNA methyltransferase and runt-related transcription factor 3 with alcohol consumption (p = 0.32 and p = 0.15). In conclusion, our results suggest that an aberrant messenger RNA expression may be the outcome of CpG, CHG, and CHH methylation in O6-methylguanine-DNA methyltransferase, whereas outcome of CHG and CHH methylation in runt-related transcription factor 3 promoters along with risk factors such as consumption of tobacco, betel nut, and smoking habits in esophageal cancer from Northeast India.
Keywords
Introduction
Esophageal cancer (EC) is the eighth most common cancer in the world, 1 with an estimated 455,800 new EC cases and 400,200 EC-caused deaths occurred in 2012 worldwide. In India, the incidence of EC is moderately high, and it is the second most common cancer among males and the fourth most common cancer among females. 2 The highest occurrence of EC has been reported from Assam in the northeast region. In this population, the promoter hypermethylation of certain genes may contribute toward greater risk at the molecular level for EC. 3 It has been reported that exposure to different carcinogens, namely, tobacco, alcohol, and betel nut consumption or smoking are associated with increased DNA methylation at certain genes. In Southeast Asia, smoking, alcohol consumption, and chewing of betel nut with or without tobacco are the major causes of oral cancer. 4
DNA methylation is the addition of a methyl (−CH3) group to nitrogenous bases. In humans, very often, high density of 5-methylcytosine has been found in the promoter regions of several genes. 5 Gene silencing through methylation has been described in genes involved in tumor suppression, DNA repair, maintenance of genome integrity, as well as tumor invasion and apoptosis. 6 Hypermethylation of CpG islands of gene promoters leading to transcriptional repression is one of the most common causes of cancer. 7 Association of messenger RNA (mRNA) expression and promoter methylation has been evaluated in various cancers. 8 The presence of hypermethylation has been associated with abnormal mRNA expression in esophageal squamous cell carcinoma (ESCC) 9 and stomach cancer. 8 Besides methylation of CpG islands, recent studies have shown the presence of 5-methylcytosine in human somatic and embryonic stem cells in CHH and CHG contexts (where H is A, T, or C).10,11 The non-CpG methylation mediated gene silencing has been observed in a subset of acute lymphatic leukemia (ALL) patients. 12 The occurrence of unique non-CpG methylation in a regulatory region of the sex determining region Y (SRY) gene has also been reported, which is unstable and promote gene expression. 13
O6-methylguanine-DNA methyltransferase (MGMT) is a direct DNA repair enzyme that removes alkyl adducts from the O6 position of guanine in DNA. MGMT prevents the mutagenic, cytotoxic, and carcinogenic effects of various alkylating agents. 14 MGMT’s inimitability is its ability to stoichiometrically repair DNA adducts (methyl group) and to self-inactivate which is also called “suicidal activity.” 14 The activity of MGMT gene is interrupted by hypermethylation in its promoter region which is mostly associated with poor prognosis and metastasis of cancer.15–17 The interruption of MGMT activity is due to hypermethylation reflected in its mRNA level of expression. mRNA expression of MGMT also indicates a direct mediating role in tumor sensitivity to different alkylating agents. 18
Runt-related transcription factor 3 (RUNX3) is a transcription factor of the transforming growth factor (TGF)-β family. It acts as a tumor suppressor gene and is involved in the TGF-β pathway through its interaction with SMADs (a family of proteins similar to the gene products of the Drosophila gene ‘mothers against decapentaplegic’ (Mad) and the C. elegans gene Sma) and P-300 proteins. RUNX3 mediates tumor suppression by cell cycle arrest, inducing apoptosis, and lowering angiogenesis. 19 RUNX3 methylation induces altered gene expression and plays an important role in disease progression in EC. 20 Several previously conducted studies reported RUNX3 mRNA expression to be reduced due to CpG hypermethylation in a variety of human malignancies, including gastric, 21 lung, 22 esophageal, 23 colon, 24 and hepatocellular carcinoma, 25 whereas, the non-involvement of RUNX3 methylation was reported in nasopharyngeal carcinoma. 26
A contrasting epigenetic role has been reported for RUNX3 and MGMT methylation in glioblastoma 27 which may also be valid for the development of EC in Northeast (NE) Indian population. This study attempts (a) to compare MGMT and RUNX3 gene promoter methylation, (b) to correlate the methylation status of MGMT and RUNX3 genes with their mRNA expression, (c) to evaluate the association of MGMT and RUNX3 mRNA expression with known risk factors in EC patients from NE India, and (d) to validate our results with trends observed in previously published data.
Materials and methods
Sample collection
Tissue samples were collected from each voluntarily participating subject diagnosed with EC based on endoscopic and histopathological evaluation with informed consent from Department of Gastroenterology, Gauhati Medical College and Hospital, Guwahati. Healthy volunteers were recruited as age and sex matched controls based on endoscopic and histopathological evaluation. History of lifestyle habits like tobacco, betel nut, alcohol consumption, and smoking was taken from both EC cases and control subjects. The collected tissue samples were stored in RNA later solution (Ambion, USA) at −80°C. Formalin-fixed, paraffin-embedded (FFPE) tissue blocks of the samples were collected from Department of Pathology, Gauhati Medical College and Hospital, Guwahati, for histopathological evaluation. Patients were categorized based on the dysphagia grade. The degree of dysphagia was determined using the modified O’Rourke grading system 28 (Table 1)
The modified O’Rourke dysphagia staging system (stage swallowing status).

DNA extraction
DNA isolation was done from the confirmed EC tissue by phenol–chloroform method. The isolated DNA was quantified by Qubit® dsDNA BR Assay Kit (Thermo Fisher Scientific, USA).
Methylation analysis by next-generation sequencing
The methylation status was determined by bisulfite sequencing primers (BSP): forward 5′-TGGTAAATTAAGGTATAGAGTTTTAGG-3′ and reverse 5′-AAAACCTAAAAAAAACAAAAAAAC-3′ with product size 191 bp for MGMT and forward 5′-GGTTTTGGGTTGTGGTATTG-3′ and reverse 5′-AACAAATCCT CCAAAATCAAATAAC-3′ with product size 109 bp for RUNX3 by next-generation sequencing (NGS). Genomic DNA (gDNA) samples were quantified using Qubit® dsDNA BR Assay Kit (Thermo Fisher Scientific, USA). EZ DNA Methylation-Gold™ Kit (Zymo research, USA) was used for bisulfite conversion using 500 ng gDNA per sample. Amplification of polymerase chain reaction (PCR) products was confirmed by agarose gel electrophoresis. The amplified PCR products were purified using Invitrogen PureLink PCR Purification Kit (Invitrogen, USA). Prepared libraries were confirmed for bisulfite conversion using Agilent 2100 Bioanalyzer (Agilent, USA).
NGS library preparation—In this step, the PCR products were end-repaired and ligated with unique barcodes. Massively parallel sequencing of barcode ligated PCR products was carried out using MiSeq® Reagent Kit v3 (Illumina, USA) at depth of 500×. After running the sequencing, an adapter trimming tool, Trim-galore (Version 0.4.0, Babraham Bioinformatics), was used to inspect and remove adapter contamination following which a quality control step was performed using the Fastqc tool (Version 0.11.4, Babraham Bioinformatics). The generated Fastqc files were used for analysis. The processed reads were aligned to the region of interest using Bismark (Version 0.16.0, Babraham Bioinformatics). Bismark is a program capable of mapping bisulfite treated sequencing reads to a genome or regions of interest in the genome and performs methylation calls in a single step. 29 The aligned files were de-duplicated to remove duplicated read alignments. The methylation calls were then contextualized as CpG, CHG, and CHH.
Quantitative real-time PCR
RNA was extracted from 25 EC and 20 normal tissue samples using the AllPrep DNA/RNA Extraction Kit (Qiagen, Germany). A total of 1 µg of RNA was converted into complementary DNA (cDNA) using SuperScript II Reverse Transcriptase (Thermo Fisher Scientific, USA). These converted cDNAs were then used for mRNA expression study in real-time PCR. All samples were run in triplicate in Rotor-Gene Q (Qiagen, Valencia, CA), and their results were normalized with housekeeping gene β-actin. The primer sequences for MGMT gene were 5′-GCTGAATGCCTATTTCCACCA-3′(forward) and 5′-CACAACCTTCAGCAGCTTCCA-3′ (reverse; product size: 123 bp) and for RUNX3 gene were 5′-GACTGTGATGGCAGGCAATGA-3′ (forward) and 5′ CGAAGCGAAGGT CGTTGAA-3′ (reverse; product size: 101 bp). The real-time amplification consisted of an initial denaturation of 5 min at 95°C, followed by 40 cycles of 95°C for 20 sec, 59.6°C for 20 sec, 72°C for 30 sec for MGMT (Figure 1 (a)), or 40 cycles of 95°C for 15 sec, 58°C for 15 sec, 72°C for 25 sec for RUNX3 (Figure 2(a)) and a final extension at 72°C for 5 min following a melting curve analysis from 55°C to 95°C for both MGMT (Figure 1(b)) and RUNX3 (Figure 2(b)).
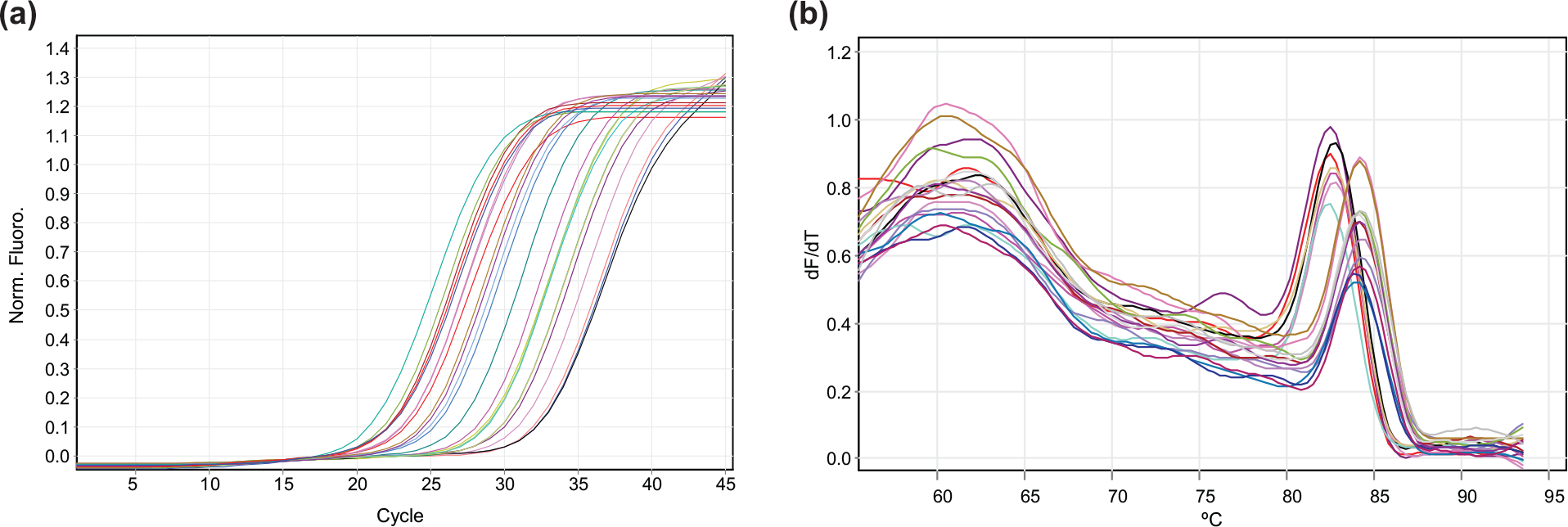
Representative (a) amplification curve (b) and melt curve for MGMT mRNA expression in real-time PCR.

Representative (a) amplification curve (b) and melt curve for RUNX3 mRNA expression in real time PCR.
Statistical analysis
Statistical software Epi Info version 3.3.2 (Centers for Disease Control and Prevention (CDC), Atlanta, Georgia, USA) was used to analyze the data. In this study, Mantel–Haenszel test was considered to compare the distribution of different variables between cases and controls. The odds ratio (OR) at 95% confidence intervals (CIs) was considered to determine the strength of association between different parameters. The statistical software SPSS (Version 16.0, IBM, USA) was used for descriptive statistics to find out the correlation between MGMT methylation and mRNA expression.
Results
This case-control study was conducted with 25 tissue samples and 20 control samples from NE India. Male:female ratio was 2:1 with a mean age of 59.6 ± 10.7 years. The mean age was 62.0 ± 9.2 years for males and 56.0 ± 11.9 years for females. The majority of the cases were from Kamrup district, Assam (71%). The promoter methylation status of MGMT and RUNX3 was evaluated by NGS technique for the DNA extracted from EC tissue samples after bisulfite conversion. In addition, we tried to examine the mRNA expression of MGMT and RUNX3 in correlation with their methylation status and different risk factors such as tobacco, betel nut, alcohol consumption, and smoking habits.
MGMT CpG methylation
The CpG methylation status of MGMT was evaluated for 25 EC cases where methylation was found to be significantly associated (p < 0.05, χ2 = 40.88, OR = 7.88) with the development of EC in relation to control samples (Table 3). MGMT promoter methylation was found to be 84% in EC patients (Table 2). While considering different characteristics of patients, a difference in methylation status of males and females was observed (male: 94.1%, female: 62.5%) in different age groups (age >50 years: 93.3%, age <50 years: 70%) as well as in different dysphagia grades (Grade 1: 40%, Grade 2: 80%, Grade 3: 100%, Grade 4: 100%, and Grade 5: 100%; Table 2).
Characteristics of patients and their MGMT and RUNX3 promoter methylation status in CpG context.
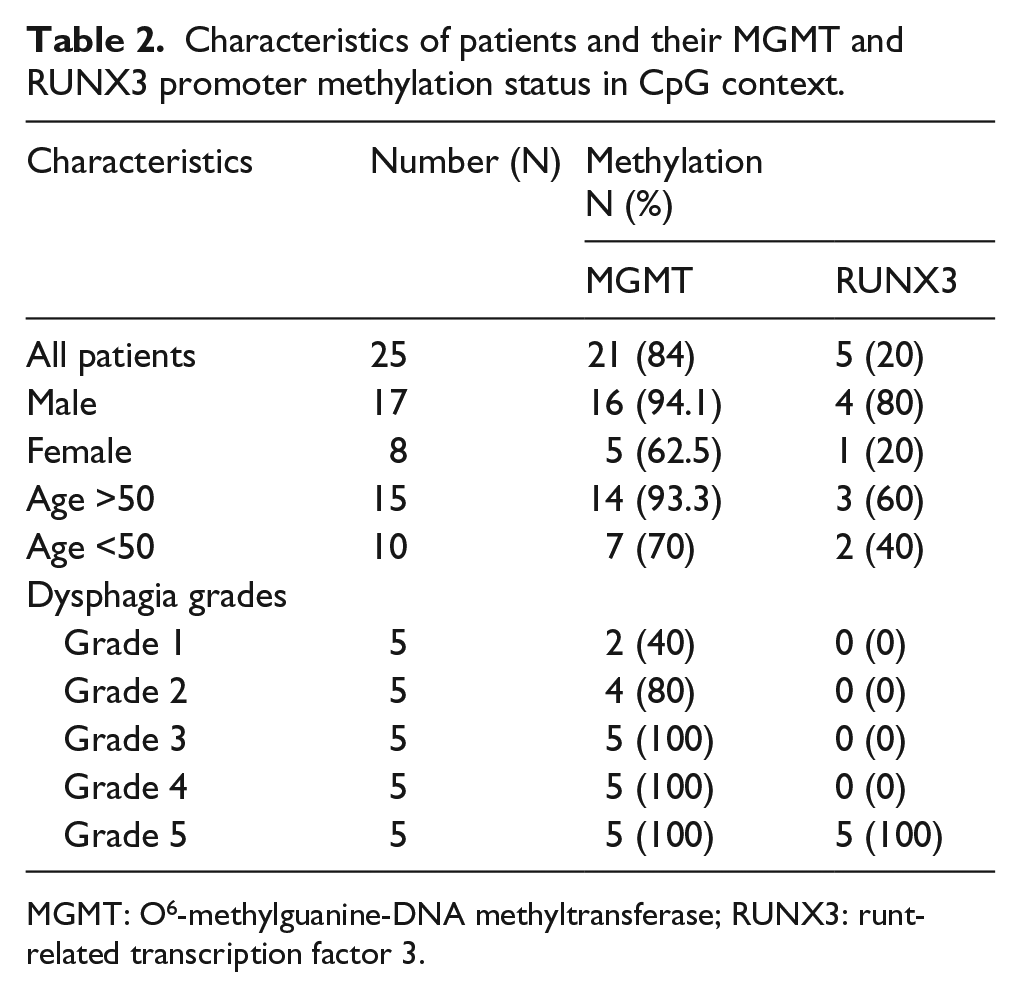
MGMT: O6-methylguanine-DNA methyltransferase; RUNX3: runt-related transcription factor 3.
RUNX3 CpG methylation
Findings showed no significant association of CpG promoter methylation of RUNX3 in the development of EC (p = 1.00, χ2 = 0.00, OR = 1.00; Table 3). NGS findings revealed the occurrence of methylation in RUNX3 promoter in 20% of the cases (Table 2). Different patient characteristics were analyzed in accordance with methylation status. RUNX3 methylation was more in males than in females (male: 80%, female: 20%). Higher CpG RUNX3 methylation was observed in age group more than 50 years (60%) compared to the age group lower than 50 years (40%). Among all dysphagia grades, methylation was observed only in Grade 5 whereas all the other grades (Grade 1, 2, 3, and 4) showed no methylation (Table 2).
CpG, CHG, and CHH methylation status in cases versus control for MGMT and RUNX3 promoters.

MGMT: O6-methylguanine-DNA methyltransferase; RUNX3: runt-related transcription factor 3; OR: odds ratio.
Bold values indicate statistical significance.
CHH and CHG methylation
NGS revealed additional asymmetric methylation in CHH islands and symmetric methylation in CHG along with CpG islands. Analysis of CHG methylation status between cases and controls showed a significant difference for both MGMT (p < 0.05, χ2 = 22.11, OR = 0.00) and RUNX3 (p < 0.05, χ2 = 12.06, OR = 0.17). However, CHH methylation status showed a significant difference in MGMT (p < 0.05, χ2 = 7.96, OR = 2.25) and RUNX3 (p < 0.05, χ2 = 22.11, OR = 0.00) promoters between case and control groups (Table 3).
Conversion of cytosine to thymine
Conversion of cytosine to thymine is the main consequence of methylation. Our study revealed a varying degree of cytosine to thymine conversions in CpG, CHG, and CHH context of methylation. MGMT promoter showed a significant difference in the conversion of cytosine to thymine in CpG, CHH, and CHG islands (p < 0.05). Whereas in RUNX3 promoter there was no significant difference of cytosine to thymine conversion in CpG, CHG, and CHH context between cases and controls (p = 0.08, p = 0.69, and p = 0.69; Table 4)
The conversion of cytosine in the context of CpG, CHG, and CHH islands in MGMT and RUNX3 promoters.

MGMT: O6-methylguanine-DNA methyltransferase; RUNX3: runt-related transcription factor 3; CI: confidence interval.
Bold values indicate statistical significance.
CpG, CHG, and CHH promoter methylation of MGMT and mRNA expression
The MGMT expression in EC cases was found to be downregulated with mean ± standard deviation (SD), 0.65 ± 0.72, relative to β-actin expression. While considering CpG methylation, out of four unmethylated samples, one showed downregulation and the rest three showed upregulation with mean ± SD, 1.95 ± 0.48, relative to β-actin expression. Whereas out of 21 methylated samples, 17 showed downregulation with mean ± SD, 0.06 ± 0.32, and the rest 4 showed upregulation with fold change, mean ± SD, 1.25 ± 0.68, relative to β-actin expression. Statistically, a significant association was observed for mRNA expression of MGMT with its methylation status (p < 0.05, χ2 = 5.01, OR = 12.75; Table 5). Similarly, methylation in CHG and CHH context was also observed to have a significant association with mRNA expression in EC cases (p < 0.05, χ2 = 6.52, OR = 18.00 and p < 0.05, χ2 = 6.56, OR = 11.00; Table 5).
Correlation of CpG, CHG, and CHH methylation and mRNA expression in MGMT and RUNX3 promoters.

mRNA: messenger RNA; MGMT: O6-methylguanine-DNA methyltransferase; RUNX3: runt-related transcription factor 3; OR: odds ratio.
Bold values indicate statistical significance.
CpG, CHG, and CHH promoter methylation of RUNX3 and mRNA expression
We observed that the expression of RUNX3 mRNA is significantly increased in tumors with fold change, 1.14 ± 1.49, relative to β-actin expression in tumor tissues. mRNA expression of RUNX3 was found to be elevated in all EC cases, but upon categorizing the cases based on CpG methylated and unmethylated, out of five meth-ylated cases, four were found to be upregulated with mean ± SD, 0.75 ± 0.54, and one case was downregulated. However, out of 20 unmethylated cases, 10 showed upregulation with mean ± SD, 2.04 ± 1.12, and the rest 10 cases showed downregulation with mean ± SD, 0.84 ± 1.89, relative to β-actin expression in tumor tissues. Statistically, no significant association was observed while correlating the CpG methylation status of RUNX3 promoter and mRNA expression level (p = 0.23, χ2 = 1.40, OR = 4.00; Table 5). Whereas mRNA expression was significantly correlated with CHG and CHH methylation of RUNX3 promoter in tumor tissues (p < 0.05, χ2 = 5.25, OR = 0.00 and p < 0.05, χ2 = 6.50, OR = 0.00; Table 5).
Correlation of risk factors and mRNA expression of MGMT and RUNX3
The known risk factors included in the study were tobacco, alcohol, betel nut consumption, and smoking habits. mRNA expression of MGMT showed significant association with tobacco (p < 0.05, χ2 = 7.96, OR = 2.25), betel nut consumption (p < 0.05, χ2 = 62.51, OR = 12.67), and smoking (p < 0.05, χ2 = 50.25, OR = 9.33). While correlating RUNX3 mRNA expression with same risk factors, a significant association was observed with betel nut (p < 0.05, χ2 = 44.0, OR = 7.71) and smoking (p < 0.05, χ2 = 40.88, OR = 7.88). Statistically, no significant association was observed between mRNA expression of MGMT and RUNX3 with alcohol consumption (p = 0.32, χ2 = 0.98, OR = 0.76 and p = 0.15, χ2 = 2.01, OR = 0.67) in NE Indian population (Table 6).
Correlation between mRNA expression of MGMT and RUNX3 genes and different risk factors associated with EC development.

mRNA: messenger RNA; MGMT: O6-methylguanine-DNA methyltransferase; RUNX3: runt-related transcription factor 3; EC: esophageal cancer; OR: odds ratio.
Bold values indicate statistical significance.
Discussion
NE India is well known global biodiversity hotspot located in the eastern region of the Himalayas, and its genetic diversity gives a sound importance to population-based case-control studies. 30 NE region of India has the highest incidence of EC when compared to other regions of the country with an increasing trend.31,32 In our study, we evaluated the promoter methylation status of MGMT and RUNX3 promoter in EC cases and controls by NGS and correlated with quantitative estimation of mRNA expression. The mRNA expression was further correlated with known risk factors such as tobacco, betel nut, alcohol consumption, and smoking habits. MiSeq sequencing system—Illumina technology was used for NGS that provides the highest accuracy over other methylome analytical methods.33,34 NGS provides the additional advantage of determining the non-CpG (CHG and CHH) methylation status along with CpG.
MGMT mediated DNA repair is the simplest form of DNA repair which is the direct removal of the methyl adducts from altered gDNA. 35 It removes alkyl groups from O6-methylguanine, appended as a result of alkylation induced by S-adenosylmethionine, different chemical agents, or dietary nitrosamines. The human MGMT gene promoter spans 762 bp (−531 to 231 from the ATG) containing 98 CpG sites. 36 However, classical tumor suppressor genes are inactivated mainly by genetic alterations on both alleles; in contrast, RUNX3 is inactivated by DNA methylation which is a reversible epigenetic modification. 37 Although EC has the highest incidence in this region, till date there is no reported data on methylation status of RUNX3 gene in the population of NE India.
The percentage of CpG methylation was observed to be more in males than in females in both MGMT and RUNX3 gene promoters. As a result, males may be more prone to develop EC in this population. Patients with higher age (>50 years) were more prone to methylation in MGMT and RUNX3 promoters. This may be because of exposure for a long duration to the risk factors that were being analyzed and may lead to the inactivity of MGMT gene to remove methyl adduct from gDNA. While the tumor suppressor gene RUNX3 which is more active in EC with lower age group as there is no methylation may have an additive effect on the progression of cancer cases to metastasis (data not shown). Similarly, the percentage of CpG methylation was observed to increase with higher dysphagia grades for both the genes which postulate that inactiveness of MGMT and RUNX3 gene after longer exposure to various carcinogens.
Our study showed a significant difference in CpG promoter methylation of MGMT promoters between EC cases and controls. In support of our findings, previous studies reported methylation of MGMT promoters in various types of cancers such as gastric cancer (36.8%), colorectal cancer (46%), lung cancer (21%), glioma (40%), head and neck cancers (60.8%), along with EC (44%). 38 Zhao et al. 39 conducted a meta-analysis reporting the contribution of abnormal promoter methylation to the risk of EC. Whereas, contradictory findings also have been reported in Turkish population where MGMT promoter methylation showed no prognostic value in primary glioblastoma. 40
The significant association of CpG methylation of RUNX3 promoter was reported with increased risk and progression in EC 20 and gastric cancer. 41 While, our study showed no significant correlation in CpG promoter methylation of RUNX3 between EC cases and controls. In support of this finding, previous data revealed the non-involvement of RUNX3 promoter methylation as a potent risk factor in the Helicobacter pylori–associated gastric cancer in the South Indian population 42 and in nasopharyngeal carcinoma. 26 The promoter methylation of MGMT and RUNX3 showed an opposite association between EC cases and control in NE population.
The overall MGMT mRNA expression level was found to be downregulated in EC cases, and it showed a positive correlation with CpG methylation. Our findings corroborate the previous data reported in glioblastoma. 43 Another supporting data stated the significant difference of MGMT mRNA expression between oral squamous cell carcinoma cases and controls in a South–Eastern Iranian population. 44 The overall mRNA expression of RUNX3 was upregulated in our findings with a negative correlation with CpG methylation of RUNX3 promoter. While previous study reported the significant alteration of RUNX3 mRNA expression and promoter hypermethylation status in cancer patients. 45 Another study also showed a significant correlation between downregulated mRNA expression and methylation of RUNX3 promoter in ovarian cancer. 46
The MGMT promoter methylation was observed in all the dysphagia grades that reflected in the downregulation of mRNA expression, whereas RUNX3 promoter methylation was observed only in dysphagia Grade 5. This may be one of the reasons for the overall increase in the RUNX3 mRNA expression.
A contrasting trend was observed between MGMT and RUNX3 gene in EC cases and control from NE India. The overall occurrence of methylation in CpG and non-CpG was more in MGMT than RUNX3 promoter. The correlation between mRNA expression and methylation status in CpG context was opposite in MGMT and RUNX3. Previous data supports the similar contrasting methylation status between MGMT and RUNX3 in glioblastoma. 27
In our study population, the CpG promoter methylation of RUNX3 did not contribute to EC, which indicated the involvement of promoter non-CpG methylation. In earlier studies, the mammalian DNA methylation was restricted to symmetrical CpG sequences, but the present focus has shifted to non-CpG sequences. DNA contains a significant number of cytosine methylation in both symmetric CHG context (where H is A, C, or T) and in asymmetric CHH context (where H is A, C, or T). 47 Significant differences were observed in CHG and CHH methylation between cases and controls in MGMT and RUNX3 genes with the development of EC. In addition, the CHG and CHH methylation showed a significant correlation with mRNA expression of MGMT and RUNX3 in EC.
The consequence of methylation is the conversion of cytosine to thymine, which occurs in varying degrees in EC cases and controls. In MGMT promoter methylation, the number of cytosine converted to thymine (C→T) was found to be significantly different between cases and controls in CpG and non-CpG context at 95% CI. In RUNX3 promoter methylation, no significant number of conversions of cytosine to thymine (C→T) was observed between cases and controls.
In India, tobacco, betel nut, alcohol consumption, and smoking are common risk factors for cancer. The betel nut and tobacco consumption have been reported as common risk factors associated with EC in Assamese population of NE India. 48 Our findings revealed the association of MGMT mRNA expression with tobacco, betel nut consumption, and smoking habits, whereas mRNA expression of RUNX3 showed significant association with betel nut consumption and smoking habit. Alcohol was not associated with the mRNA expression of MGMT and RUNX3.
Conclusion
The alteration of mRNA expression of MGMT and RUNX3 genes in EC distinctly varies in correlation with methylation status in CpG and non-CpG (CHG and CHH) context. MGMT showed a positive correlation between mRNA expression and its methylation status in both CpG and non-CpG context. RUNX3 provides a significant correlation in CHG and CHH context but not in CpG context. Thus, aberrant methylation of RUNX3 promoter in non-CpG context could be a useful adjunct to histopathological evaluation in the prediction of the risk of EC in NE Indian population. Tobacco, betel nut consumption, and smoking habits were associated with altered mRNA expression of MGMT, and in terms of RUNX3, mRNA expression showed a significant association with only consumption of betel nut and smoking habits. Statistically, no significant association was observed between mRNA expression of MGMT and RUNX3 with alcohol consumption. A limitation of the study was that the DNA methylation sites other than the promoters were not evaluated. Moreover, the methylation status of MGMT and RUNX3 gene promoter may reflect its effect not only in transcriptional level but also in protein expression.
Footnotes
Acknowledgements
The authors thank Department of Biotechnology (DBT), Government of India for financial support; all the participants of this study and members of the Department of Bioengineering and Technology, Gauhati University Institute of Science & Technology, Gauhati University; and the Department of Gastroenterology, Gauhati Medical College and Hospital; especially all the members of Endoscopy OT. S.S. and A.U.R. contributed equally to the study.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical approval
All the laboratory methods and sample collection were done in accordance with relevant regulations and approved by Gauhati University ethical committee (GU/ACA/Ethics/2012/1564; dated 12 December 2012). Informed consent was obtained from all individual participants included in the study.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by DBT, Government of India (sanction number BCIL/NER-BPMC/2013/367; dated 26 March 2013).
