Abstract
Non-small-cell lung cancer is one of the most lethal cancers in the worldwide. Although Paclitaxel-based combinational therapies have long been used as a standard treatment in aggressive non-small-cell lung cancers, Paclitaxel resistance emerges as a major clinical problem. It has been demonstrated that Curcumin from Curcuma longa as a traditional Chinese medicine can inhibit cancer cell proliferation. However, the role of Curcumin in Paclitaxel-resistant non-small-cell lung cancer cells is not clear. In this study, we investigated the effect of Curcumin on the Paclitaxel-resistant non-small-cell lung cancer cells and found that Curcumin treatment markedly increased the sensitivity of Paclitaxel-resistant non-small-cell lung cancer cells to Paclitaxel. Mechanically, the study revealed that Curcumin could reduce the expression of metastasis-associated gene 1 (MTA1) gene through upregulation of microRNA-30c in Paclitaxel-resistant non-small-cell lung cancer cells. During the course, MTA1 reduction sensitized Paclitaxel-resistant non-small-cell lung cancer cells and enhanced the effect of Paclitaxel. Taken together, our studies indicate that Curcumin increases the sensitivity of Paclitaxel-resistant non-small-cell lung cancer cells to Paclitaxel through microRNA-30c-mediated MTA1 reduction. Curcumin might be a potential adjuvant for non-small-cell lung cancer patients during Paclitaxel treatment.
Keywords
Introduction
Non-small-cell lung cancer (NSCLC) is one of the most aggressive cancers around the world with high mortality and occurrence. Despite improvements in chemotherapy, radiotherapy, and surgery, the prognosis of NSCLC still remains poor. 1 Paclitaxel-based combinational chemotherapies exhibit a better curative effect for NSCLC patients.2,3 However, the resistance of NSCLC cells to Paclitaxel continues to be a major problem in clinic.4,5
Curcumin is an active phenolic compound extracted from the rhizome of the plant “Curcuma longa” as a traditional Chinese medicine. Extensive studies have revealed various biological functions of Curcumin.6–10 Reportedly, Curcumin is able to attenuate cancer cell proliferation and promotes apoptosis in vivo and in vitro.11–14 As an anti-cancer agent, Curcumin has been demonstrated to regulate transcriptional factors, signaling molecules, and microRNAs (miRNAs), such as signal transducer and activator of transcription 3 (STAT3), activation protein 1 (AP-1), Forkhead box O1 (FOXO1), nuclear factor kappa B (NF-κB), Akt, reactive oxygen species (ROS), miR-208, and miR-21.14–18 Our current study revealed that Curcumin could increase the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel in vitro. Further studies need to be done to explore the potential mechanism by which Curcumin enhances the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel.
Metastasis-associated gene 1 (MTA1) is ubiquitously expressed protein that markedly increases metastasis and aggressiveness of human cancers such as malignant pleural mesothelioma (MPM), colorectal cancer, hepatocellular carcinoma, gastric cancer, and prostate cancer.19–23 MTA1 has been reported to be upregulated in NSCLC and be involved in cancer cell growth.24,25 However, MTA1 expression can be downregulated as a target gene by drug-induced microRNA, and the reduction of MTA1 can inhibit the proliferation of cancer cells.26,27 Therefore, in this study, the role of MTA1 in Curcumin-enhanced sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel and its upstream regulation by microRNA were explored subsequently.
Materials and methods
Cell culture
NSCLC cell lines of A549 and H460 (ATCC, Manassas, VA, USA) was cultured in RPMI 1640 medium with 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, USA) and penicillin (100 U/mL). Cells were cultured at 37°C with 5% CO2. Paclitaxel-resistant NSCLC cell lines A549 and H460 were generated using previously described protocols.28,29 HEK293T cells were provided by American Type Culture Collection (ATCC) and maintained in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FBS (vol/vol), 100 µg/mL penicillin G, and 100 µg/mL streptomycin.
Generation of overexpression plasmid
The plasmid of pcDNA3.1/MTA1 was constructed by inserting the Open Reading Frame (ORF) of human MTA1 complementary DNA (cDNA) into the mammalian expression plasmid of pcDNA3.1. The MTA1 gene was amplified by polymerase chain reaction (PCR) from cDNA of normal human alveolar epithelial cells. Both PCR products and pcDNA3.1 vector were further digested with the two restriction enzymes of Hind III and BamH I and then ligated with T4 DNA ligase.
miRNA mimic and locked nucleic acid-anti-miRNA synthesis
The miR-30c-5p mimic, negative control mimic, and mutant miR-30c-5p mimic were purchased from GenePharma (Shanghai, China). Locked nucleic acid (LNA)-anti-miR-30c-5p and negative control LNA were purchased from Exiqon (Vedbæk, Denmark).
Short interfering RNA synthesis
To silence human MTA1 gene, a short interfering RNA (siRNA) sequence (GGGAGGATTTCTTCTTCTATTCT) against human MTA1 messenger RNA (mRNA) was designed with siDirect (http://sidirect2.rnai.jp/) and synthesized by GenePharma. Meanwhile, a universal negative control siRNA (ATTCTTGACGTGTTACTGTTGTC) was also produced by GenePharma.
3′-untranslated region luciferase reporter construction and luciferase assays
The plasmid of pGL3-Promoter/wild-type (WT)-MTA1 (pGL3-Promoter/WT-MTA1) was constructed by inserting the 460 kb 3′-untranslated region (UTR) of human MTA1 mRNA into pGL3-Promoter vector at the Xba I restriction enzyme site. Mutation of MTA1 3′-UTR was introduced into the miRNA-binding site by the QuikChange Mutagenesis Kit (TaKaRa, Kusatsu, Japan), and the plasmid of pGL3-Promoter/mutant-MTA1 was constructed successfully. For WT or mutant MTA1 3′-UTR reporter assay, HEK293T cells in each well of 24-well cell culture plates were transfected with a mixture of 0.5 µg pGL3-Promoter/MTA1, 2.5 ng pRL-SV40, and 100 nmol miRNA mimic by using Lipofectamine 2000 (Invitrogen) according to the instructions. Cells were lysed at 48 h after transfection, and then, the luciferase activity of MTA1 3′-UTR reporter was measured through dual-luciferase reporter gene assay.
Isolation of total RNA and quantitative reverse transcription polymerase chain reaction
Total RNA was extracted from cells using TRIzol (Invitrogen), and then, both miRNA and mRNA were reversely transcribed to cDNA. The stem-loop primer for miR-30c was 5′-GTCGTATCCAGTGCAGGGTCCGAGTATTCGCACTGGATACGACGCTGA-3′. U6 small nuclear RNA was used for normalization. PCR reactions were performed with the following primers: for miR-30c, forward 5′-GCCGCTGTAAACATCCTACACT-3′ and reverse 5′-GTGCAGGGTCCGAGGT-3′ and for U6, forward 5′-CTCGCTTCGGCAGCACA-3′ and reverse 5′-AACGCTTCACGAATTTGCGT-3′. Relative expression level of MTA1 mRNA was examined by SYBR Green real-time PCR and normalized to glyceraldehyde 3-phosphate dehydrogenase (GAPDH). The primers were as follows: for MTA1, forward 5′-AGCTACGAGCAGCACAACGGGGT-3′ and reverse 5′-CACGCTTGGTTTCCGAGGAT-3′ and for GAPDH, forward 5′-CGTGGGCCGCCCTAGGCACCA-3′ and reverse 5′-TTGGCTTAGGGTTCAGGGGGG-3′. Primers were synthesized by Genscript (Nanjing, China). PCR was performed by using the ABI 7500 Fast Real-Time PCR system (ABI, Foster City, CA, USA).
Western blot
Total cellular protein was prepared and quantified with a bicinchoninic acid (BCA) protein assay (Beyotime, Nantong, China). Protein was fractionated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene fluoride (PVDF) membrane. The PVDF membrane was blocked in 5% dry milk at room temperature for 1 h and immunostained with antibodies of anti-MTA1 (1:2000, ab50263; Abcam, Cambridge, UK) and anti-β-actin (1:5000, ab8226; Abcam) at 4°C overnight. After washing, the PVDF membrane was incubated with horseradish peroxidase (HRP)-conjugated anti-mouse IgG (1:2000, #7076; Cell Signaling Technology, Danvers, MA, USA) at room temperature for 1 h. All results were visualized through a chemiluminescent detection system (Thermo Fisher Scientific, Waltham, MA, USA) and then exposed with regular X-ray film. The integrated density of the band was analyzed by using the software of Quantity One (Bio-Rad, Hercules, CA, USA).
Cell survival assay
Cells were seeded into 96-well plates at 2000 cells/well. After different treatment, 500 µg/mL of 3-(4,5)-dimethylthiahiazo(-z-y1)-3,5-di-phenytetrazoliumromide (MTT; Sigma-Aldrich, St. Louis, MO, USA) was added into each well at 4 h before detection. 200 µL of dimethyl sulfoxide (DMSO) was added to each well to dissolve the precipitate at 10 min before detection. Finally, optical density (OD) value was measured at the wavelength of 490 nm. Cell survival rate was calculated with the formula: OD value of treated group/OD value of non-treated group × 100%.
Statistical methods
All statistical analyses were carried out by using SPSS 13.0 software. All data are expressed as mean ± standard deviation (SD). The statistical significance of the groups was defined as p < 0.05 and evaluated by one-way analysis of variance (ANOVA) with simultaneous multiple comparisons between groups by the Bonferroni method.
Results
Curcumin increases the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel
Paclitaxel-resistant A549 cells were cultured with Paclitaxel and Curcumin alone or together to examine whether Curcumin can increase the cytotoxicity of Paclitaxel to Paclitaxel-resistant A549 cells. The result showed that Paclitaxel had no significant cytotoxicity to Paclitaxel-resistant A549 cells (Figure 1(a) and (d)), but Curcumin had slight cytotoxicity to Paclitaxel-resistant A549 cells in a dose-dependent manner (Figure 1(b) and (e)). More importantly, the combinational use of Paclitaxel and Curcumin exhibited much stronger cytotoxicity to Paclitaxel-resistant A549 cells compared with Paclitaxel or Curcumin treatment alone (Figure 1(c) and (f)), indicating that Curcumin is able to increase the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel.

Curcumin increases the sensitivity of Paclitaxel-resistant A549 cells to Paclitaxel. (a and b) Paclitaxel-resistant A549 cells were cultured with (a) Paclitaxel and (b) Curcumin at different dose, respectively, and then MTT was used to detect the cell survival at 48 h after treatment (**p < 0.01 vs 0 µM group). (c) Paclitaxel-resistant A549 cells were cultured with 30 µM Paclitaxel or 30 µM Curcumin alone or the combination of 30 µM Paclitaxel and 30 µM Curcumin. MTT was used to detect the cell survival at 48 h after treatment (**p < 0.01 vs control group, ##p < 0.01 vs Paclitaxel group and Curcumin group). The data are from one experiment, representative of three independent experiments. Results were represented as means ± SD (n = 4 for each group). (d–f) The microscopic photographs of cells in (a–c) were shown.
Curcumin reduces the expression of MTA1 in Paclitaxel-resistant NSCLC cells
Given that MTA1 upregulation is closely related to NSCLC cell proliferation,24,25 to explore the potential mechanism by which Curcumin increases the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel, and the expression of MTA1 was further detected in Paclitaxel-resistant A549 cells after Curcumin treatment. The result showed that Curcumin obviously reduced the expression of MTA1 in Paclitaxel-resistant A549 cells at both mRNA and protein levels in a dose-dependent manner (Figure 2), suggesting MTA1 might be a downstream target of Curcumin in Paclitaxel-resistant NSCLC cells.
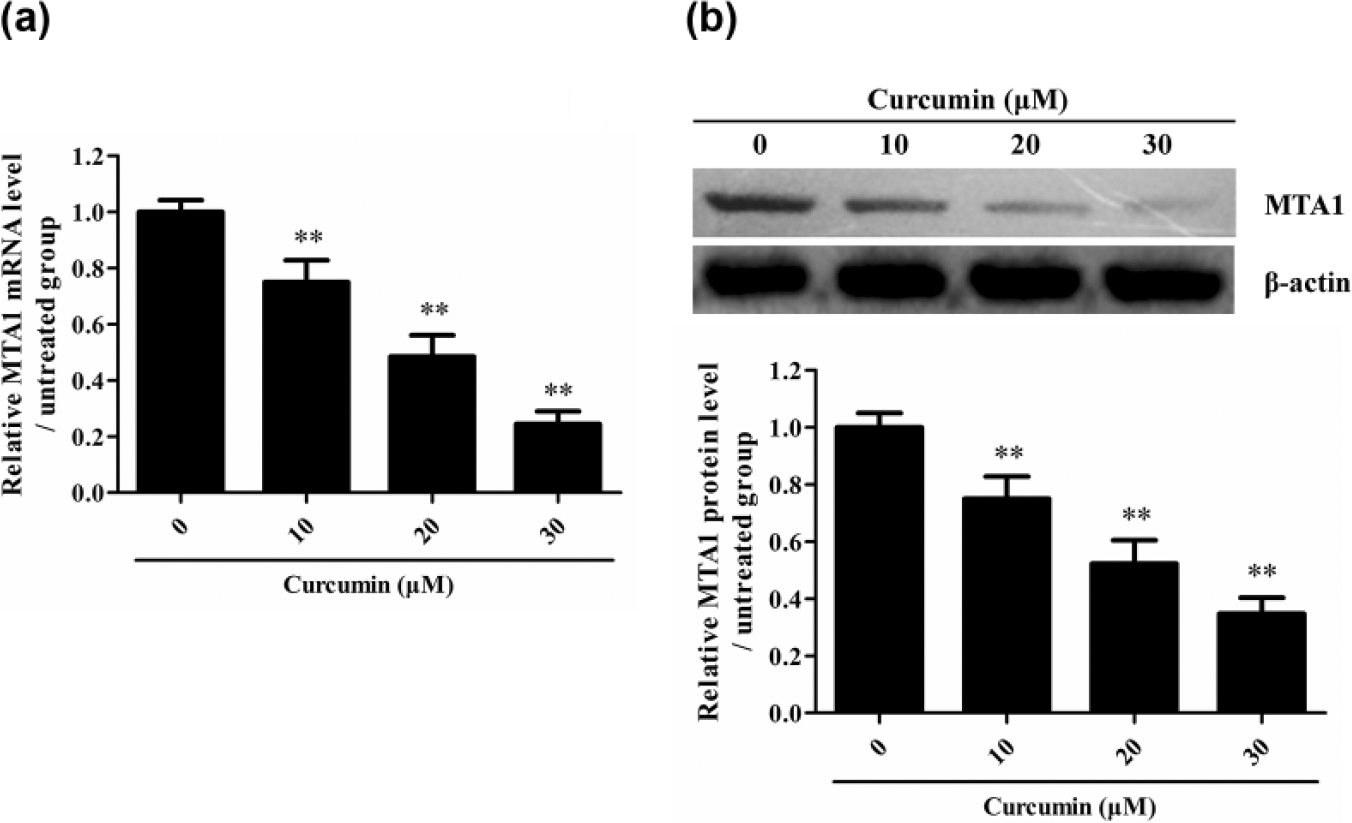
Curcumin reduces the expression of MTA1 in Paclitaxel-resistant A549 cells. Paclitaxel-resistant NSCLC cells were cultured with Curcumin at different dose for 48 h, and then, the mRNA and protein expression of MTA1 was detected by (a) real-time PCR and (b) Western blot, respectively (**p < 0.01 vs 0 µM group). The data are from one experiment, representative of three independent experiments. Results were represented as means ± SD (n = 4 for each group).
MTA1 reduction is involved in Curcumin-increased sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel
To check whether the downregulation of MTA1 contributes the Curcumin-increased sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel, siMTA1 was used to silence MTA1 gene, and then, the effect of Paclitaxel alone on Paclitaxel-resistant A549 cells was assayed. The result showed that knockdown of MTA1 gene could enhance the sensitivity of Paclitaxel-resistant A549 cells to Paclitaxel (Figure 3(a) and (b)). However, overexpression of MTA1 could markedly reduce Curcumin-enhanced sensitivity of Paclitaxel-resistant A549 cells to Paclitaxel (Figure 3(c) and (d)). These data indicate that MTA1 downregulation is involved in Curcumin-enhanced sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel.

Downregulation of MTA1 is involved in Curcumin increased the cytotoxicity of Paclitaxel to Paclitaxel-resistant A549 cells. (a) siMTA1 and siCTR were transfected into the Paclitaxel-resistant A549 cells for 48 h, and the protein level of MTA1 was detected by Western blot (**p < 0.01 vs siCTR group). (b) siMTA1 and siCTR were transfected into the Paclitaxel-resistant A549 cells for 24 h, and then, the cells were treated with 30 µM Paclitaxel for 48 h. Subsequently, MTT was used to detect the cell survival. (**p < 0.01 vs siCTR + Paclitaxel group). (c and d) pcDNA3.1-MTA1 and pcDNA3.1 were transfected into the Paclitaxel-resistant A549 cells for 48 h, and then, the cells were treated with 30 µM Paclitaxel for 48 h. Western blot was used to detect the protein level of (c) MTA1, and MTT was used to detect the (d) cell survival (**p < 0.01 vs pcDNA3.1 + Paclitaxel + Curcumin group). The data are from one experiment, representative of three independent experiments. Results were represented as means ± SD (n = 4 for each group). (e) siMTA1 and siCTR were transfected into the Paclitaxel-resistant A549 cells for 24 h, the cells were then treated with 30 μM Paclitaxel and/or 30 μM Curcumin for 48 h. MTT assay was performed to determine the cell survival (**p < 0.01 vs siCTR + Paclitaxel group).
Curcumin reduces the expression of MTA1 gene through upregulation of miR-30c-5p in Paclitaxel-resistant NSCLC cells
To explore the potential mechanism by which Curcumin reduces the expression of MTA1 gene, the microRNA targeted at human MTA1 3′-UTR was predicted with TargetScanHuman (http://www.targetscan.org/vert_70/). We got five potential miRNAs targeted at human MTA1 3′TR, including miR-30a-5p, miR-30b-5p, miR-30c-5p, miR-30d-5p, and miR-30e-5p. Subsequent study demonstrated that Curcumin could upregulate the level of miR-30c-5p in NSCLC cells, but had no significant effect on the levels of miR-30a-5p, miR-30b-5p, miR-30d-5p, and miR-30e-5p (Figure 4(a)). Furthermore, miR-30c-5p mimic could obviously reduce the expression of MTA1 (Figure 4(b)).
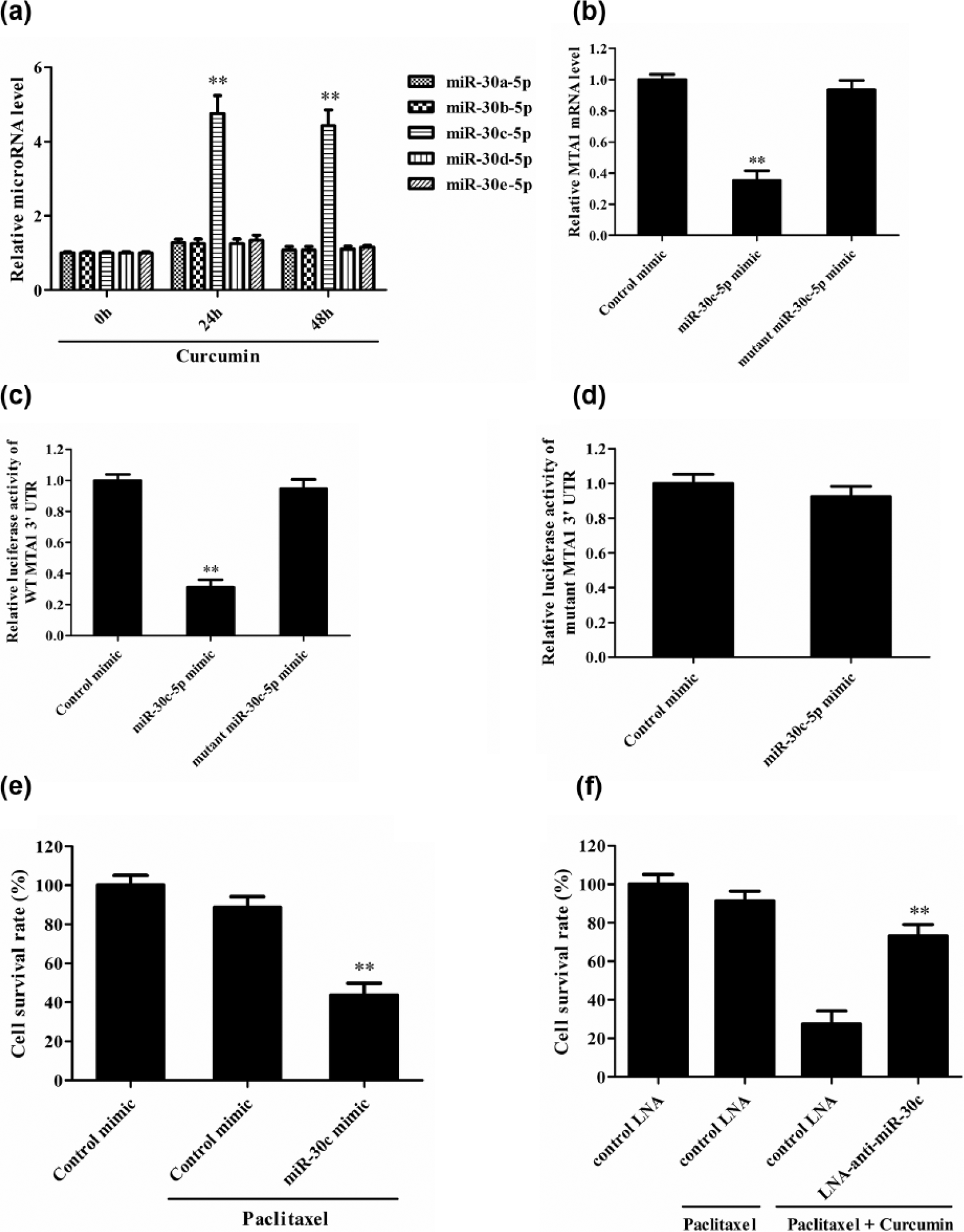
Curcumin reduces the expression of MTA1 gene through upregulation of miR-30c-5p in Paclitaxel-resistant A549 cells. (a) The expression of mircoRNAs was detected in Paclitaxel-resistant A549 cells at 24 and 48 h after treatment (**p < 0.01 vs 0 h group). (b) miR-30c-5p mimic, control mimic, and mutant miR-30c-5p mimic were transfected into the Paclitaxel-resistant A549 cells for 48 h, and then, the expression of MTA1 mRNA was detected by real-time PCR (**p < 0.01 vs control mimic group). (c) pGL3-Promoter/WT-MTA1 and miRNA mimic were co-transfected in HEK293T cells, and then, the luciferase activity was assayed at 48 h after transfection (**p < 0.01 vs control mimic group). (d) pGL3-promoter/mutant-MTA1 and miRNA mimic were co-transfected in HEK293T cells, and then, the luciferase activity was assayed at 48 h after transfection. (e) miR-30c-5p mimic and control mimic were transfected into the Paclitaxel-resistant A549 cells, and then, MTT was used to detect the cell survival at 48 h after transfection (p < 0.01 vs control mimic + Paclitaxel group). (f) miR-30c-5p inhibitor was transfected into the Paclitaxel-resistant NSCLC cells for 24 h, and then, the cells were cultured with the combination of 30 µM Paclitaxel and 30 µM Curcumin for 48 h (**p < 0.01 vs control LAN + Paclitaxel + Curcumin group). MTT was used to detect the cell survival. The data are from one experiment, representative of three independent experiments. Results were represented as means ± SD (n = 4 for each group).
In order to confirm the regulatory effect miR-30c-5p on MTA1 gene, the luciferase reporter plasmids of WT and mutant 3′-UTR of MTA1 mRNA were used. Transfection of miR-30c-5p mimic led to a significant suppression of luciferase activity of pGL3-Promoter/WT MTA1 reporter, but transfection of mutant miR-30c-5p mimic had no significant effect (Figure 4(c)). As for pGL3-Promoter/mutant MTA1 reporter, transfection of miR-30c-5p mimic had no significant effect on its luciferase activity (Figure 4(d)). These data indicate that miR-30c-5p does degrade MTA1 mRNA through targeting at 3′-UTR of MTA1 mRNA.
Further functional study showed that miR-30c-5p mimic could increase the sensitivity of Paclitaxel-resistant A549 cells to Paclitaxel (Figure 4(e)), while miR-30c-5p inhibitor could reduce Curcumin-enhanced sensitivity of Paclitaxel-resistant A549 cells to Paclitaxel (Figure 4(f)). Subsequent experiment was done to study the effect of Curcumin-microRNA-30c-MTA1 on the sensitivity of Paclitaxel-resistant 95D NSCLC cells to Paclitaxel, and similar results were obtained (Figure 5). Taken together, these data indicate that Curcumin increases the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel through microRNA-30c-mediated MTA1 reduction.

Curcumin increases the sensitivity of Paclitaxel-resistant H460 cells to Paclitaxel through microRNA-30c-mediated MTA1 reduction. (a) The expression of MTA1 was detected by western blot in Paclitaxel-resistant H460 cells treated with Paclitaxel or Curcumin or together (**p < 0.01 vs control group). (b) MTT was used to determine the cell survival of Paclitaxel-resistant H460 cells treated with Paclitaxel or Curcumin or together (**p < 0.01 vs control group; ##p < 0.01 vs Curcumin group). (c) pcDNA3.1/MTA1 or pcDNA3.1 and LAN-anti-miR-30c-5p or control LAN were transfected into the Paclitaxel-resistant H460 cells for 24 h, and then, the cells were cultured with the combination of 30 µM Paclitaxel and 30 µM Curcumin for 48 h. Finally, MTT assay was done to detect the cell survival (**p < 0.01 vs pcDNA3.1 + control LAN + Paclitaxel + Curcumin group). The data are from one experiment, representative of three independent experiments. Results were represented as means ± SD (n = 4 for each group).
Discussion
The resistance of NSCLC cells to Paclitaxel is a major problem in clinic during Paclitaxel-based combinational chemotherapies. Notably, Curcumin has been reported to suppress the proliferation and invasion of NSCLC cells.30,31 Our present studies revealed that Curcumin treatment alone had only slight cytotoxicity to Paclitaxel-resistant NSCLC cells; however, the combinational use of Curcumin and Paclitaxel had obvious cytotoxicity to NSCLC cells, indicating that Curcumin treatment does increase the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel. This finding might provide an insight into the combinational use of Curcumin and Paclitaxel for treating Paclitaxel-resistant NSCLC in clinic.
Further mechanical studies revealed that Curcumin reduced the expression of MTA1 in Paclitaxel-resistant NSCLC cells. MTA1 gene knockdown could imitate the effect of Curcumin on Paclitaxel-resistant NSCLC cells. However, overexpression of MTA1 reduced the effect of Curcumin on Paclitaxel-resistant NSCLC cells. Collectively, these data indicate that the downregulation of MTA1 is involved in Curcumin-enhanced sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel. Further studies need to be performed to find the potent mechanism by which Curcumin induces the downregulation of MTA expression.
As post-transcriptional regulation molecules, microRNAs have been identified as an abundant class of small non-coding RNAs that play important roles in various biological processes through negative regulation of target genes.32–36 Reportedly, Curcumin is able to upregulate the levels of microRNAs such as miR-192-5p, miR-365, and miR-21.37–39 Therefore, the potent upstream regulation of MTA1 by microRNAs was subsequently explored in the current studies. As a result, five potential miRNAs including miR-30a-5p, miR-30b-5p, miR-30c-5p, miR-30d-5p, and miR-30e-5p were predicted to target at human MTA1 3′-UTR through TargetScanHuman. Further study demonstrated that Curcumin could induce the upregulation of miR-30c-5p rather than other four microRNAs in NSCLC cells. More importantly, miR-30c-5p mimic could markedly reduce the expression of MTA1 and increases the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel, while miR-30c-5p inhibitor could decrease Curcumin-enhanced sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel. Luciferase assay demonstrated that Curcumin-upregulated miR-30c-5p targeted at 3′-UTR of MTA1 mRNA directly and degraded MTA1 mRNA. Taken together, these data indicate that Curcumin reduces the expression of MTA1 gene through upregulation of miR-30c-5p in Paclitaxel-resistant NSCLC cells. Since Curcumin has been reported to regulate signaling molecules and transcriptional factors,15–17 further studies can be done to explore which signaling molecule and/or transcriptional factor is required for Curcumin-increased miR-30c-5p expression in Paclitaxel-resistant NSCLC cells.
In summary, the effect of Curcumin on the Paclitaxel-resistant NSCLC cells was investigated, and study revealed that Curcumin treatment could markedly increase the sensitivity of Paclitaxel-resistant NSCLC cells to Paclitaxel. Mechanically, Curcumin reduced the expression of MTA1 gene through upregulation of miR-30c in Paclitaxel-resistant NSCLC cells. Furthermore, MTA1 reduction was able to sensitize Paclitaxel-resistant NSCLC cells and enhance the effect of Paclitaxel to the cells. Our studies indicate that Curcumin might be a potential adjuvant for NSCLC patients during Paclitaxel treatment.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by a Grant from Department of Public Health of Jiangsu Province (YG201403), Science and Technology Developmental Grant from Suzhou City of Jiangsu Province (SYSD2013162), Social Developmental Grant from Kunshan City of Jiangsu Province (KSZ1309), and Suzhou city diagnosis and treatment special project for clinical key diseases (LCZX201518).
