Abstract
Glioma is the most common primary malignant tumor of the central nervous system, which results in both a poor prognosis and outcome because of the aggressive progression of disease, growth and resistance to surgery, chemotherapy, and radiotherapy. MiR-140-5p is a small, non-coding single-stranded RNA molecule, which was previously studied in the settings of human tongue cancer, hepatocellular carcinoma, and colorectal cancer. However, detailed data that formally demonstrate the contribution of miR-140-5p to glioma development are missing. Similarly, relatively little is known about the relationship of miR-140-5p, vascular endothelial growth factor A, and matrix metalloproteinase-2 in glioma progression. In this study, we found that miR-140-5p expression was significantly decreased in glioma tissues and in the glioma cell-lines U87 and U251 as compared with non-cancerous brain tissues by quantitative real-time polymerase chain reaction. In addition, miR-140-5p inhibited glioma cell proliferation and invasion and promoted glioma cell apoptosis both in vivo and in vitro. Interestingly, while the expression levels of miR-140-5p were higher in glioma cells, the messenger RNA or protein expression levels of vascular endothelial growth factor A and matrix metalloproteinase-2 were lower in glioma cells as determined by quantitative real-time polymerase chain reaction, western blot assay, and immunohistochemistry. By contrast, downregulation in the expression levels of miR-140-5p augmented the messenger RNA and protein expression levels of both vascular endothelial growth factor A and matrix metalloproteinase-2. These findings suggested that miR-140-5p inhibited glioma proliferation and invasion by regulating the vascular endothelial growth factor A/matrix metalloproteinase-2 signaling pathway both in vitro and in vivo.
Keywords
Introduction
Glioma is the most common primary malignant tumor of the central nervous system (CNS) that displays a series of malignant features including widespread, rapid, and aggressive progression and growth and resistance to surgery, chemotherapy, and radiotherapy, all of which result in poor prognosis and outcomes in glioma patients previously treated with both traditional and newer therapeutic approaches. Newly diagnosed glioma patients typically have a median survival time of approximately only 14 months. 1 In addition, there is evidence of inter-individual variability in glioma patients. Thus, new therapeutic strategies and targets are urgently needed to consistently manage patients who presented with this troubling condition.
MicroRNAs (miRNAs) are a class of small, non-coding, and single-stranded RNA molecules with an approximate length of 19–25 nucleotides, 2 which play important roles in a variety of biological processes by regulating gene expression though binding to the 3′-untranslated region (3′-UTR) of target messenger RNAs (mRNAs). This process results in translational repression or degradation of target mRNAs. 3 Although most miRNAs have not been sufficiently studied, ample evidence informs us that miRNA exerts regulatory functions in different tumor types. This includes stimulating tumor growth or suppressing tumor development and regulation of cell cycle kinetics, differentiation, proliferation, invasion, and programmed cell death by apoptosis. 4 In addition, miRNA plays a vital role in the development and plasticity of the CNS, especially being closely associated with the emergence of CNS disease and progression of those diseases.5,6 An ever increasing number of studies have suggested the capacity of miRNAs to therapeutically target glioma in the clinic. For example, miR-16, miR-494-3p, and miR-184 regulate glioma cell proliferation, invasion, and apoptosis and do so by differential mechanisms that target oncogenes or anti-oncogenes and by mechanisms that have been demonstrated in a xenograft animal model.7–9
In this study, we found that miR-140-5p suppressed progression of human tongue cancer, hepatocellular carcinoma, and colorectal cancer10–12—processes that have not previously been investigated extensively. Specifically, there is an absence of previously published data that have demonstrated the contributions of miR-140-5p in the development of glioma. Furthermore, the molecular mechanism of miR-140-5p in human brain glioma is also very poorly understood.
Since glioma formation is partly dependent on tumor characteristics like angiogenesis, invasion, and migration, vascular endothelial growth factor (VEGF) is known to play a crucial role in tumorigenesis. Accordingly, previously published studies in nude mouse models have shown that suppressed expression of VEGF in tumor cells by antisense RNA technology blocked the growth of glioma cells. 13 Moreover, VEGF plays a central role in glioma progression, especially in the context of glioma stem cells.14–16 VEGFA is an important member of the VEGF family and includes VEGFA, VEGFB, VEGFC, VEGFD, and placental growth factor (PIGF).
VEGFA is a key regulator of developmental neoangiogenesis, which mainly interacts with the VEGF receptors FLT1 and KDR, thereby modulating downstream signaling pathways. 17 Activation of downstream signaling pathways promotes endothelial cell proliferation, angiogenesis, and glioma cell growth. 16 By contrast, matrix metalloproteinases (MMPs) are a family of zinc-dependent enzymes that are secreted in an inactive precursor form. The proenzyme form of MMP is activated autocatalytically or by other contributing proteinases. 18
The MMP family currently comprises 25 related and distinct vertebrate gene products, of which 24 products are found in mammals. 19 In addition, MMP2 degrades cleaved collagen more efficiently than do other gelatinolytic MMPs, with the notable exception of MMP9. 20 Indeed, one study demonstrated that MMP2 promoted the formation of tumor vessels and accelerated glioma invasion and metastasis. 21 Thus, it is formally possible that inhibiting malignant glioma invasion and metastasis may contribute to a more positive prognostic outcome.
It was previously found that MMP2 was regulated by VEGFA in the mitogen-activated protein kinase (MAPK)-dependent signaling pathway. 22 By contrast, VEGFA was predicted as a target gene of miR-140-5p by relative forecast software such as TargetScan, miRanda, and miRWalk.
In this study, we explored how miR-140-5p played an effective role in glioma development, proliferation, invasion, and tumor cell migration.
Materials and methods
Human tissue samples
Human brain glioma tissue specimens were obtained from January 2000 through December 2013 at the Department of Neurosurgery of the First Affiliated Hospital of Soochow University, People’s Republic of China. These samples were obtained from 58 glioma patients, including 21 cases of World Health Organization (WHO) low-grade I and II tumors, 37 cases of WHO high-grade III and IV, and seven patients presented with craniocerebral injuries whose brain tissues were partially resected to reduce intracranial pressure according to decompression treatment guidelines.
In this study, there were 38 male patients with a mean age of 49.2 years at the time of surgery, including 5 cases of brain trauma, 13 cases of diffuse astrocytoma (II), 7 cases of anaplastic astrocytoma (III), and 13 cases of primary brain glioblastoma. In addition, this study included 27 women with a mean age of 47.3 years at the time of surgery including 2 cases of brain trauma, 8 cases of diffuse astrocytoma (II), 10 cases of anaplastic astrocytoma (III), and 7 cases of primary brain glioblastoma. 18 The tissue samples that were obtained from glioma and brain trauma patient were immediately placed into liquid nitrogen after resection from those patients. Written and informed consent to use specimens for future research was obtained from each patient. Additionally, this study was authorized by the local ethics committee of our hospital.
Cell cultures and cell transfection
The glioma cell-lines U87 and U251 were purchased from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). All cell lines were maintained in Dulbecco’s modified Eagle’s medium (DMEM; Hyclone, Thermo Fisher Scientific, Waltham, MA, USA) that was supplemented with 100 U/mL of penicillin, 100 µg/mL of streptomycin, and 10% fetal bovine serum (FBS; Gibco, Invitrogen, Carlsbad, CA, USA) at 37°C under a fully humidified atmosphere of 5% CO2 in air. The glioma cell-lines U87 and U251 were transfected with miR-140-5p mimics, the miR-140-5p inhibitor, and negative control oligonucleotides by Lipofectamine 2000 (Invitrogen Inc., Carlsbad, CA, USA). Specific sequences were as follows—miR-140-5p mimics had the following sequences: 5′-ACCAUAGGGUAAAACCACUGUU-3′; miR-140-5p inhibitor: 5′-CUACCAUAGGGUAAAACCACUG-3′; and the negative control oligonucleotide had the following sequences: 5′-ACGUGACACGUUCGGAGAATT-3′. The glioma cell-lines U87 and U251 were transfected with miR-140-5p mimics, and these negative oligonucleotides were washed with phosphate-buffered saline (PBS), and then, cells were harvested and transfection efficiency was determined by flow cytometry (Cytomics FC 500; Beckman Coulter, Brea, CA, USA), and the results were analyzed with CXP Cytometer and CXP Analysis.
Quantitative real-time polymerase chain reaction analysis of mRNA and miRNA expression
TRIzol reagent (Invitrogen Inc., Carlsbad, CA, USA) was applied to extract the total RNA of the tissue specimens and cell lines that were transfected or left non-transfected for the following study. The mRNA expression levels of miR-140-5p, VEGFA, and MMP2 were analyzed by an All-in-One miRNA quantitative real-time polymerase chain reaction (qRT-PCR) detection kit (GeneCopoeia, Rockville, MD, USA) and a SYBR Green I qRT-PCR kit (LightCycler 480; Roche Applied Science, Basel, Switzerland) according to the manufacturer’s instructions. The expression level of glyceraldehyde 3-phosphate dehydrogenase (GAPDH) served as an internal control for mRNA expression, and the endogenous U6 small nuclear RNA (snRNA) served as an internal control for miRNAs. The following primers were used: VEGFA forward primer: 5′-GTGCCCGCTGCTGTCTAATG-3′; VEGFA reverse primer: 5′-TAACTCAAGCTGCCTCGCCT-3′; MMP2 forward primer: 5′-AGGATGGCAAGTACGGCTTC-3′; MMP2 reverse primer: 5′-CTTCTTGTCGCGGTCGTAGT-3′; GAPDH forward primer: 5′-AGAAGGCTGGGGCTCATTTG-3′; and GAPDH reverse primer: 5′-AGGGGCCTCCACAGTCTTC-3′. The relative expression of each gene was calculated and normalized by the 2−ΔΔCT method. Each sample was tested in triplicate.
Analysis of cell proliferation, cell cycle, and cell apoptosis
Cells (at a density of 2 × 105/100 µL) were seeded into a 96-well plate after being transfected with miR-140-5p mimics or the negative control oligonucleotides for 12 h. Then, the cell counting kit-8 (CCK-8; Beyotime, Shanghai, China) reagent was added to each well at 24, 48, and 72 h post transfection, following which the plates were incubated for 4 h at 37°C. Absorbance values were measured at a wavelength of 450 nm, and the proliferation rates of each group were analyzed.
To analyze the cell cycle, transfected cells were fixed in 75% ethanol at 4°C, and then, cells were collected and washed with ice-cold PBS and resuspended in 100 µL binding buffer containing 5 µL of 7-AAD (7-amino-actinomycin D) and 5 µL of PI at room temperature in the dark for 10–15 min. Then, cell cycle was analyzed by flow cytometric assay (Cytomics FC 500) with CXP Cytometer, CXP Analysis, and Multicycle AV for Windows. All experiments were performed in triplicate. The proportion of glioma cells undergoing apoptosis that had been transduced with miR-140-5p mimics or negative control oligonucleotides was detected by the Annexin V Apoptosis Detection Kit PE (eBioscience, San Diego, CA, USA). Cells transfected with miR-140-5p mimics or negative oligonucleotides were collected and resuspended in 400 µL 1× binding buffer containing 5 µL 7-AAD (7-amino-actinomycin D) and 5 µL PI at room temperature in the dark for 10–15 min, and then, cells were analyzed by flow cytometric assay (Cytomics FC 500) with CXP Cytometer and CXP Analysis.
Analysis of the invasion assay and scratch wound assay
Matrigel (BD Biosciences, San Jose, CA, USA) was used to cover transwell chambers (40 µL per well) at 37°C for 2 h after being diluted with serum-free medium. In this assay, 5 × 104 cells that had been suspended in 100 µL serum-free medium were added to the upper chambers of each well, while the lower chambers were filled with 750 µL DMEM that was supplemented with 10% FBS, followed by incubation at 37°C under a humidified atmosphere of 5% CO2 in air for 48 h. Then, non-invading cells from the top chambers were removed by a cotton wrap, and cells on the lower membrane surface were fixed in 4% formaldehyde for 30 min and stained with 0.2% crystal violet for 5 min. Under a microscope, the migrating cells that had penetrated the matrigel were imaged and counted across six randomly chosen fields. All experiments were performed in triplicate. For scratch wound assay, cells were seeded in six-well plates and cultured to 70%–80% confluence. A 200 µL pipette tip was applied to generate a linear wound. The floating cells were rinsed with PBS. Cells cultured for 48 h and were recorded under a microscope (Olympus, Tokyo, Japan) at 0, 6, 12, 24 and 48 h. All experiments were performed in triplicate.
Western blot analysis
Cells were transduced for 48 h with miR-140-5p mimics, the miR-140-5p inhibitor, and negative control oligonucleotides. Next, total protein was extracted from the cells and quantified by radioimmunoprecipitation assay (RIPA) and a bicinchoninic acid (BCA) assay kit (Beyotime, Shanghai, China). Equal protein samples were separated through a 12% sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) gel and then transferred onto nitrocellulose membranes. These membranes were incubated with primary antibodies overnight at 37°C, and then, membranes were washed with phosphate-buffered saline with Tween 20 (PBST) before incubating them with horseradish peroxidase (HRP)-conjugated anti-rabbit secondary antibodies (1:5000; Prosci Inc., Poway, CA, USA) or HRP-conjugated anti-mouse secondary antibodies (1:5000; Prosci Inc., Poway, CA, USA) for 2 h at room temperature. Finally, enhanced chemiluminescence (ECL) western blotting detection reagents were used to detect and record the detected proteins on the immune cross-reacted membranes. The primary antibodies used were rabbit anti-VEGFA (Abcam, Tokyo, Japan), mouse anti-VEGFA (Abcam), and rabbit anti-GAPDH (Santa Cruz Biotechnology, Santa Cruz, CA, USA), all at a dilution of 1:1000.
Xenograft experiments and analysis
Cell lines were stably transfected with pLenti-miR-140-5p-GFP and pLenti-NC-GFP (Shanghai GenePharma Co, Ltd, China) and were diluted with proper cell culture medium. For formation of intracranial xenografts, 1 × 105 cells in 3 µL DMEM/F12 were injected 3 mm deep into the frontal lobe, 1.8–2.0 mm right of sagittal suture and 1.8–2 mm anterior to coronal suture, of 4–5 week old BALB/c female nude mice (n = 6 per group, Shanghai SLAC Laboratory Animal Co., Ltd., Shanghai, China) using a stereotactic frame. Mice were observed daily, weighted two times a week until moribund signs (hunchback posture and/or 20% weight loss) appeared, and mice were killed. An amount of 1 × 106 cells was injected subcutaneously into the armpit subcutaneous area of the left forelimb (n = 6 per group). Caliper measurements were performed to assess subcutaneous tumor growth during mice development. The tumors were obtained from nude mice after about 1 month and then stored by fixing in formalin for subsequent research.
Immunohistochemistry analysis
U87-glioma-cell-transplanted tumor specimens were paraffin-embedded and precision sliced into 6 µm sections with a microtome. Antigen retrieval was performed in 10 mM sodium citrate buffer at pH 6 for 10 min at 37°C, and then, slides were incubated with primary antibodies against VEGFA and MMP2 at 4°C overnight. On the following day, sections were incubated with the HRP-DAB Cell and Tissue Staining Kit (R&D Systems, Minneapolis, MN, USA). Results between positive and negative control groups were analyzed and photographed by light microscopy.
Statistical analysis
Statistical analyses were carried with the SPSS version 13.0 statistical software program (SPSS, Chicago, IL, USA). All results were expressed as the mean ± standard deviation (SD). The data were analyzed by two-tailed Student’s t-test and analysis of variance (ANOVA). Data were considered statistically significant at an alpha value of p < 0.05.
Results
MiR-140-5p and VEGFA were negatively correlated in glioma
Previous research had adequately demonstrated that miR-140-5p played vital roles in partial tumor progression, but not in the setting of glioma. In addition, the functional relationship of miR-140-5p and VEGFA in glioma was not previously reported; qRT-PCR was carried out to detect their expression levels. In 65 tissue samples, miR-140-5p expression was markedly decreased in glioma tissues as compared to non-cancerous brain tissues. Obviously, the expression levels of miR-140-5p decreased gradually with the degree of tumor malignancy (Figure 1(a)). By contrast, VEGFA had higher expression levels in glioma specimens than did non-tumor brain tissues and the expression of VEGFA increased with the degree of glioma malignancy (Figure 1(b)). In the glioma cell-lines U87 and U251, miR-140-5p displayed lower expression levels than non-cancerous brain tissues, although the expression of VEGFA was significantly upregulated as compared to non-cancerous brain tissues (Figure 1(c) and (d)). These results suggested that miR-140-5p might be negatively associated with VEGFA in the setting of glioma progression.

Expression of miR-140-5p and VEGFA in glioma tissues and cells. (a and b) The expression of miR-140-5p and VEGFA in different degree glioma tissues and non-cancerous brain tissues. (c and d) The expression of miR-140-5p and VEGFA in non-cancerous brain tissues and glioma cell-lines U87 and U251.
Up-regulation of miR-140-5p suppressed cell proliferation and cell cycle
To comprehend the function of miR-140-5p in glioma development, the glioma cell-lines U87 and U251 were transfected with miR-140-5p mimics and negative control oligonucleotides, and the transfection efficiency was detected by flow cytometry and fluorescence microscopy (Figure 2(a) and (b)). Transfection efficiency of glioma cell-lines U87 and U251 transfected with miR-140-5p mimics and negative oligonucleotides was 93.6%, 81.9%, 88.2%, and 81.5%; qRT-PCR showed that the blank and negative groups had lower expression of miR-140-5p than miR-140-5p mimic groups in U87 and U251 (Figure 2(c)). We also explored the role of miR-140-5p in cell proliferation by the CCK-8 assay. MiR-140-5p overexpression inhibited glioma cell-line proliferation as compared with their empty vector or untreated cells (Figure 3). Meanwhile, enhanced expression of miR-140-5p induced U87 and U251 cell cycle arrest, resulting in a higher frequency of cells (65.308% and 65.322%) in the G0/G1 phase and fewer cells (23.672% and 20.703%) in the S phase of the cell cycle as compared to the groups that had been transfected with negative oligonucleotides and blank groups without treatment (Figure 4). Thus, miR-140-5p possibly served as a tumor growth inhibitor in the development of glioma.

The transfection efficiency, fluorescence image, and qRT-PCR of the glioma cell-lines U87 and U251 were transfected with miR-140-5p mimetics and negative control oligonucleotides. (a) The transfection efficiency of U87 and U251 transfected with miR-140-5p mimics and negative oligonucleotides. (b) Fluorescence images of U87 and U251 transfected with miR-140-5p mimics and negative oligonucleotides. (c and d) The expression level of miR-140-5p in U87 and U251 transfected with miR-140-5p mimics, blank, and negative groups with GraphPad Prism 5 analysis, and error bars correspond to the SD of triplicate measurements.

The cell proliferation of U87 and U251. (a) The cell proliferation rate in U87 tansfected with miR-140-5p mimics, negative oligonucleotides, and the untreated cells group. (b) The cell proliferation rate in U251 tansfected with miR-140-5p mimics, negative oligonucleotides, and the untreated cells group, and error bars correspond to the SD of triplicate measurements.
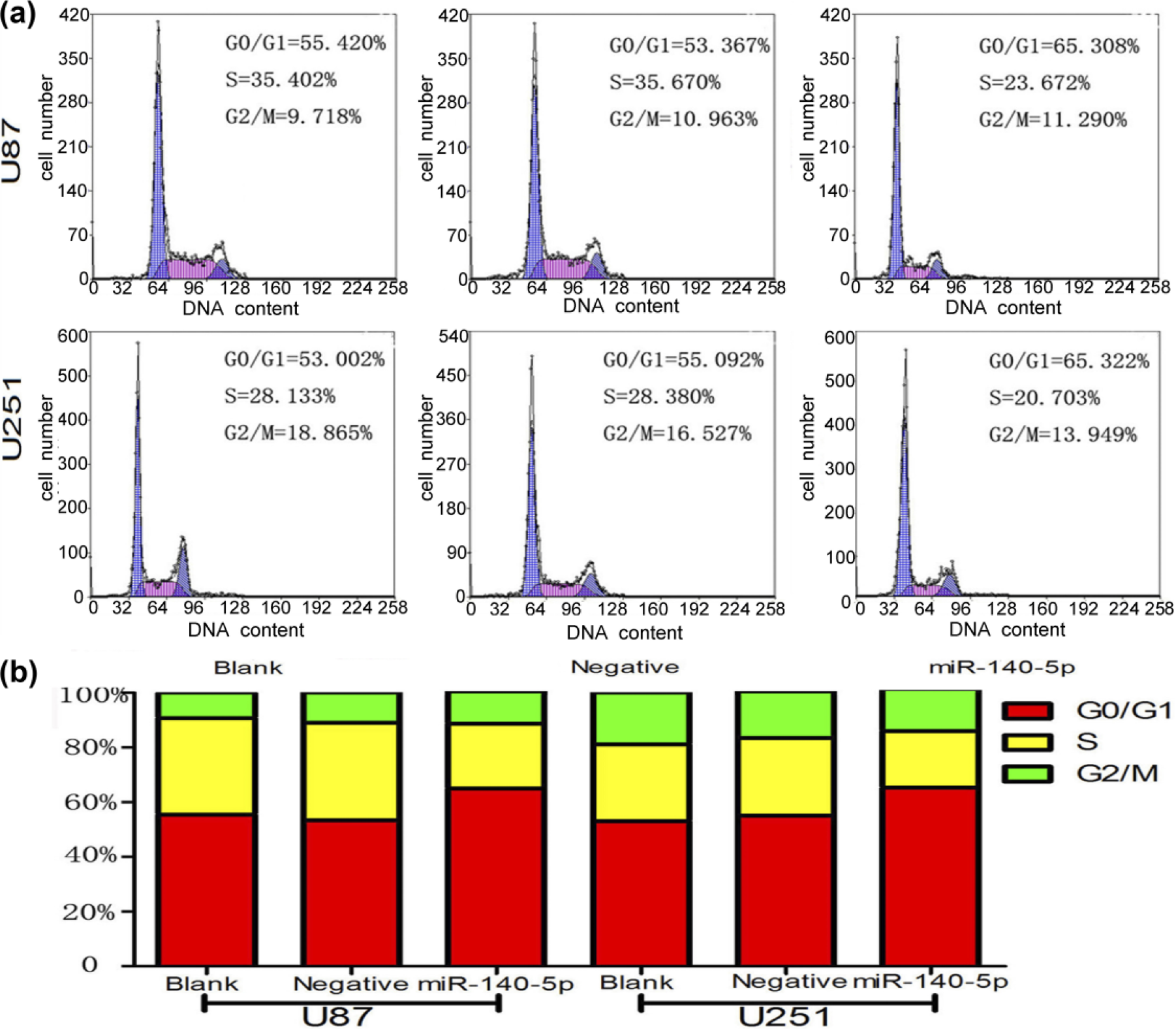
Cell cycle distribution of U87 and U251 tansfected with miR-140-5p mimics, negative oligonucleotides, and blank groups. (a) Flow cytometry results of each phase cell rate in U87 and U251 tansfected with miR-140-5p negative oligonucleotides and blank groups. (b) Column chart of each phase cell rate of U87 and U251 tansfected with miR-140-5p negative oligonucleotides and blank groups.
Overexpression of miR-140-5p promoted cell apoptosis and inhibited cell invasion and migration
There were obvious differences in apoptosis among cells that were transfected with miR-140-5p mimetics and negative and blank controls. According to the results of flow cytometry, the apoptosis rates of the blank, negative, and miR-140-5p mimic groups were 7.6%, 3.3%, and 26.9%, respectively, in U87, whereas 2.3%, 1.2%, and 31.5% in U251 (Figure 5(a)). Thus, we suggested that upregulation of miR-140-5p in the glioma cell-lines U87 and U251 had higher apoptosis rates as compared with empty vector or untreated cells, which suggested that apoptosis was obviously influenced by miR-140-5p expression. To further explore the anti-oncogenic function of miR-140-5p in vitro, the invasion and migration capacity of glioma cells was evaluated by using a transwell invasion system. As shown below, overexpression of miR-140-5p clearly inhibited cell invasion and migration (Figure 6(a) and (b)). In addition, the number of invasive cells with miR-140-5p mimics was significantly reduced as compared to empty vector and untreated cells.
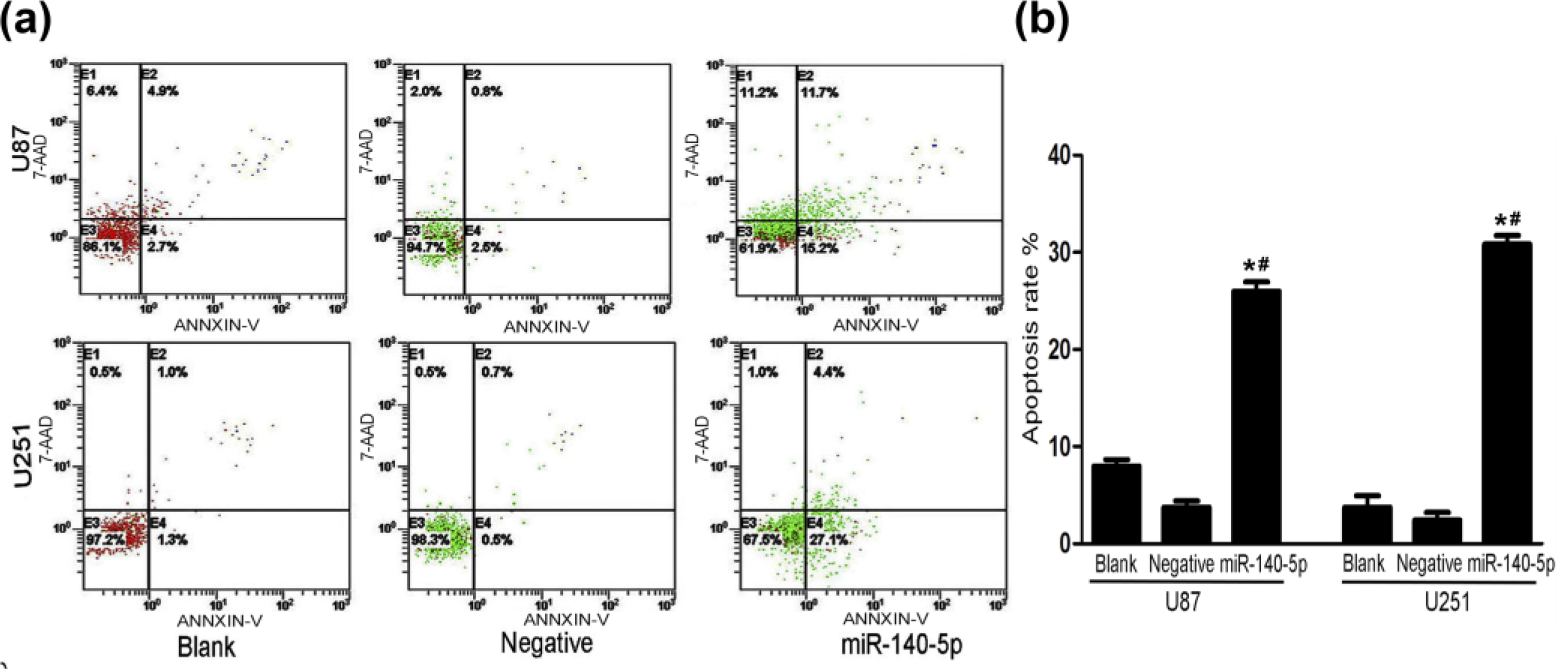
The apoptosis rates of U87 and U251 tansfected with miR-140-5p mimics, negative oligonucleotides, and blank groups. (a) Flow cytometry results of the apoptosis rate in U87 and U251 tansfected with miR-140-5p mimics, negative oligonucleotides, and blank groups. (b) Column chart of the apoptosis rate in U87 and U251 tansfected with miR-140-5p mimics, negative oligonucleotides, and blank groups with GraphPad Prism 5 analysis, and error bars correspond to the SD of triplicate measurements.

MiR-140-5p mimics inhibited cell invasion and migration of U87 and U251. (a) Transwell assay exhibits the invasion of U87 and U251 cells transfected with miR-140-5p mimics, negative oligonucleotides, and blank groups. (b) Column chart of the invasion of U87 and U251 cells tansfected with miR-140-5p mimics, negative oligonucleotides, and blank groups with GraphPad Prism 5 analysis, and error bars correspond to the SD of triplicate measurements (*compared to the blank group, p < 0.01; #compared to the negative group, p < 0.01). (c) Scratch wound assay results of U87 and U251 transfected with miR-140-5p mimics, negative oligonucleotides, and blank groups.
MiR-140-5p regulated VEGFA and MMP2 levels in U87 and U251
To study the molecular mechanisms of miR-140-5p, we explored publicly available databases (e.g. miRbase, miRanda, and TargetScan) to search potential target genes (Figure 7(a)). VEGFA, as one potential target gene in the biological information software, had been checked as a direct target gene of miR-140-5p in colorectal cancer using the luciferase reporter assay. 12 A series of experiments were applied to determine the relationship between miR-140-5p and VEGFA in glioma; qRT-PCR analysis and western blot analysis were used to detect glioma cells U87 and U251 that were transfected with miR-140-5p mimics, the inhibitor, and negative oligonucleotides. The results showed that overexpression of miR-140-5p reduced VEGFA protein expression as compared to empty vector or untreated cells. By contrast, low expression of miR-140-5p increased VEGFA protein expression as compared to empty vector or untreated cells (Figure 7(b)).

MiR-140-5p regulated the protein levels of VEGFA and MMP2 in glioma cell-lines. (a) The binding sites between miR-140-5p and VEGFA were predicted with different gene databases. (b) Western blotting analysis of MMP2 and VEGFA expression in U87 and U251 cells. (c)–(f) Relative gray value of VEGFA and MMP2 expression in U87 and U251 with ImageJ analysis.
In addition, the blank and negative groups had higher RNA expression of VEGFA than did the miR-140-5p mimic groups and lower RNA expression level of VEGFA than miR-140-5p inhibitor groups in U87 and U251 (Figure 8(a) and (b)). MMP2 was a downstream gene of VEGFA, and they were positively correlated in the glioma cell-line A172. 22 In U87 and U251 cells that were transfected with miR-140-5p mimics, miR-140-5p inhibitor, and negative oligonucleotides, RNA expression of MMP2 was also examined by qRT-PCR. We found that overexpression of miR-140-5p in U87 and U251 cells significantly reduced MMP2 levels and dampened miR-140-5p served to increase the RNA expression of MMP2 (Figure 8(c) and (d)). In conclusion, we demonstrated that miR-140-5p regulated the expression of VEGFA and MMP2.
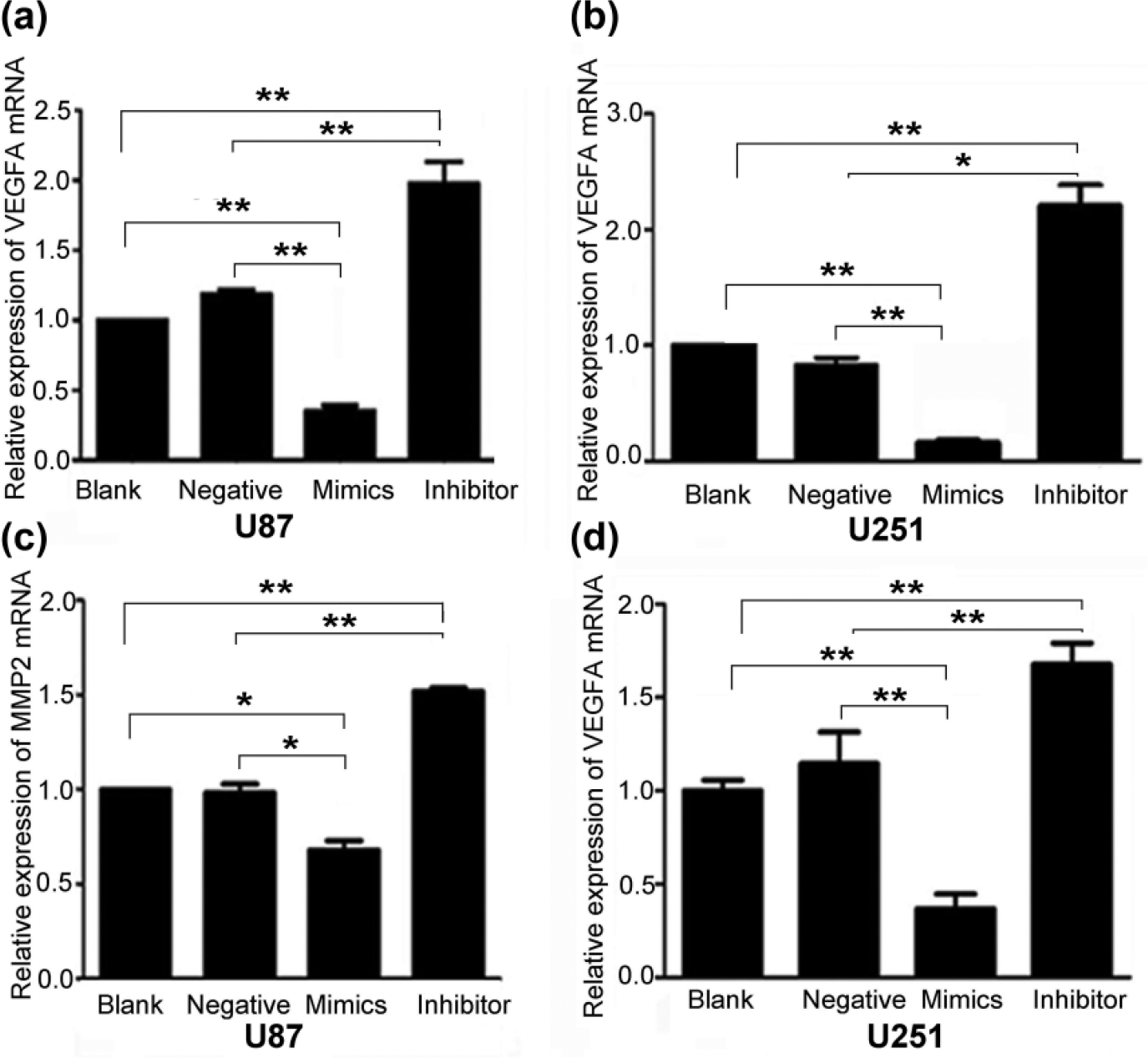
MiR-140-5p regulated the RNA levels of VEGFA and MMP2 in U87 and U251. (a and b) The expression level of VEGFA in U87 and U251. (c and d) The expression level of MMP2 in U87 and U251 (*p < 0.05; **p < 0.01).
miR-140-5p suppressed glioma cell proliferation and invasion in vivo
A xenograft model helped to determine whether miR-140-5p was involved in glioma tumorigenesis and invasiveness. The tumors that were resected from the armpit subcutaneous area of the left forelimb of nude mice showed clear differences in size and weight by visual inspection. The subcutaneous tumor mass from mice injected with U87 cells transduced with pLenti-miR-140-5p-GFP was significantly smaller and lighter than tumors that were found in mice that had been adoptively transferred with U87 cells transduced with pLenti-NC-GFP (Figure 9(a) and (b)). With nude mouse development, the volume of subcutaneous tumors in the subcutaneous area was smaller in those groups showing increased miR-140-5p than in xenograft tumors of negative control groups (Figure 9(c) and (d)). Furthermore, U87 tansfected with pLenti-miR-140-5p-GFP and pLenti-NC-GFP was injected into nude brain to form the intracranial xenograft. Hematoxylin and eosin staining showed that the tumor of the mice injected with U87 tansfected with pLenti-miR-140-5p-GFP has lower malignancy (Figure 9(e)). Immunohistochemistry showed that the expression of VEGFA and MMP2 was lower than the negative group (Figure 9(f)).
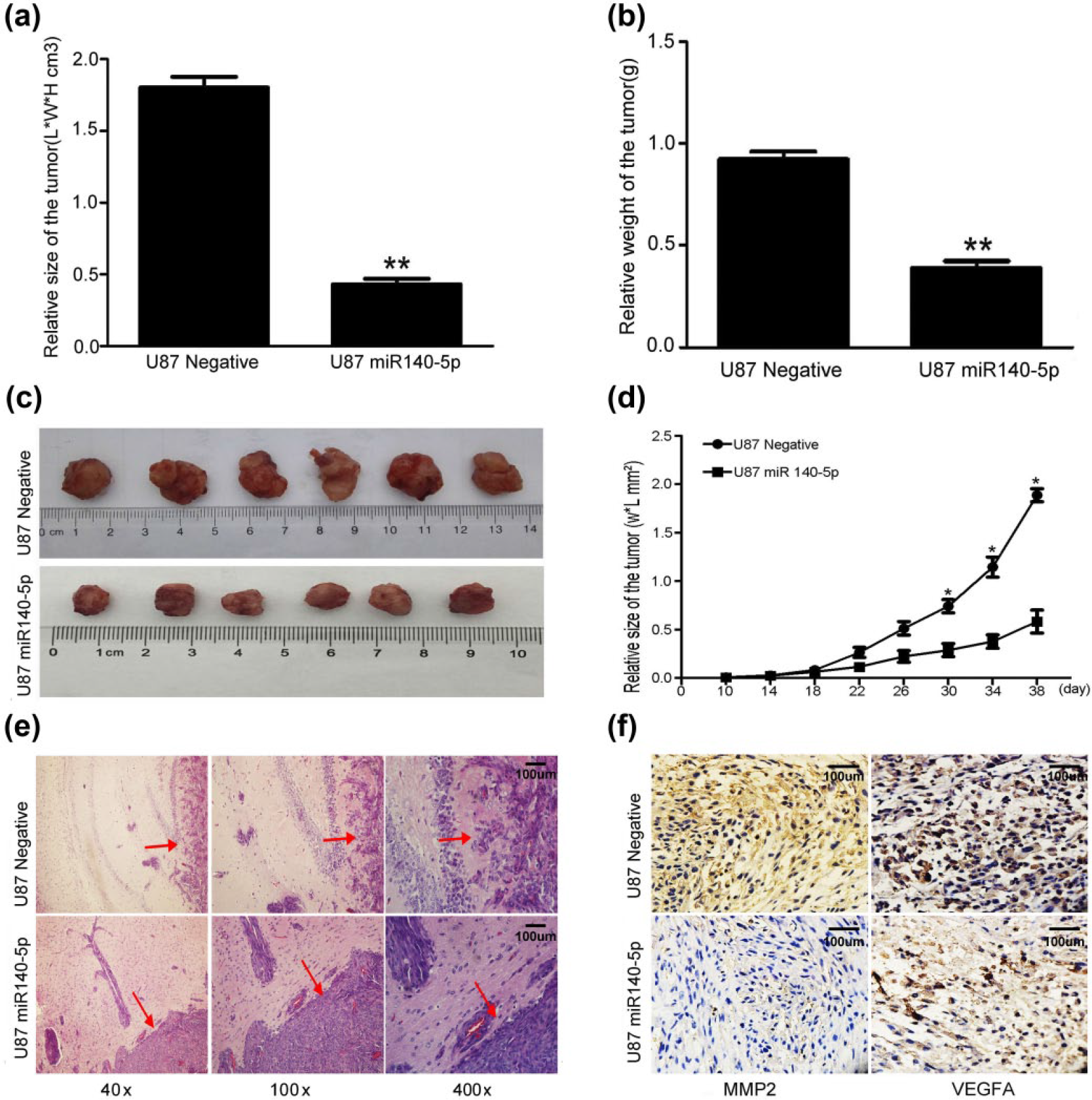
MiR-140-5p inhibited tumor proliferation and invasion in vivo. (a)–(c) The volume and weight of subcutaneous tumors of the mice in both groups (**p < 0.01). (d) The size of subcutaneous tumors of mice in both groups in survival time (*p < 0.05). (e) Hematoxylin–eosin staining of the intracranial tumors of the mice in both groups. (f) Immunohistochemistry staining of VEGFA and MMP2 of the intracranial tumors of the mice in both groups (400×).
Discussion
MiRNA represents one of the most intensely studied research areas and has been shown to play various roles in different tumors and various biological processes. MiRNA also displays a variety of mechanisms that are involved in tumor progression including regulating mRNA stability or negatively blocking mRNA translation by binding to its target gene. 23 Moreover, miRNA behaves as an oncogene or an anti-oncogene in many tumor types; for example, miR-140-5p, as has been previously reported.10–12,24 However, miR-140-5p has not been sufficiently explored though experiment research in terms of its effects on cellular proliferation, angiogenesis, differentiation, and invasion in malignant glioma cells.
In our study, we suggested that miR-140-5p inhibited proliferation and invasion and promoted apoptosis in glioma development. Furthermore, miR-140-5p expression was inversely correlated with the degree of malignancy in glioma as determined by qRT-PCR. To clarify the molecular mechanisms of miR-140-5p, we paid close attention to VEGFA depending on the prediction software databases (e.g. TargetScan, miRanda, and miRBase). As we know, glioma is characterized by increased proliferation, robust angiogenesis, and invasion into the surrounding normal brain tissues, VEGFA may reveal itself as a potential therapeutic target in the clinical treatment and management of glioma. 25 There is abundant evidence to show that the anti-VEGF antibody bevacizumab has been widely used in cancer therapy and that the expression of VEGFA is influenced by bevacizumab in the chemotherapeutic treatment of metastatic colorectal cancer.26,27 In addition, VEGFA could also promote de novo endothelial cell formation to provide a vascular supply for the growth and development of osteosarcoma and glioblastoma, which suggested that VEGFA has an important role in tumor growth and metastasis. 28 The importance of the relationship between miR-140-5p and VEGFA is self-evident, and understanding the importance of this relationship is urgently needed to validate in studies of glioma both in vivo and in vitro. It can also provide a new therapeutic strategy for the treatment of glioblastoma.
VEGFA and MMP2 have the inherent capacity to enhance invasiveness and angiogenesis in glioblastoma.21,29,30 MMP2 is an important member of the MMP family, which was positively correlated with the extent of glioma and was not expressed or only weekly expressed in healthy brain tissues. 21 VEGFA and MMP2 play a synergistic role in tumor invasion of normal brain tissues in glioma. We found that VEGFA is an upstream gene of MMP2 in the extracellular signal–regulated protein kinase (ERK)/MAPK signaling pathway and could influence the expression of MMP2. 22 Moreover, the results showing that overexpression of miR-140-5p simultaneously reduced the expression levels of VEGFA and MMP2 suggested that miR-140-5p influenced glioma progression and did so by regulating the VEGFA/MMP2 signaling pathway. Moreover, miR-140-5p directly affected glioblastoma development in intracranial and subcutaneous implantation models by regulating the functional expression of VEGFA and MMP2.
In conclusion, our results have shown that VEGFA displayed higher expression levels in glioma specimens and glioma cell-lines than did non-tumorigenic brain tissues. Meanwhile, VEGFA was positively correlated with the degree of malignancy of glioma. Overexpression of miR-140-5p directly bound to the 3′-UTR region of VEGFA remarkably reduced the mRNA and protein expression of VEGFA, which inversely downregulated miR-140-5p and enhanced VEGFA mRNA and protein expression. Clearly, miR-140-5p regulates glioma cell proliferation, invasion, apoptosis, and cell cycle both in vivo and in vitro and does so by targeting VEGFA.
Footnotes
Acknowledgements
The authors thank their central laboratory for providing technical instructions and assistance. Y.H. and Y.L. are equal contributors.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was partly supported by the National Science Foundation of China (Grant Nos 81372689 and 81572475), the Anti-Cancer Association Foundation of China (Grant CSNO-2013-MSD-009) and the Foundation of Suzhou Science and Technology Development Plan Item (Grant No. SYS201324).
