Abstract
MicroRNAs are short non-coding RNAs, which have been implicated in several biological processes. Aberrant expression of the microRNA miR-122 has frequently been reported in malignant cancers. However, the mechanism underlying the effects of miR-122 in renal cell carcinoma remains unknown. The aim of this study was to determine the biological function of miR-122 in renal cell carcinoma and to identify a novel molecular target regulated by miR-122. We measured the expression levels of Sprouty2 in six renal cell carcinoma tissue samples and adjacent non-tumor tissues by western blot analysis. We then used reverse transcription polymerase chain reaction to measure miR-122 levels in 40 primary renal cell carcinoma and adjacent non-malignant tissue samples. The effects of miR-122 down-regulation or Sprouty2 knockdown were evaluated via Cell Counting Kit-8 assay, flow cytometry, and western blot analysis. The relationship between miR-122 and Sprouty2 was determined using dual-luciferase reporter assays. Sprouty2 was down-regulated in renal cell carcinoma tissue samples compared with adjacent normal tissue. In contrast, miR-122 was up-regulated in primary renal cell carcinoma tissue samples compared with adjacent normal tissue samples. Down-regulation of miR-122 substantially weakened the proliferative ability of renal cell carcinoma cell lines in vitro. In contrast, Sprouty2 knockdown promoted the in vitro proliferation of renal cell carcinoma cell lines. The spry2 gene could therefore be a direct target of miR-122. In conclusion, miR-122 could act as a tumor promoter and potentially target Sprouty2. MiR-122 promotes renal cell carcinoma cell proliferation, migration, and invasion and could be a molecular target in novel therapies for renal cell carcinoma.
Introduction
Renal cell carcinomas (RCCs) are the third most common urological neoplasm after prostate and bladder cancer and account for 2% of all cancer-related deaths. 1 Clear cell renal cell carcinoma (ccRCC) is the most common subtype of RCC, accounting for approximately 70% of all cases. 2 RCC is resistant to radiotherapy and chemotherapy, and therefore usually treated by surgical intervention. 3 Moreover, an unusually high proportion of patients with RCC are diagnosed with progressive metastasis; however, there is a dearth of biomarkers for the early detection of this condition. A few well-established lifestyle risk factors for RCC, such as cigarette smoking, obesity, hypertension, and diabetes, have been identified; however, the exact etiology remains unclear. 4 Therefore, research on the molecular mechanisms involved in the initiation and progression of RCC is of great importance and may provide a new approach for targeted RCC therapy.
MicroRNAs (miRNAs) form a large family of 21–22 nucleotide RNAs, which affect post-transcriptional regulation by binding to the 3′-untranslated region (UTR), leading to inhibition of translation, or messenger RNA (mRNA) cleavage. 5 Recently, there has been increasing evidence implicating miRNAs in various aspects of tumorigenesis. 6 Many miRNAs, including miR708, miR-199a, miR-204, and miR141, are aberrantly expressed in RCC7–10 and likely contribute to the development and progression of RCC tumorigenesis. Liver-specific miR-122 is the most abundant miRNA isolated from the liver and is involved in the regulation of hepatocyte development, differentiation, cholesterol metabolism, and stress response, as well as the inhibition of hepatocellular carcinomas (HCCs).11,12 MiR-122 is abnormally expressed in breast cancer (BC) and cutaneous T-cell lymphoma (CTCL)13,14 and overexpressed in RCC and could be a reliable diagnostic marker and effective therapeutic target.15–17
Sprouty2 (Spry2) is a member of the mammalian Sprouty family of signal transduction proteins that inhibit the Ras/mitogen-activated protein kinase signaling pathway.18–20 Spry2 plays an important role in modulating pathways that are crucial to the development or progression of neoplasms, including cell proliferation, invasion, migration, and apoptosis.21–23 Spry2 may serve as an inhibitor of tumors, including BC, prostate cancer, HCC, lung cancer, gliomas, and RCC.24–29 The purpose of this study was to investigate the function of miR-122 in RCC and to determine whether Spry2 is a direct target of miR-122.
Materials and methods
Ethics statement
The study protocol was in accordance with the ethical standards of the Declarations of Helsinki and Istanbul. The protocol of this study was approved by the local Ethics Committee of the First Affiliated Hospital of Nanjing Medical University, and written informed consent was obtained from all patients.
Cell culture and tissue specimens
We used two human cell lines, CAKI-1 and 786-0, for our in vitro research. CAKI-1 and 786-0 cells were cultured in RPMI-1640 and McCoy’s 5A media, respectively, supplemented with 10% fetal bovine serum (FBS), 100 µg/mL penicillin, and 100 µg/mL streptomycin. All cell lines were incubated at 37°C in a 5% CO2 chamber.
Primary RCC and adjacent non-malignant tissues were obtained from patients who underwent partial or radical nephrectomy in our center. The specimens were immediately frozen in liquid nitrogen after surgery and stored at −80°C until further analysis. Only samples containing >70% tumor tissue were used for the extraction of total RNA. Specific information of each tissue donor is exhibited in Table 1.
Clinical characteristics of patients.

RCC: renal cell carcinoma.
Cell infection
Cells were seeded in six-well plates at 70% confluence on the day before transfection. The infection was carried out using Lipofectamine 2000 reagent (Invitrogen, CA, USA) according to the manufacturers’ protocol. Six hours after transfection, the culture medium was replaced with RPMI-1640/McCoy’s 5A and FBS. The mimic sequences of miR-122 were as follows: sense 5′-UGGAGUGUGACAAUGGUGUUUG-3′ and antisense 5′-AACACCAUUGUCACACUCCAUU-3′. In addition, RNA with no sequence homology to any human genomic sequence was used as the negative control (NC): sense 5′-UUCUCCGAACGUGUCACGUTT-3′ and antisense 5′-ACGUGACACGUUCGGAGAATT-3′. The miR-122 inhibitor sequence was 5′-CAAACACCAUUGUCACACUCCA-3′. The NC inhibitor sequence was 5′-CAGUACUUUUGUGUAGUACAA-3′.
The small interfering RNA (siRNA) sequences targeting Spry2 were as follows: sense 5′-CCCAGCAGGUACAUGUCUUTT-3′ and antisense 5′-AAGACAUGUACCUGCUGGGTT-3′. All sequences were purchased from GenePharma (Shanghai, China).
RNA extraction and quantitative real-time polymerase chain reaction
Total RNA was extracted from tissues and cell lines using Trizol reagent (Invitrogen) according to the manufacturers’ protocol. RNA concentrations were measured using a NanoDrop (Thermo Fisher Scientific, Waltham, MA, USA). To analyze the expression of miR-122, total RNA (10 ng) was transcribed into complementary DNA (cDNA) using the TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Carlsbad, CA, USA) according to the manufacturers’ protocol. Real-time polymerase chain reaction (PCR) was carried out with the TaqMan MicroRNA Assay Kit (Applied Biosystems). U6 expression was used as an internal control. For Spry2, RNA was reverse-transcribed into cDNA using a PrimerScript One-Step RT-PCR Kit (Takara, Dalian, China) according to the manufacturers’ instructions. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used as an internal reference. The sequences of primers were as follows: Spry2: forward 5′-ATCCAGAGACAAGACATGTAC-3′, reverse 5′- TTCAGATGTGTTCTAAGCC-3′; GAPDH: forward 5′-GAAGGTGAAGGTCGGAGTC-3′, reverse 5′-GAAGATGGTGATGGGATTTC-3′ (synthesized by GeneRay, Shanghai, China). All reactions were performed in triplicate using a 7900HT Fast Real-Time PCR System (Applied Biosystems). Every experiment described above was repeated at least three times.
Western blotting assay
Cell lines and human RCC tissues were prepared in ice-cold radioimmunoprecipitation assay buffer (Keygene, Nanjing, China). Using BCA protein assay, the total cell proteins were measured. Equivalent amounts of protein samples were resolved by 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred onto a polyvinylidene fluoride membrane (Millipore, Billerica, MA, USA), which was then blocked for 1 h with 5% nonfat milk in PBST. After incubation overnight with primary antibodies at 4°C, the polyvinylidene fluoride membranes were washed three times with tris buffered saline with tween-20 (TBST) (20 mM Tris–HCl, pH 7.6, 137 mM NaCl, 0.01% Tween-20) and then incubated with horseradish peroxidase–conjugated goat anti-rabbit or horse anti-mouse secondary antibody at room temperature for 2 h. Anti-β-tubulin (Abcam, Cambridge, MA, USA) and mouse anti-Spry2 (Abcam) were used. The protein bands were visualized using chemiluminescence (Thermo Fisher Scientific, USA). β-Actin was used as an internal reference. Every experiment described above was repeated at least three times.
Cell proliferation assay
Cell Counting Kit-8 (CCK-8) assay (Beyotime, Shanghai, China) was used to estimate the proliferation, according to the manufacturers’ instructions. Cells were seeded in 96-well plates at a density of 2000 cells per well. At 24, 48, 72, and 96 h after transfection, CCK-8 assays were carried out. Absorbance detected at a wavelength of 450 nm was used to determine viable cell numbers. Each treatment group was composed of three wells.
Cell cycle analysis
Forty-eight hours post-transfection, cells were collected, washed twice with ice-cold phosphate-buffered saline, and fixed with 70% ethanol at −20°C overnight. The cells were then blocked in 50 mg/mL of propidium iodide and 1 mg/mL of RNase for 30 min at room temperature. The treated cells were measured using flow cytometry (Becton Dickinson, Franklin Lakes, NJ, USA). Each sample contained at least 20,000 cells. Every experiment described above was repeated at least three times.
Luciferase reporter assay
A fragment of the Spry2 3′-UTR and a Spry2 3′-UTR that contained the putative miR-122-binding sites were cloned downstream of the luciferase gene in the pGL3-REPORT luciferase vector (Invitrogen). For reporter assays, cells were co-transfected with pGL3-3′-UTR or control reporter plasmid, miR-122 mimics or NC, and miR-122 inhibitor or NC inhibitor. Luciferase activities were measured via dual-luciferase assays (Promega, Madison, WI, USA) 48 h after co-transfection and normalized against the activity of the Renilla/firefly luciferase gene.
Statistical analysis
The results are expressed as mean ± standard deviation (SD). Differences between groups were subjected to Student’s t-test. p < 0.05 was considered statistically significant. All statistical calculations were performed using SPSS 11.0 (Chicago, IL, USA).
Results
MiR-122 and Spry2 are inversely expressed in resected RCC specimens
MiR-122 expression was analyzed in 40 primary RCC and 40 non-malignant samples using TaqMan quantitative reverse transcription polymerase chain reaction (qRT-PCR). The expression of miR-122 was significantly decreased in tumors compared with non-malignant control tissues (Figure 1(a)). In contrast, western blot analysis revealed that Spry2 protein expression was significantly lower in tumor tissues than in adjacent patient-matched normal tissues (Figure 1(b) and (c)). This inverse expression suggested that miR-122 and Spry2 were potential to be coordinately regulated during tumor progression.
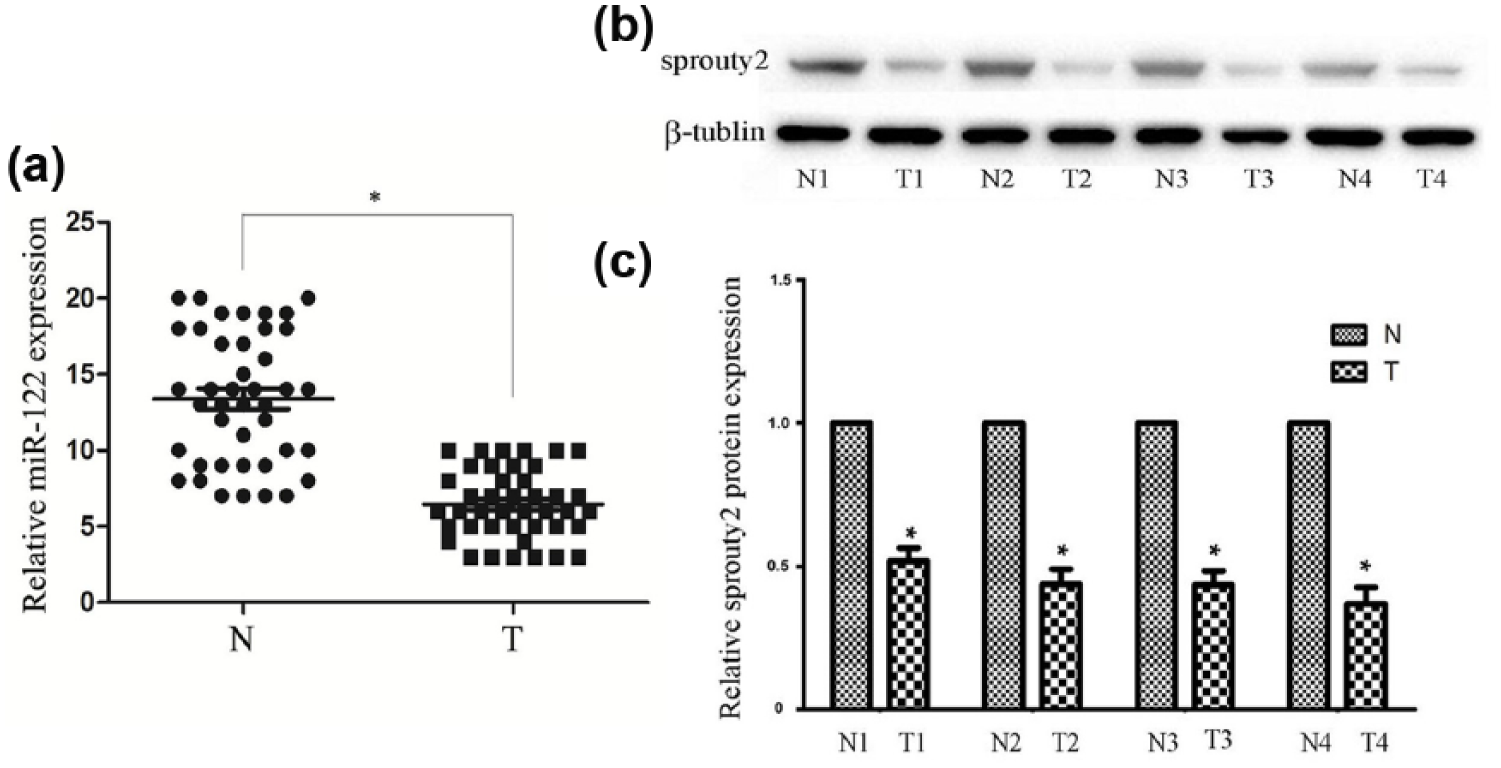
The expression of miR-122 and Sprouty2 in human samples: (a) the expression of miR-122 in cancerous specimens (n = 40) was significantly lower than the normal group (n = 40) by qRT-PCR assay, (b) western blotting assay also suggested that the Sprouty2 protein expression was significantly lower than the normal group, and (c) quantitative results of western blotting assay were expressed as the fold-change relative to normal group. N and T indicate normal and tumor tissues, respectively. The vertical axis represents the ΔCT. U6 was used as an internal standard.
MiR-122 down-regulation suppresses RCC cell proliferation in vitro
To explore the influence of miR-122 on cell proliferation, CAKI-1 and 786-0 cells were infected with a miR-122 inhibitor or a control miRNA inhibitor. In a CCK-8 assay, reduced miR-122 expression distinctly reduced the proliferative potential of RCC cells in vitro at 96 h (Figure 2(a) and (b)). Cell cycle assay was performed to analyze the cell cycle distribution in transfected cells, to further explore the effects of miR-122 on cell proliferation. Compared with the NC inhibitor- and mock-treated cells, the percentages of miR-122 inhibitor-infected CAKI-1 and 786-0 cells in the G0/G1 phase were notably higher, suggesting that down-regulation of miR-122 caused G1 cell cycle arrest in vitro (Figure 2(c) and (d)). Thus, miR-122 enhances the proliferative potential of RCC cells in vitro.

Down-regulation of miR-122 promotes proliferation in RCC cell lines. (a, b) Cell activity post-transfection with miR-122 inhibitor or NC was evaluated by CCK-8 assay at four time points (24, 48, 72, and 96 h). UV–visible absorbance at 450 nm was used for measurement (n = 3). (c, d) Analysis of the cell cycle in 786-0 and CAKI-1 cells after transfection with miR-122 inhibitor or negative control (NC; n = 3). Comparison of the proportion of cells in G1 phase arrest is shown.
MiR-122 inhibitor enhances Spry2 protein expression
Then, CAKI-1 and 786-0 cells were transfected with a miR-122 inhibitor or a control miRNA inhibitor, and western blot analysis and qRT-PCR were performed to determine whether down-regulation of miR-122 had an impact on Spry2 expressions at the protein or mRNA levels. As shown in Figure 3(a), compared with the cells transfected with a control miRNA inhibitor, the expression of miR-122 in CAKI-1 and 786-0 cells transfected with miR-122 inhibitor was significantly lower. In Figure 3(c) and (d), expression of Spry2 protein was significantly higher in cells transfected with miR-122 inhibitor than that transfected with control miRNA inhibitor or mock cells at 72 h post-transfection. However, there was no significant difference in Spry2 mRNA expression among three groups (Figure 3(b)). These findings suggested that miR-122 reduced Spry2 expression at the translational level.

Effect of miR-122 on Spry2. (a) Expression of miR-122 in transfected 786-0 and CAKI-1 cell lines measured by real-time RT-PCR. U6 was used as an internal control. (b) Spry2 mRNA expression 48 h post-transfection with miR-122 inhibitor or NC. The expression was normalized to that of β-actin. (c, d) Spry2 protein expression 72 h post-transfection with miR-122 inhibitor or NC. β-tubulin was used as an internal control.
Spry2 deficiency facilitates cell proliferation in vitro
To explore whether miR-122 promotes cell proliferation via Spry2 deficiency, CAKI-1 and 786-0 cells was transfected with si-Spry2. We performed qRT-PCR and western blotting assays to determine whether si-Spry2 reduced Spry2 expression at the mRNA as well as protein levels in CAKI-1 and 786-0 cells. As presented in Figure 4, the expressions of Spry2 protein and mRNA in cells transfected with si-Spry2 were significantly lower than that in cells infected with control siRNA or mock cells’ transfection. In addition, CCK-8 assay indicated that compared with cells transfected with control siRNA and mock cells, cells transfected with si-Spry2 showed enhanced proliferative potential (Figure 5(a) and (b)). To investigate the roles of si-Spry2, cell cycle assay was also performed. Compared with the NC and mock cells, the percentage of cells in the G1 phase was much decreased in cells transfected with si-Spry2, 48 h after transfection (Figure 5(c) and (d)). All these findings hinted that Spry2 down-regulation promotes the proliferation of RCC cell lines in vitro.

Effect of si-Spry2 on Spry2. (a) Spry2 mRNA expression 48 h after transfection with si-Spry2 or negative control (NC; n = 3). The expression was normalized to that of β-actin. (b, c) Spry2 protein expression 72 h post-transfection with si-Spry2 or NC (n = 3). β-tubulin was used as an internal reference.

Effect of Spry2 deficiency on proliferation in vitro. (a, b) Cell activity post-transfection with miR-122 inhibitor or negative control (NC) was evaluated by CCK-8 assay at four time points (24, 48, 72, and 96 h; n = 3). UV–visible absorbance at 450 nm was used for measurement. (c, d) Analysis of cell cycle in 786-0 and CAKI-1 cells after transfection with miR-122 inhibitor or NC (n = 3). Comparison of the proportion of cells in G1 phase arrest is shown.
Spry2 is a direct target of miR-122 in RCC cells
To investigate the mechanism underlying the observed effects of miR-122, we determined whether it could directly target Spry2. Spry2 contains putative complementary miR-122-binding sites in its 3′-UTR (Figure 6(a)). We observed the relative activity of a reporter containing the wild Spry2 3′-UTR. In both cell types, the results showed that compared with cells transfected with miR-122 mimics NC and blank (or miR-122 inhibitor NC and mock), co-transfection with miR-122 mimics significantly inhibited the luciferase reporter activity; co-transfection with miR-122 inhibitor significantly promoted the luciferase reporter activity. Furthermore, for the reporter containing a mutant Spry2 3′-UTR, there was no obvious difference in the relative luciferase activity of the reporter between groups (Figure 6(b)). In conclusion, miR-122 could directly regulate Spry2 in RCC cell lines.
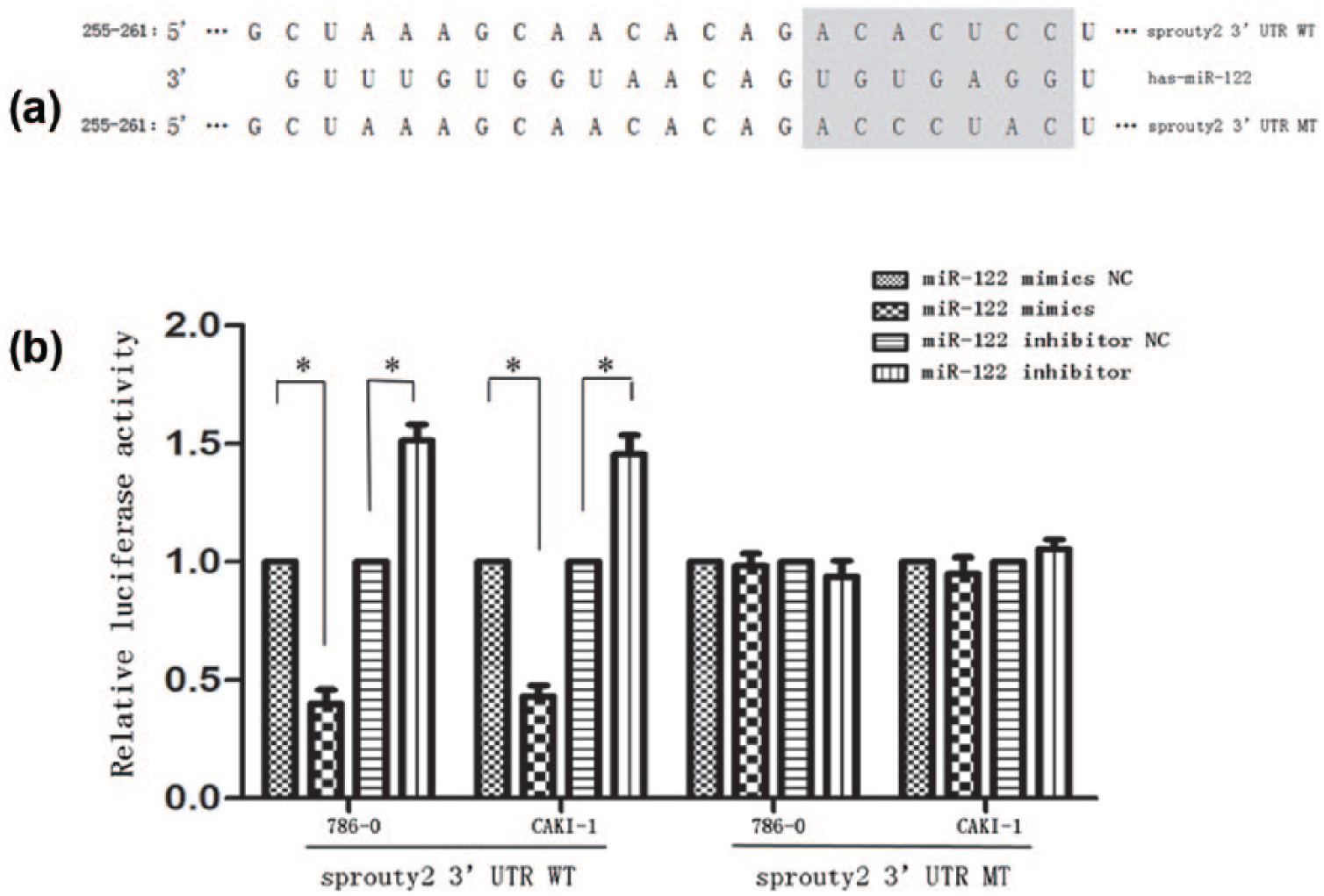
Sprouty2 is a direct target of miR-122. (a) Graphical representation of miR-122-binding sites in the wild Spry2 3′-UTR and mutant Spry2 3′-UTR. (b) Dual-luciferase reporter assay was performed in 786-0 and CAKI-1 cells. All experiments were performed in triplicate. For the relative luciferase reporter activity, there was a marked difference between groups of the reporter containing wild Spry2 3′-UTR not of the reporter containing mutant Spry2 3′-UTR.
Discussion
In this study, we found that miR-122 was overexpressed in primary RCC tissue samples compared to their adjacent non-carcinoma tissue samples. This confirms previous findings that miR-122 is up-regulated in RCC tissue samples compared to adjacent normal tissue samples.15–17 Recently, accumulating evidence has suggested that miR-122 plays either an oncogenic or a tumor-suppressing role in the progression of carcinomas. In HCC, the restoration of miR-122 can induce cell cycle arrest and apoptosis by inhibiting Bcl-W and/or CCNG1 expression. 30 Forced miR-122 expression could also inhibit HCC cell growth and promote cell cycle arrest via the Wnt/B-catenin-TCT signaling pathway. 12 Additionally, the overexpression of miR-122 could repress tumorigenesis by targeting IGF1R and modulating the PI3K/AKT/Mtor/p70s6k pathway in BC. 13 These findings support the hypothesis that miR-122 acts as a tumor suppressor in HCC and BC. In contrast, miR-122 was overexpressed in advanced CTCL compared with stationary phase T-cells in CTCL, suggesting that miR-122 likely acts as an oncogene in T-cell lymphoma. 14 Consistent with this finding, we demonstrated herein that transfection with the miR-122 inhibitor repressed the proliferation, invasion, and migration of both 786-0 and CAKI-1 cell lines, indicating that miR-122 plays an oncogenic role in renal carcinoma cell lines. Together, these data indicate that the biological activity of miR-122 is likely intimately involved in the tumorigenesis and progression of human cancers.
To elucidate the mechanism by which miRNAs function, it is critical to identify targets involved in their regulation. In this study, we used computational analysis to predict that Spry2 was a latent target of miR-122. To support this speculation, we identified a binding site that was complementary to the miR-122 subsequence in the Spry2 3′-UTR. The inhibition of endogenous miR-122 by the miR-122 inhibitor significantly increased the activity of a luciferase reporter gene compared with co-transfection with the NC inhibitor of the wild-type reporter. This increase was not observed with a mutated reporter in either renal carcinoma cell line. Furthermore, overexpression of miR-122 suppressed activity upstream of the wild-type 3′-UTR of Spry2. However, the mutated reporter was not regulated by miR-122. Additionally, transfection of the miR-122 inhibitor promoted the expression of Spry2 at the protein level. Therefore, we concluded that miR-122 regulated and controlled Spry2 expression by directly binding its 3′-UTR.
Emerging evidence implicates sprouty proteins in various cancers.22,25–29 Some sprouty proteins can specifically inhibit the Ras/MAPK signaling pathway and regulate pivotal signaling pathways that are essential to cancer oncogenesis or neoplasia, including angiogenesis, cytokinesis, cell proliferation, invasion, and migration.18,20–22,24,26,31,32 We found that Spry2 expression level was significantly lower in tumors than in adjacent non-cancer tissue samples, supporting its role as a tumor suppressor in RCC. Additionally, we confirmed that the inhibition of miR-122 promoted the expression of Spry2, suggesting that miR-122 may be involved in regulating malignant biological activity. Therefore, we hypothesize that Spry2 knockdown induced by miR-122 facilitates cell proliferation or inhibits cell apoptosis through the abnormal activation of the Ras/MAPK signaling pathway in carcinoma cells. Spry2 hypermethylation has been reported in various cancers, such as prostate cancer, B-cell diffuse lymphomas, and endometrial carcinoma.33–35 Thus, Spry2 methylation could play a role in its low expression. Additionally, both gene methylation and miRNA action may lead to the Spry2 suppression observed in renal cancer tumorigenesis.
MiR-122 has recently been implicated in renal carcinoma due to its modulation of the PI3K/AKT signaling pathway, which supports the oncogenic role of miR-122 in renal neoplasms. 36 Our results are consistent with those of this previous study. The multiple roles of miR-122 can be explained by the fact that miRNAs can modulate various target genes in different signaling pathways in one type of cancer or affect the same target genes in similar signaling pathways in different tumors. Additionally, the correlation between miRNAs and their target genes may reflect the complexity of miRNA function in various tumors. In conclusion, miR-122 may modulate different target genes in RCC, collaboratively contributing to the tumorigenesis of this type of malignancy.
Conclusion
In summary, our study is the first to hypothesize that miR-122 plays an oncogenic role in renal carcinoma cell lines by directly targeting Spry2. However, the carcinogenic effect of miR-122 on the tumorigenesis of renal tumor cell lines observed in this study could be partially attributed to its modulation of Spry2. The purpose of this study was to explore the possible molecular mechanism underlying the role of miR-122 in the progression of renal carcinoma in an attempt to provide new strategies to treat this tumor type. Therefore, miR-122 is a potential diagnostic and prognostic marker of renal cancer and may offer a novel target in the treatment of this neoplasm.
Footnotes
Acknowledgements
Z.W., C.Q., and J.Z. contributed equally to this work and share the first authorship.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the National Natural Science Foundation of China (grant nos 81570676, 81270685); the Science and Education Health Project of Jiangsu Province for Important Talent (grant no. RC2011055); the “333 High Level Talents Project” in Jiangsu Province, China (grant no. BRA2015469, 2011); the Jiangsu Province Six Talents Peak from Department of Human Resources, Social Security Office of Jiangsu Province, China (grant no. 2010WSN-56); the General Program of Department of Health of Jiangsu Province, China (grant no. H2009907); and the Priority Academic Program Development of Jiangsu Higher Education Institutions (grant no. JX10231801).
