Abstract
Colorectal cancer is among the most lethal of malignancies, due to its propensity to metastatic spread and multifactorial-chemoresistance. The latter property supports the need to identify novel therapeutic approaches for the treatment of colorectal cancer. MicroRNAs are endogenous non-coding small RNA molecules that function as post-transcriptional regulators of gene expression. Recently, programmed cell death 4 has been identified as a protein that increases during apoptosis. This gene is among the potential targets of miR-21 (OncomiR). Locked nucleic acid–modified oligonucleotides have recently emerged as a potential therapeutic option for targeting microRNAs. The aim of this study was to explore the functional role of locked nucleic acid-anti-miR-21 in the LS174T cell line in vitro and in vivo models. LS174T cells were treated with locked nucleic acid-anti-miR-21 for 24, 48, and 72 h in vitro. The expression of miR-21 and PDCD4 at messenger RNA (mRNA) level was evaluated by quantitative real-time polymerase chain reaction, while the protein level of PDCD4 was determined by Western blotting. Cell migratory behavior and the cluster-forming ability of cells were assessed before and after therapy. The disseminated tumor cells were assessed in the chick chorioallantoic membrane model by Alu quantitative polymerase chain reaction. Locked nucleic acid-anti-miR-21 was transfected successfully into the LS174T cells and inhibited the expression of miR-21. Locked nucleic acid-anti-miR-21 inhibited the migration and the number of cells forming clusters. Moreover, we found that locked nucleic acid-anti-miR-21 transfection was associated with a significant reduction in metastatic properties as assessed by the in ovo model. Our findings demonstrated the novel therapeutic potential of locked nucleic acid-anti-miR-21 in colon adenocarcinoma with high miR-21 expression.
Introduction
Colorectal cancer (CRC) is the third leading cause of cancer-related death. 1 The America Cancer Society has predicted that there would be 95,270 cases of colon cancer and 39,220 cases of rectal cancer in 2016 in the United States. Moreover, it is expected that more than 49,190 deaths from CRC will occur in 2016 in the United States. 2 Despite extensive clinical and preclinical developments, the prognosis of CRC is still extremely poor, because of its aggressive properties and its intrinsic resistance to most chemotherapeutic agents. 3 Our data provide a proof of principle for developing a new therapeutic approach and identifying the molecular mechanisms underlying the drug resistance in CRC.
Recently, programmed cell death 4 (PDCD4) has been identified as a tumor suppressor gene and is being suggested as a potential therapeutic target. This gene is among the potential targets of microRNA-21 (miR-21), although the molecular mechanism is still obscure. 4 It has been shown that PDCD4 can interact with several translational factors, including eukaryotic translation initiation factor-4G (eIF4G) and eukaryotic translation initiation factor-4A (eIF4A), and could suppress translation of mRNA.5,6 High levels of PDCD4 may be associated with reduced levels of dUTPase and might contribute to the tumor suppressor function of PDCD4 in Bon-1 cells. 4 The regulatory role of PDCD4 is mediated via P21, 7 Cdk4, ornithine decarboxylase, 8 carbonic anhydrase II, 9 JNK/c-Jun/AP-1,5,10 and tissue inhibitor of metalloproteinase 2 (TIMP-2). 11 The downregulation of PDCD4 in lung and colorectal cancers has been found to be associated with a poor prognosis.12,13
Chen et al. 14 evaluated the prognostic value of PDCD4 using two molecular profiling techniques complementary DNA (cDNA) and tissue microarray analysis in 124 patients with primary lung carcinomas. They identified PDCD4 as a potential prognostic factor for the prediction of disease outcome. Another study has shown that PDCD4 suppresses tumor phenotype via inhibiting c-Jun and c-Fos activation in transformed (Tx) mouse epidermal JB6, RT101, cells. 15 It has also been reported that enforced expression of PDCD4 inhibited tumorigenesis and malignant progression in K14-PDCD4 transgenic mice. 8 However, it has also been shown that miR-21 can act as cooperative repressors of PDCD4 in pancreatic ductal adenocarcinoma. 16
MiR-21 is one of the most prominent microRNAs (miRNAs) implicated in the development and progression of different cancers, which has been shown to be involved in tumor progression, 17 proliferation 18 and with an anti-apoptotic effect 19 as well as its association with tumor resistance to chemotherapy.20,21 There is a growing body of evidence showing miR-21 aberrant expression in CRC,19,20,22 suggesting its value as a prognostic biomarker.23,24
Locked nucleic acid (LNA)-anti-miRs are a new class of modified oligonucleotides 25 that bind to miRNA and can modulate miRNA expression. They are very similar to RNAs, methylene bridge connecting the 2′-oxygen atom and the 4′-carbon atom (2′-O 4′-C methylene) in the ribose ring. This bridge forms a bi-cyclic structure that locks the ribose conformation and is integral to the high stability and low toxicity in biological systems and affinity of the LNA to its complementary RNA sequences. 26 LNA oligonucleotides are stable in the modulation of RNAs, have a good aqueous solubility, and do not cause an immune response, suggesting their value for antisense-based gene silencing in gene therapy.23,27 Therefore, in the current study, we explored the anti-tumor activity of LNA-anti-miR-21 in colon adenocarcinoma cells and the modulation of PDCD4 expression, through the analysis of cell migration and metastatic features of LS174T cells. This cell line is mucin-producing colorectal cell line mimicking the condition usually happens in younger patients with interference of high amounts of mucin in treatment effectiveness. The fact that mucin production hampers effective therapy remained obscure in previous works.
Materials and methods
Cell culture
The human colorectal adenocarcinoma cell line LS174T was obtained from the National Cell Bank of Iran (Pasteur Institute, Tehran, Iran) and was maintained in Dulbecco’s modified Eagle’s medium (DMEM; Gibco, Grand Island, NY, USA) with 4.5 g/L glucose, supplemented with 10% fetal bovine serum (FBS; Bio-IDEA, Tehran, Iran), 2-mM
Cell transfection with LNA-anti-miR-21
Nucleotide sequences of miR-21 and related scientific data were obtained from a reputable site of www.mirbase.org. Human miR-21 has the accession number MIMAT0000076, with a sequence of nucleotides in the form of 5'- UAGCUUAUCAGACUGAUGUUGA-3'. The inhibition interaction between miRNAs on mRNA PDCD4 was obtained by using EMBL-EBI (MicroCosm Targets version 5), target scan program, and miRTarBase database. The LS174T cells were transfected with LNA-anti-miR-21, miRCURY LNA inhibitor (anti-miR), or scrambled LNA (Exiqon, Copenhagen, Denmark) using the X-tremeGENE siRNA Transfection Reagent Kit™ (Roche, Mannheim, Germany) according to the manufacturer’s instructions. 5′ end of both nucleotides was marked with 6-carboxyfluorescein (6-FAM). Sequences of LNA-anti-miR-21 and scrambled LNA were as follows: 5′-3′/56-FAM/ACA TCA GTC TGA TAA GCT and 5′-3′/56-FAM/GTG TAA CAC GTC TAT ACG CCC A. The sequence of negative control was analyzed using BLAST search to exclude potential hits in the human transcriptome. Both oligonucleotides were purified by high-pressure liquid chromatography (HPLC). In brief, for cell transfection, 2 × 105 LS174T cells were cultured in each six-well culture plates (SPL Life Sciences, Pocheon, Korea), and plates were kept in a cell culture incubator in 37°C and 5% CO2 for 24 h. For standard transfection, the culture medium was completely removed from each well and the cells were washed with phosphate-buffered saline (PBS; 1×) and serum-free medium for three times to decrease mucus layer on the cells, and subsequently, 1.8 mL of fresh medium containing 2.7% serum without antibiotics was added to each well containing cells. 2-µL of scrambled LNA or LNA-anti-miR (50 pmol/µL) was added separately with 5-µL X-tremeGENE siRNA transfection reagent in 200 µL Opti-MEMI (Gibco), followed by incubation in a dark room for 20 min at room temperature. The resulting complex was added to each well and slowly swirled while adding the complex to spread entirely in all parts of the plate. The plate was incubated for 10 h. Then, 200-µL FBS was added to each well, and the cells were incubated for 24, 48, and 72 h. The reverse transfection method was used for evaluation of cluster formation assay. In this method, LS174T cells were transfected in a state of suspension. Briefly, 4 × 105 cells were added to each well of 6-well low attachment plates (SPL Life Sciences), and subsequently, 1.8-mL DMEM with 2.7% FBS without antibiotics was added to each well. Transfection complexes of 200 µL were added to the cells. After 10 h, 200-µL 10% FBS was added to each well to maintain maximum viability.
Transfection efficiency was examined by fluorescence microscope (Olympus, Tokyo, Japan) and then analyzed by FACSCalibur flow cytometer (Becton-Dickinson Immunocytometry Systems, Franklin Lakes, NJ, USA).
Quantitative real-time polymerase chain reaction
Reverse transcriptase real-time polymerase chain reaction qRT-PCR was used to evaluate the efficiency of miR-21 inhibition by LNA-anti-miR-21 after 24, 48, and 72 h of transfection. Total RNA was extracted from LS174T cells after 24, 48, and 72 h of transfection by the miRCURY RNA Isolation Kit™ (Exiqon) according to the manufacturer’s instructions. The concentration of RNA was measured using a NanoDrop Epoch (BioTek Instruments, Winooski, VT, USA). cDNA was synthesized by the Universal cDNA Synthesis Kit (Exiqon). Relative levels of miRNA were examined using the SYBR Green Master Mix Kit™ (Exiqon) with specific primers which hsa-miR-21-5p (Product No. 204230; Exiqon) and hsa-let-7a-5p (Product No. 205727; Exiqon) 28 as an internal control (Exiqon) to normalize quantitative real-time PCR (qRT-PCR). The relative miR-21 and let-7a expression were calculated using the ΔΔCT method. Moreover, for evaluation of PDCD4 gene expression, total RNA was extracted as described above and cDNA was synthesized by the Fermentas kit (Fermentas Life Science, Vilnius, Lithuania) according to the manufacturer’s instructions. qRT-PCR was performed using the SYBR® Green Master Mix Kit (Exiqon) according to the manufacturer’s instructions. The primers’ sequences for PDCD4 were as follows: forward, 5′-AAAGGGAAGGTTGCTGGATAGG-3′; reverse, 5′-CACAGTTCTCCTGGTCATCATCA-3′. GAPDH was used as the internal control gene (forward, 5′-AGTCCACTGGCGTCTTCAC-3′; reverse, 5′-AGGCATTGCTGATGATCTTGAG-3′). The relative PDCD4 and GAPDH expression were calculated using the ΔΔCT method. The StepOnePlus™ (Applied Biosystems, Foster City, CA, USA) instrument was used for real-time PCR experiments under the following conditions: polymerase activation at 95°C for 10 min followed by 40 cycles at 95°C for 10 s and 60°C for 1 min.
Western blot analysis
The level of protein expression of PDCD4 was determined after 24, 48, and 72 h following transfection in the three groups of cells (transfected by LNA-anti-miR, transfected by scrambled LNA, and untreated cells) by using Western blot analysis. Total protein was extracted using RIPA (Radioimmunoprecipitation) lysis buffer (Santa Cruz Biotechnology, Santa Cruz, California, USA) containing RIPA, phenyl methyl sulfoxide (PMSF), sodium orthovanadate solution, and protease inhibitor cocktail solution. Protein concentrations were determined by the Quick Start™ Bradford Protein Assay Kit™ (Bio-Rad, Hercules, CA, USA). Electrophoresis was performed in discontinuous system of Laemmli 29 at 15°C in a vertical slab filled with electrophoresis buffer (25-Mm Tris, 192-mM glycine, and 0.1% sodium dodecyl sulfate (SDS), pH 8.3). Equal aliquots of solubilized total protein sample (100 µg per lane) were resolved on 12% polyacrylamide gel electrophoresis with 6% stacking gel. A prestained protein marker (9–170 kDa; CinnaGen, Tehran, Iran) was used to estimate the molecular weight of proteins resolved by electrophoresis. Electrophoresis was applied with 20 mA per gel until the bromophenol blue had reached the separation gel. The current was then increased to 30 mA until the dye had migrated through the separation gel. Afterward, the gels were removed and the proteins were transferred onto the nitrocellulose membrane (Amersham, Piscataway, NJ, USA). The membranes were then blocked with buffer containing 5% skimmed milk (Sigma, St. louis, USA) and 0.05% Tween-20 (polyoxyethylene-20 sorbitan monolaurate)(Merck, Darmstadt, Germany) at 4°C. Mouse monoclonal antibody anti-human PDCD4 (Santa Cruz Biotechnology) and mouse monoclonal antibody anti-human β-actin (Santa Cruz Biotechnology) were used at 1:200 dilution. The secondary antibody, horseradish peroxidase–conjugated goat anti-mouse IgG (Santa Cruz Biotechnology) was used at 1: 2000 dilutions. The position of the protein was visualized using diaminobenzidine (DAB; Sigma-Aldrich Company Ltd, Gillingham, UK) reagent. Results of densitometric analyses of Western blots were measured by ImageJ software ver. 1.42q (National Institute of Health, Bethesda, MD, USA). The β-actin was utilized as an internal control for normalization.
Scratch cell migration assay
For the migration assay, 2 × 105 cells/well were seeded into six-well plates (SPL Life Sciences) and grown to 90% confluence at 37°C with 5% CO2. Then, culture medium on the cells was removed; cells were washed with PBS solution and exposed overnight in medium containing 0.5% serum. Afterward, a direct scratch on the bottom plate using three vertical scratches was created by using a sterilized glass rod with a sharp tip; the cells were gently washed with PBS to remove non-adherent cells separated from the plate’s bottom area and replaced with 1.8 mL of fresh medium containing 2.7% FBS solution. After 24, 48, and 72 h following transfection in three groups of cells (transfected by LNA-anti-miR, transfected by scrambled LNA, and untreated cells), cell migration was evaluated by counting cells at 24, 48, and 72 h post transfection that migrated from the scratch edge. The average numbers of migratory cells in three random fields were counted at 200× magnification using an inverted light microscope (Nikon TS100, Tokyo, Japan).
Cluster-formation assay
For the cluster formation assay, cell suspensions were resuspended in DMEM supplemented with 2.7% FBS. 10h after reverse transfection, 2 × 104 cells were seeded into 24-well ultra-low attachment plates (Corning, Corning, NY, USA) and incubated at 37°C with 5% CO2 for 48 h. Suspensions of cell clusters were transferred into six-well plates (SPL Life Sciences). After 24-h incubation at 37°C with 5% CO2, cell clusters attached to the bottom of the plate for evaluation of biomass. The clusters were stained with cell stain solution (Cell Biolabs, San Diego, CA, USA) for 10 min and then washed with PBS. The average numbers of cells forming the cluster in three random fields were counted at 200× magnification using an inverted light microscope (Nikon TS100) in the three groups (transfected with LNA-anti-miR, scrambled LNA, and untreated cells).
In ovo experiments
To assess spontaneous metastasis, fertilized chicken eggs were randomly allocated to one of four groups: healthy chick embryos, scrambled LNA, LNA-anti-miR, and control (untreated cells). Fertilized chicken eggs were incubated in a rotating incubator at 38°C and 65% humidity for 10 days. Afterward, the air sac in the blunt end was determined using light source to candle the eggs. The chorioallantoic vein was detected and marked; 1-cm by 0.5-cm rectangle away from the area of capillary network connections was drawn with a pencil on the egg shell. Then, the area including blunt end and around the square was then cleaned using an iodine swab. A hole was made using a sterile pin in air sac and central part of the rectangle. Following suction made by the pump in the air sac, the new air pocket was formed under the drawn rectangle hole showing that chorioallantoic membrane (CAM) successfully separated from the egg shell. After CAM coming down, a small window above the rectangle, drawn on the egg shell, was removed with a sterile forceps. CAM was abraded using a cotton swab. A sterile plastic ring was placed on CAM area. All procedures were performed in accordance with the ethical standards of the local ethics committee of Mashhad University of Medical Sciences (permit number: 921797).
Chicken model for metastasis quantification
To study the metastatic features of tumor cells, burden in the livers of chicken embryo was determined by real-time quantitative PCR (qPCR) to detect human Alu repeats. Alu sequences are found in 500,000–1,000,000 copies in the haploid genome. They make an excellent target or marker for human DNA. After 48-h transfection, 105 cells were injected into an approximately 0.8-cm plastic ring placed on the 10-day-old chick CAM. After 8 days, the eggs were placed on ice for 3 h to anesthetize and euthanize. To assess metastasis, the livers were harvested and genomic DNA was extracted using the DNA Isolation Kit (Genet Bio, Cheonan, Korea). DNA concentration and purity were measured spectrophotometrically at an optical density (OD) of 260 to 280 nm with NanoDrop Epoch (BioTek). The concentration of genomic DNA extracted was regulated to 50 ng/µL. Genomic DNA was diluted 1:10 in nuclease-free water before using it in the subsequent PCR reaction for amplification. 1-µL diluted genomic DNA was amplified in a total volume of 10-µL reaction, containing 1× RealQ Plus Master Mix Green (Ampliqon, Odense, Denmark), dH2O, and 200 nmol/L of human Alu primers (Alu-F 5′-ACGCCTGTAATCCCAGCACTT-3′ and Alu-R 5′-TCGCCCAGGCTGGAGTGCA-3′) and chick glyceraldehyde-3-phosphate dehydrogenase (chGAPDH) specific primers (chGAPDH-F 5′-GAGGAAAGGTCGCCTGGTGGATCG-3′and chGAPDH-R 5′-GGTGAGGACAAGCAGTGAGGAACG-3′) as endogenous controls. Metastasis was assessed using an ABI StepOnePlus™ instrument under the following conditions for Alu sequences: polymerase activation at 95°C for 15 min followed by 30 cycles at 95°C for 30 s, 63°C for 30 s, and 72°C for 30 s. Quantitative analysis by the ΔΔCT method for comparisons between groups was performed. Each assay included a negative control (no template controls, NTC), genomic DNA extracted from the liver of healthy chick embryos to determine the threshold between LS174T cells invaded and non-invaded cells.
Data analysis
The experiments were carried out in triplicate and repeated at least twice. Data were expressed as mean values ± standard deviation (SD) and analyzed by two-way analysis of variance (ANOVA) with Fisher’s test followed by Scheffé’s multiple comparisons test and compare means test-Dunnett. Data were analyzed using SPSS version 22 statistical software (IBM, Chicago, IL, USA). Statistical significance was set at **p < 0.01.
Results
LNA-anti-miR-21 reduced the expression of miR-21
We aimed to transfect the LS174T cells with FAM-labeled scrambled oligonucleotides (scrambled LNA group) to evaluate the transfection efficiency. Based on primary optimization experiments, different concentrations of the scrambled LNA were added to the cells and the transfection efficacy was assessed by fluorescence microscopy and flow cytometry. The best results were obtained at a concentration of 50 pmol of scrambled LNA and 5-µL X-tremeGENE siRNA transfection reagent. We found that the transfection efficacy was >85% (Figure 1(a) and (b)).

Transfection of LS174T cells. (a) LS174T cells were transfected with scrambled LNA 6-carboxyfluorescein (6-FAM)-labeled, as detected by fluorescent microscopy (×200); scale bars: 50µm; (b) fluorescence-activated cell sorting (FACS) analysis; (c) miR-21 expression after 24-, 48-, and 72-h transfection by qRT-PCR; data were mean ± 2SE of three independent experiments in the graph (**p < 0.01).
In order to see whether the target was inhibited, the expression level of miR-21 was evaluated by qRT-PCR after transfection for the 24, 48, and 72 h. This analysis revealed the downregulation of miR-21 in the LNA-anti-miR group. At all three time points, the expression of miR-21 was significantly lower in the LNA-anti-miR group compared with scrambled LNA and untreated groups (**p < 0.01; control: 1 ± 0.0, scrambled LNA: 1.01 ± 0.088, and LNA-anti-miR: 0.018 ± 0.013; Figure 1(c)).
LNA-anti-miR-21 modulated the PDCD4 expression in LS174T cells
The expression pattern of PDCD4 was determined in cells before and after transfection with LNA-anti-miR-21 compared with scrambled LNA and untreated groups at mRNA and protein levels. We observed that the level of mRNA PDCD4 was significantly increased at 48 h after transfection with LNA-anti-miR-21, although this effect was more pronounced after 72 h (Figure 2(a)). Notably, when LS174T cells were transfected with LNA-anti-miR, a significant increase in PDCD4 mRNA expression levels was apparent compared with the control and scrambled LNA groups (**p < 0.01).

PDCD4 expression. (a) The mRNA PDCD4 level 24, 48, and 72 h after transfection was analyzed by qRT-PCR. The expression was normalized to GAPDH. Mean ± 2SE of three independent experiments in the graph (**p < 0.01). (b) Protein expression of PDCD4 after 24-, 48-, and 72-h transfection as detected by Western blot analysis. The expression was normalized to beta-actin protein. (c) The regression curve was created with the distance from surface of the membrane and logarithmic of the molecular weight. (d) Density ratio of PDCD4/β-actin by Western blotting performed 24, 48, and 72 h after transfection; bar diagram, mean ± 2SE of three independent experiments in the graph (**p < 0.01).
There was a statistically significant inverse correlation (r = −0.502; **p < 0.01) between miR-21 inhibition and PDCD4 mRNA, which may be due to translational repression instead of mRNA degradation by miR-21 because an r indicates a weak correlation between the variables. We also evaluated the protein expression of PDCD4 (Figure 2(b)). A strong inverse correlation was found between miR-21 inhibition and PDCD4 protein (r = −0.853). The regression equation was used to evaluate the molecular weight with respect to the distance from the surface of membrane (Figure 2(c)). At all three time points, these data showed that the highest amount of PDCD4 protein was associated to LNA-anti-miR-21-transfected cells (control: 0.16 ± 0.25, scrambled LNA: 0.14 ± 0.21, and LNA-anti-miR: 2.85 ± 1.37), and this effect was greater after 48 h.
After 24 h in the LNA-anti-miR-transfected cells, the expression level of PDCD4 protein was increased, but in the untreated cells and cells transfected with scrambled LNA, PDCD4 protein was not detected. At 24 h, this increasing pattern of PDCD4 protein was continued at 48 h and decreased at 72 h after transfection, but this reduction was not less than that of 24 h after transfection (Figure 2(b)). In untreated and transfected cells treated with scrambled LNA, a very low level of PDCD4 protein was detected after 48 and 72 h transfection that may be because of high expression of miR-21 in the cells that leads to significant translational repression instead of mRNA degradation (Figure 2(d)).
LNA-anti-miR-21 reduced cell migration
A cell scratch assay was performed to estimate the migratory behaviors of the cells after 24-, 48-, and 72-h transfection. This analysis showed that LNA-anti-miR-21 reduced the migratory properties of LS174T cells compared with scrambled LNA and untreated cells (Figure 3(a)). In addition, the average numbers of cells moved across the scraped edge were counted at 24, 48, and 72 h post transfection. The average number of LS174T cells transfected with LNA-anti-miR-21 that migrated for all three time points was 47.44 ± 14.82 cells per site. Notably, this average number was significantly lower than that for the scrambled LNA (97.77 ± 36.97 cells per site) and untreated (98.78 ± 38.87 cells per site) cells (Figure 3(b)).

LNA-anti-miR-21 decreased cell migration. (a) Cell migration after 0, 24, 48, and 72 h in three groups: control, scrambled LNA, and LNA-anti-miR-transfected cells (magnification: ×200). (b) Different migration abilities of LS174T cells transfected with LNA-anti-miR compared with scrambled LNA–transfected cells and control groups in all three time points (**p < 0.01).
LNA-anti-miR-21 decreased cluster-forming ability of LS174T cells
We investigated the effect of LNA-anti-miR-21 on cluster-forming ability of LS174T cells. The average number of cells forming the cluster was lower in the LS174T cells transfected by LNA-anti-miR-21 after 48-h transfection, compared with scrambled LNA and untreated cells. In particular, the average number of cells forming the cluster in transfected cells with LNA-anti-miR-21 was 20.67 ± 3.21, compared with 51.33 ± 1.53 cells per site for scrambled LNA and 50.0 ± 3.0 cells per site for untreated cells (Figure 4).

Transfection of the cells with LNA-anti-miR-21 decreased cluster-forming ability of LS174T cells. Different number of counted cells in each cluster transfected with LNA-anti-miR and scrambled LNA compared with control. In cells transfected with LNA-anti-miR, the cluster was significantly smaller compared with the other groups (**p < 0.01).
Inhibition of miR-21 reduced liver metastasis
LS174T cells were successfully grafted onto the CAM, outlined in Figure 5(a), to investigate the functional significance of metastasis in vivo assay. LS174T cells grafted CAM model were treated on day 10 with LNA-anti-miR-21, scrambled LNA, and untreated cells, exposed till day 18. After incubation, a primary tumor as the primary CAM tumor developed, allowing access to the chick vasculature for metastasis to more distant chick tissues. Then, livers were harvested and analyzed for metastases (Figure 5(b)). OD260/OD280 ratios of 1.4–1.8 were accepted to be adequate for qPCR for human Alu repeat sequences in DNA. The number of tumor cells in the liver was determined by comparison with a standard curve in which we plotted the qPCR signal for Alu repeats against known numbers of LS174T cells (Figure 6(a)). Interestingly, we observed no metastasis in embryos treated with LNA-anti-miR-21 (Figure 6(b)). Moreover, the average number of metastases was higher in the control group (2024.67 ± 1271.22 cells/sample) or scrambled LNA group (1490.33 ± 571.24 cells/sample) compared with the embryos treated with LNA-anti-miR-21 groups (0.05 ± 0.032 cells/sample; Figure 6(c)). Human genomic DNA was higher in the control group (0.41 ± 0.3 cells/sample) or scrambled LNA group (0.3 ± 0.13 cells/sample) compared with the embryos treated with LNA-anti-miR-21 groups (0.00 ± 0.00001 cells/sample; Figure 6(d)). However, cells treated with scrambled LNA or untreated cells had a significant higher rate of metastasis to liver.

Inhibition of miR-21 reduced liver metastasis in CAM model. (a) Scheme of the spontaneous metastasis chick model. (b) The inhibition of spontaneous metastasis cells in a chick model treated with the LNA-anti-miR group compared with the untreated and scrambled LNA–treated cells groups. Cells were grafted in CAM at day 10; tumors and livers were harvested on day 18. MiR-21 knockdown by LNA-anti-miR-21 reduced metastasis of LS174T cells in the chick embryo spontaneous metastasis model.
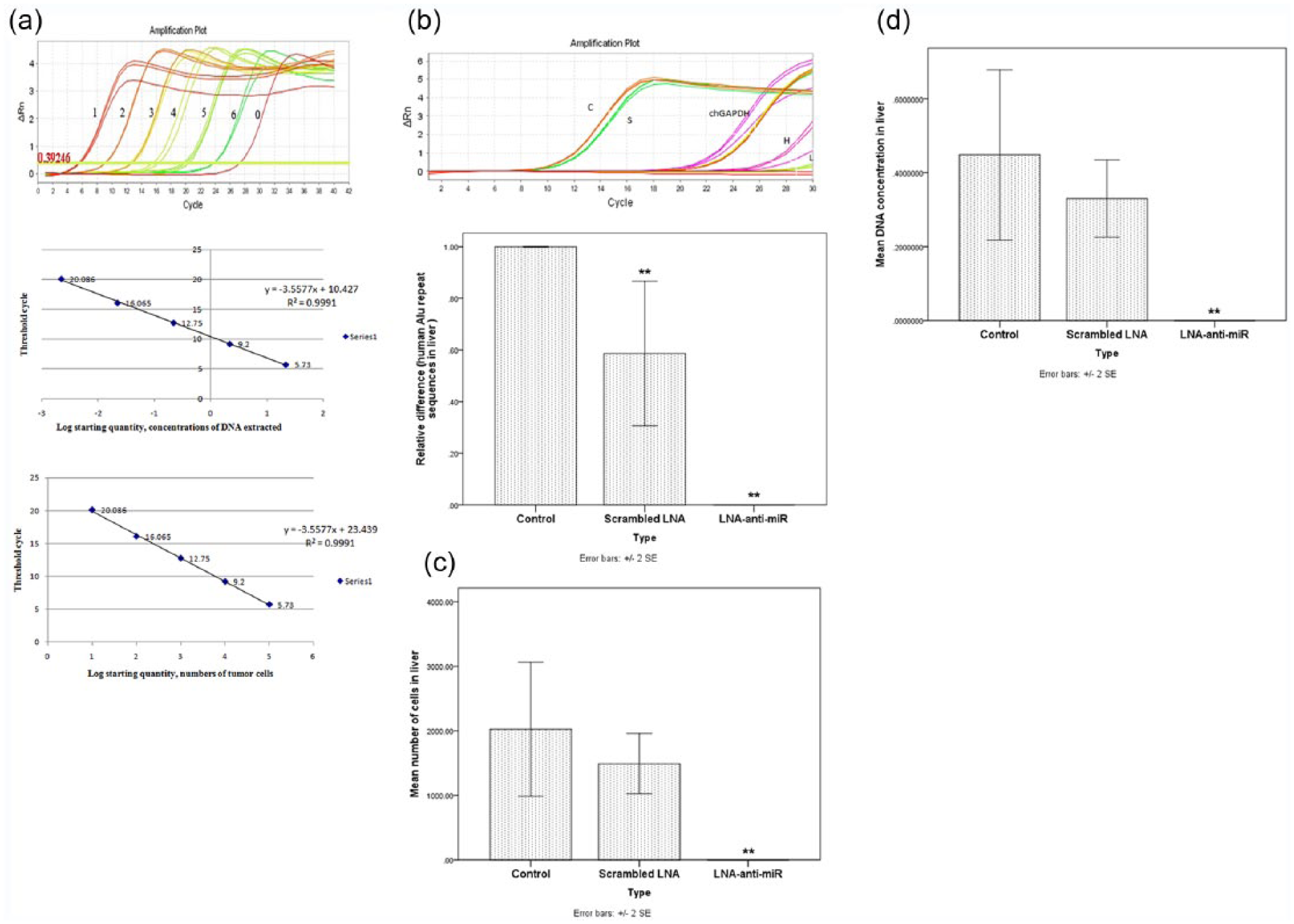
Validation of Alu PCR assays. (a) Human genomic DNA was extracted from LS174T cells, which had been serially diluted by 10-fold (lines 1, 22 ng/100,000 cells; lines 2, 2.2 ng/10,000 cells; lines 3, 0.22 ng/1000 cells; lines 4, 0.022 ng/100 cells; and lines 5, 0.00022 ng/10 cells), and then, Alu PCR was performed to detect human-specific Alu repeats in the liver lobes of chick embryo. The Alu standard curve was generated with the concentration of total DNA/numbers of tumor cells and threshold cycle of the 10-fold serially diluted human genomic DNA samples. These results were used to estimate the number of LS174T cells in liver by comparison with a standard curve. (b) The livers from each chick embryo were harvested 8 days later and subjected to quantitative real-time Alu PCR analysis. chGAPDH was used as an internal control. The ΔΔCT method was used for data analysis, and the untreated group was considered as a reference for each time point. Data were mean ± SD of three independent experiments (**p < 0.01). (c) Number of tumor cells metastasized to the liver was counted. Data were mean ± SD of three independent experiments (**p < 0.01). (d) Concentration of human genomic DNA metastasized to the liver was estimated. Data were mean ± 2SE of three independent experiments in the graph (**p < 0.01).
Discussion
To the best of our knowledge, this is the first study evaluating the inhibitory effect of LNA-anti-miR-21 and exploring its molecular mechanisms in vitro and in vivo, in the chick CAM model derived from human colorectal adenocarcinoma LS174T cells. Our results suggest that miR-21 expression could be of relevance to the critical biological behavior of colorectal neoplasia. We demonstrated that inhibition of miR-21 using novel LNA-anti-miR-21 suppressed its expression in human colorectal adenocarcinoma LS174T cells and reduced cell migration and liver metastasis in the CAM model, most likely through modulation of PDCD4.
There is mounting evidence showing the important role of miR-21 in different types of cancer, including CRC,17,18,25,26,30–33 which has been suggested to be associated with poor prognosis and chemoresistance of CRC,34,35 indicating its value as a potential therapeutic target. Furthermore, there appears to be an inverse correlation between miR-21 expression and PDCD4 expression15,36,37 which is in agreement with our data. It has also been documented that reduced expression of this tumor suppressor gene in CRCs is associated with poor prognosis.13,15 In particular, Fassan et al. 36 evaluated PDCD4 expression in 300 polypoid lesions of the colon mucosa (50 hyperplastic polyps, 50 serrated adenomas, 50 tubular adenomas with low-grade intraepithelial neoplasia, 50 tubular adenomas with high-grade intraepithelial neoplasia), 50 colon adenocarcinomas, and in 50 biopsy samples obtained from patients with irritable bowel syndrome as normal controls. They showed that miR-21 expression was upregulated in pre-neoplastic/neoplastic samples, consistent with PDCD4 downregulation. Moreover, it has been reported that PDCD4 could affect on tumor progression, malignant transformation, tumorigenesis,8,15 and apoptosis. 38 Wei et al. 39 investigated the role and mechanism of miR-21 in promoting the 5-fluorouracil (5-FU) resistance of pancreatic cancer cells in 5-FU resistance cell line PATU8988/5-FU. They found that miR-21 regulated 5-FU drug resistance PATU8988 and PANC-1 cells via decreasing the expression of PTEN and PDCD4. In turn, enforced expression of PDCD4 attenuated the effects of miR-21 in 5-FU resistance cell line PATU8988/5-FU. Shi and colleagues 40 found that exosomal levels of miR-21 from cerebrospinal fluids were related with poor prognosis and tumor recurrence of glioma patients, whose expression was inversely correlated with PDCD4 expression. A similar study also showed that the downregulation of PDCD4 was correlated with aromatase inhibitor resistance and poor prognosis in estrogen receptor–positive breast cancer and forced overexpression of PDCD4 resensitized aromatase-resistant cells to aromatase inhibitors or hormone deprivation. 41 Zhang et al. 42 have reported that modulation of miR-21 could affect on PDCD4 in hepatocellular carcinoma.
These observations provide a proof of principle for the modulation of PDCD4 expression via targeting miR-21 in CRC as a novel therapeutic option. Therefore, in this study, we investigated the inhibitory effects of LNA-anti-miR-21 in LS174T cells as a possible treatment for CRC. The LS174T cell line is able to produce large amounts of mucin and inhibit effective transfection.43,44 At the optimized level, we found that transfection efficiency was more than 85%. This amount of transfection is considered to be important because one of the key challenges in the improvement of effective miRNA-based therapy is the delivery of stable and efficient agents into target cells. 45 Although the role of mucin in the behavior of colon cancer cells is not clear, other reports have suggested that mucinous adenocarcinoma is associated with a poorer outcome. 43 In addition, mucinous carcinoma has a higher incidence in the proximal colon and among younger patients than non-mucinous adenocarcinoma. 43 Our results showed that the expression of miR-21 was high, while the level of PDCD4 protein was lower in LS174T cells before transfection with LNA-anti-miR-21. However, we found that PDCD4 protein was markedly enhanced after transfection with LNA-anti-miR-21.
According to this study, miR-21 has a negative regulatory effect on PDCD4 and it has anti-apoptotic effects. 38 LNA-anti-miR-21-transfected cells significantly increased PDCD4 protein levels, but it almost caused weak alternations in PDCD4 mRNA levels. These findings showed that miR-21 inhibits PDCD4 protein production. It further suggests that miR-21 regulates PDCD4 at the level of translational repression, instead of mRNA degradation. This finding strongly suggests that a high level of miR-21 expression in LS174T cells downregulated PDCD4 protein expression at post-transcription level, and enhanced tumor cell invasion in vitro 38 and distant metastasis in vivo. Our findings showed that LNA-anti-miR-21 reduced cell migration and cluster-forming ability of LS174T cells. In particular, we successfully developed a CAM model using LS174T CRC cells. Also, the average number of tumor cells in the liver was significantly reduced in the eggs treated with LNA-anti-miR compared with the scrambled LNA and untreated cells. It seems that this performance is due to the effect of inhibiting the activity of matrix-metalloproteinase and inhibition of u-PAR, which is in agreement with several previous studies.35,38,46–51
Conclusion
Our study confirms that miR-21 triggers carcinogenic progression. Nowadays, miRNA-based therapeutic drug has moved to clinical trials, and it is possible to become a potential therapeutic target for cancers. In aggregate, our data illustrated that the inhibition of oncogenic role of miR-21 impairs the metastatic characteristic of colon adenocarcinoma cells, supporting further investigation on the therapeutic potential of LNA-anti-miR-21 as a new option for treatment of CRC.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by the Mashhad University of Medical Sciences and Isfahan University of Medical Sciences (grant no. 921797).
