Abstract
Objectives
To investigate the allele and genotype frequencies of the warfarin-related genes VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 among healthy Saudis.
Materials and Methods
This cross-sectional study involved 125 unrelated healthy Saudis ages 18–60 years visiting the King Abdulaziz University Hospital (KAUH) in Jeddah, Western Saudi Arabia. The Oragene™ DNA saliva collection kits were used to collect and extract DNA from saliva samples. A polymerase chain reaction-restriction fragment length polymorphism analysis was used to detect the mutant alleles.
Results
Over 51.4% of the Saudi participants carried one or more mutant alleles. The frequency of the VKORC1 (-1639G>A) allele in Saudi was relatively high at 54.8%. The frequencies of the CYP2C9 allele were 19.6% and 54% for the CYP2C9*2 and CYP2C9*3 alleles, respectively, which are substantially more abundant than in other populations.
Conclusion
The observed high frequencies of VKORC1 (-1639G>A) and CYP2C9*2 and CYP2C9*3 polymorphisms suggest that genetic testing should be considered before initiating warfarin therapy to predict the optimal initial dose of warfarin and minimize warfarin-related side effects.
Keywords
Introduction
Warfarin is the most frequently prescribed oral anti-coagulant. It is an effective medication for managing deep-vein thrombosis, pulmonary embolism, and prophylaxis in patients with atrial fibrillation or a prosthetic heart valve replacement. 1 However, warfarin has a higher incidence of iatrogenic complications than other anti-coagulant. This is mainly due to the significant inter-individual variability in dosing requirements, which can result in overdosing or underdosing.2, 3 The selection of the appropriate initial and maintenance doses relies on various patient-related factors, such as body weight, diet, disease states, and concomitant medications. 4 Furthermore, genetic factors are important determinants that further influence warfarin dose variability. 5
Warfarin is a racemic mixture of two enantiomers (R and S), with the (S) isomer having a higher potency than the (R) isomer. Warfarin exerts its anticoagulation effects by inhibiting the vitamin K epoxide reductase (VKORC) enzyme in the liver, thereby preventing the formation of vitamin K necessary to activate several clotting factors. 6 The VKORC enzyme is the rate-limiting enzyme that converts vitamin K-epoxide to vitamin K. The protein of vitamin K-epoxide reductase complex subunit 1 (VKORC1) is encoded by the VKORC1 gene. Several single nucleotide polymorphisms (SNPs) in the VKORC1 gene have been reported. A common SNP of the VKORC1 gene is VKORC1*2 -1639G>A (rs9923231). 7 The 1639G>A polymorphism causes the VKORC1 transcription factor to bind differently, resulting in lower mRNA levels and decreased protein synthesis. 8 As a result, patients with VKORC1 (-1639) AA and (-1639) GA genotypes require lower initial and maintenance warfarin doses than (-1639) patients with GG genotypes. 9
Warfarin is mainly metabolized in the liver by cytochrome P450 (CYP) enzymes. The primary enzyme that metabolizes S-warfarin is CYP2C9, encoded by the CYP2C9 gene. 10 More than 60 variants in the CYP2C9 gene have been reported. 11 A clinically crucial SNP in the CYP2C9 gene occurs at locations 430 and 1075. A change of cytosine with thymine at position 430 (430C>T, rs1799853) is known as the CYP2C9*2 allele, while adenine to cytosine substitution at position 1075 (1075A>C, rs1057910) is known as the CYP2C9*3 allele. 12 The CYP2C9*2 and CYP2C9*3 alleles have lower CYP2C9 enzyme activity than the wild-type allele. As a result, S-warfarin is metabolized more slowly in patients with one or two copies of the CYP2C9*2 or CYP2C9*3 alleles. Patients with the mutant alleles have a higher risk of bleeding and require lower warfarin doses and more time to attain a stable international normalized ratio (INR) than patients with the normal allele CYP2C9*1. Polymorphisms in the VKORC1 and CYP2C9 genes have been reported to be responsible for 35%–50% of the variation in warfarin’s initial and maintenance doses. 13
The frequencies of the VKORC1 and CYP2C9 alleles exhibit significant intra- and inter-ethnic variations. 11 A limited number of studies have been conducted in Saudi Arabia to investigate the frequency of VKORC1(-1639G>A), CYP2C9*2, and CYP2C9*3 alleles in different regions. According to the study by Alzahrani et al. (2013), the prevalence of the VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 alleles were 42.4%, 13.3%, and 2.3%, respectively, among the population of the Eastern region of Saudi Arabia. 14 A cohort study conducted in the Central Region of Saudi Arabia found that the frequencies of polymorphisms were 45%, 11%, and 8% for VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 alleles, respectively. 15 There is currently no research on the prevalence of these genetic variations in the Western Saudi populations. Therefore, this study aimed to investigate the allele and genotype frequencies of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 among healthy Saudi participants visiting King Abdulaziz University Hospital (KAUH) located in Jeddah. Better knowledge of each patient’s VKORC1 (-1639) and CYP2C9 genotypes will assist in choosing their optimal warfarin dose.
Materials and Methods
Patient Selection Criteria and Sample Size Calculation
A cross-sectional study of 125 participants, consisting of 62 females and 63 males, was conducted in Jeddah, the Western Region of Saudi Arabia. Participants were visiting KAUH for routine checkups. The inclusion criteria were Saudi, aged between 18 and 60, healthy, and not on any prescribed medications. The exclusion criteria were participants with a known family history of cancer or mitochondrial DNA-related diseases that may affect the DNA. The study protocol was revised and approved by the KAU Human Research Ethics Committee (076-03-23). Before enrollment, the study objectives were explained to participants, and they were asked to sign a written consent form. Based on the study by Alzahrani et al. (2013), the prevalence of the CYP2C9*3 allele was 13.3% in Saudi Arabia’s Eastern region. 14 Using the open-source calculator OpenEpi, Version 3, we determined that a sample size of 125 subjects would be required to detect the frequencies of CYP2C9*3 alleles in the Western Saudi population with 90% confidence, assuming that the prevalence is in the range of 13.3% ± 5%. 16
Genomic DNA Extraction
Oragene™ DNA Sample Collection Kits (OGR-250, DNA Genotek Inc., Canada) were used to collect around 2 mL of saliva samples from each participant. Genomic DNA was extracted from Oragene DNA/saliva samples using the previously described protocol. 17 Briefly, Oragene DNA/saliva samples were inverted for a few seconds. Then, the samples were incubated overnight at 50°C in an air incubator. The next day, 500 µL of the Oragene DNA/saliva samples were added to new 2-mL microcentrifuge tubes with 20 µL of prepIT®•L2P reagent (DNA Genotek Inc. Canada), vortexed for 5 s, and incubated for 10 min on ice. The samples were centrifuged at room temperature (RT) at 15,000×g for 10 min. Around 300 µL of the supernatants that contain the DNA were transferred to a fresh microcentrifuge tube containing 500 µL of 100% ethanol. The genomic DNAs were allowed to precipitate at RT for 10 min. The samples underwent a 2 min, 15,000×g centrifugation at RT. The DNA pellets were allowed to air-dry for 5 min before being suspended in 100 µL of Tris-EDTA buffer. After extraction, the DNA concentrations were determined using spectrophotometry and stored in a freezer at −20°C.
Genotyping of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Polymorphisms
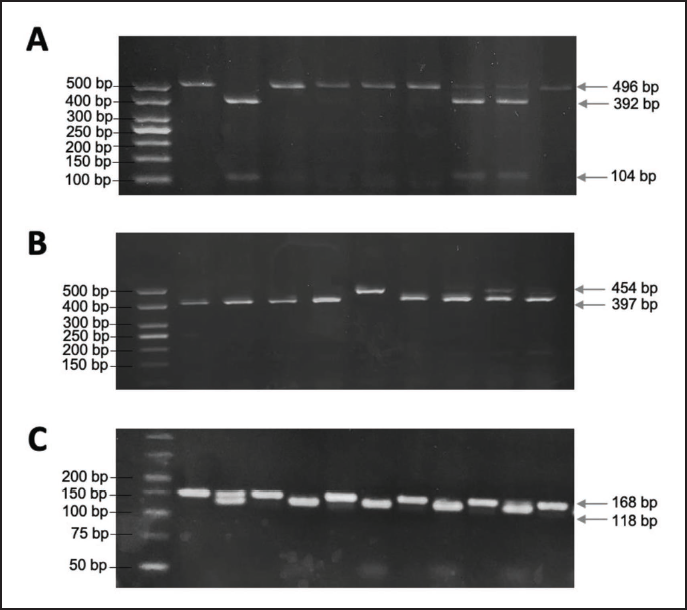
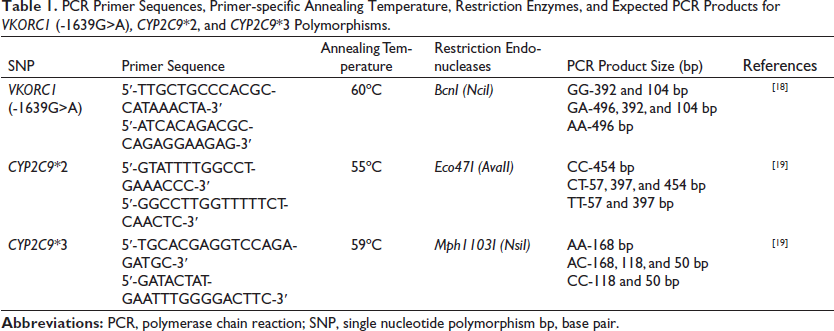
Polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) analysis was used to detect VKORC1 and CYP2C9 genetic polymorphisms, as previously prescribed.18, 19 The PCR reaction included 10 µL of 10X PCR Master Mix (Beijing Solarbio Science & Technology Co., China), 0.8 µL of each primer (0.5 µM) (Macrogen, Korea), 0.8 µL of genomic DNA (100 nM), and nuclease-free PCR-grade water (Jena Bioscience GmbH, Germany) to bring the final volume to 20 µL. The PCR protocol was 95°C for 7 min, followed by 35 cycles of 95°C for 30 s, primer-specific annealing temperature for 30 s, and 72°C for 1 min, followed by 72°C for 3 min. The PCR products (10 µL) were digested overnight at 37°C using a specific restriction endonuclease (Thermo Fisher Scientific, USA) to detect each polymorphism. The digested PCR products were analyzed by agarose gel electrophoresis at 100 V for 90 min using the ENDURO™ Gel XL Electrophoresis System (Labnet International, USA) and visualized using UVP BioDoc-It® Imaging Systems (Analytik Jena, Germany). The representative agarose gel images of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 PCR-RFLP assays were used to interpret the results of the current study (Figure 1). All the PCR primer pairs, primer-specific annealing temperatures, restriction endonucleases, and expected size of DNA fragments for each polymorphism are listed in Table 1.
(A) PCR-Based RFLP Results of VKORC1 Polymorphism. Agarose Gel Electrophoresis for VKORC1 Polymorphism After Digestion with BenI (Ncil). Digestion of the Homozygous GG Genotype Leads to Two Bands of 392 and 104 bp, the Heterozygous GA Genotype Results in Three Bands of 496, 392, and 104 bp, and the Homozygous Mutant AA Genotype Results in a Single Band of 496 bp. Lane 1: 100 bp DNA Marker. (B) PCR-RFLP Results of the CYP2C9*2 Polymorphism. Agarose Gel Electrophoresis for CYP2C9*2 After Digestion with Eco47I (AvaII). Digestion of the Homozygous Wild-Type CC Genotype Yields a Single Band of 454 bp. In Contrast, the Heterozygous CT Genotype Results in Three Bands of 57, 397, and 454 bp, and the Homozygous TT Genotype Shows Two Bands at 57 and 397 bp. Lane 1:100 bp DNA Marker. (C) PCR-based RFLP Results of the CYP2C9*3 Polymorphism. Agarose Gel Electrophoresis for CYP2C9*3 Polymorphism After Digestion with Mph1103I (NsiI). Digestion of the Homozygous AA Genotype Results in a Single Band of 168 bp. The Heterozygous AC Genotype Results in Three Bands of 168, 118, and 50 bp, and the Homozygous CC Genotype Leads in Two Bands of 118 and 50 bp. Lane 1: 50 bp DNA Marker. Abbreviation: PCR-RFLP, polymerase chain reaction restriction fragment length polymorphism.
PCR Primer Sequences, Primer-specific Annealing Temperature, Restriction Enzymes, and Expected PCR Products for VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Polymorphisms.
Data Analysis
Direct counting was used to calculate the genotype and allele frequencies of polymorphisms. The results were then compared to the expected values predicted by the Hardy–Weinberg equilibrium (HWE) using the Chi-Square (χ 2 ) test. For each genotype and allele, the frequencies are provided along with 95% confidence intervals (CI). The CI was calculated using the normal approximation with continuity correction. The χ 2 test compared allelic frequencies between Saudis in the Western area and other populations. p < 0.05 was considered statistically significant.
Results
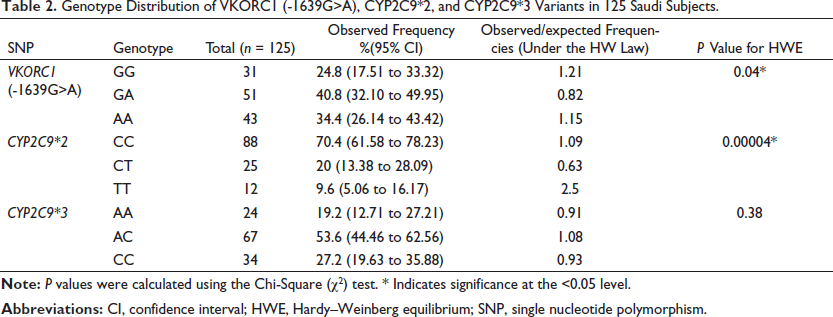
In the current study, allele and genotype frequencies for VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 were investigated in 125 Saudi participants. For VKORC1 (-1639G>A), the genotype distributions were as follows: the homozygous wild-type (GG) genotype was present in 24.8% (95% CI 17.51−33.32) of the sample, whereas the heterozygous genotype (GA) was present in 40.8% (95% CI 32.10−49.95). Thus, the homozygous mutant (AA) compromised the remaining 34.4% (95% CI 26.14−43.42) (Table 2). The frequency of the G allele was 45.2% and the A allele was 54.8%, which was not significantly different (p = 0.14) (Table 3). The genotype distribution for VKORC1 (-1639G>A) did not agree with the HWE (χ 2 = 3.89, p = 0.04) (Table 2).
Genotype Distribution of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Variants in 125 Saudi Subjects.
Allele Frequency of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Variants in 125 Saudi Subjects.
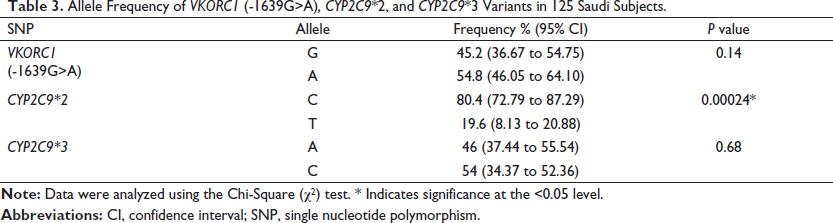
The genotype distributions of CYP2C9*2 and CYP2C9*3 were also investigated. The homozygous CYP2C9*2 wild-type (CC) was observed in 70.4% (95% CI 61.58−78.23) of the subjects, while the heterogeneous CYP2C9*2 (CT) was observed in 20% (CI 13.38−28.09) of the subjects. Hence, the homozygous mutant CYP2C9*2 (TT) genotype represented the remaining 9.6% (95% CI 5.06−16.17) (Table 2). The frequency of the C allele was present in 80.4%, which is significantly higher (p = 0.00024*) than the T allele (19.6%) (Table 3). The CYP2C9*2 genotype distribution did not agree with the expected distribution based on an HWE (χ 2 = 16.69, p = 0.00004) (Table 2). The CYP2C9*3 (AA) was present in 19.2% (95% CI 12.71−27.21) of the subjects, whereas the heterogeneous CYP2C9*3 (AC) was present in 53.6% (CI 44.46−62.56), and the homozygous mutant CYP2C9*3 (CC) genotype was present in 27.2% (95% CI 19.63−35.88) of the subjects (Table 2). There was no statistically significant difference (p = 0.68) between the A allele (46%) and the C allele (54%) (Table 3). This genotype distribution for the CYP2C9*3 polymorphism was consistent with the HWE (χ 2 = 0.77, p = 0.38) (Table 2).
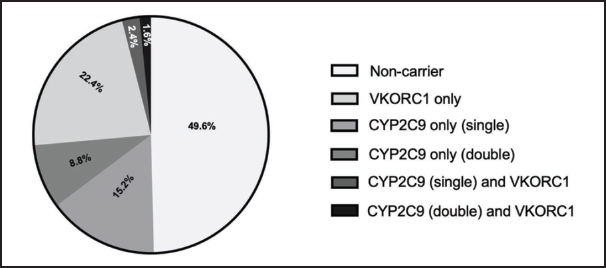
Next, we investigated the combination of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 polymorphisms (Figure 2). Individuals with no polymorphism in either gene (noncarriers) composed 49.6% of the tested population. The percentages of subjects with a single polymorphism in only one gene were 22.4% for VKORC1 and 15.2% for CYP2C9. Double-carrier individuals with two polymorphisms on the CYP2C9 genes composed 8.8% of the tested population. Double-carrier individuals with single polymorphisms on the CYP2C9 and VKORC1 genes accounted for 2.4% of the tested samples. Triple-carrier individuals with two polymorphisms in the CYP2C9 gene and a single polymorphism in VKORC1 accounted for only 1.6%.
Carrier Prevalence of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Polymorphisms Among the Saudi Population.
Discussion
The present study aimed to assess the allele and genotype frequencies of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 polymorphisms in healthy Saudi residents of the Western region of Saudi Arabia using the PCR-RFLP technique. The frequencies of the VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 alleles in Saudi were 54.8%, 19.6%, and 54%, respectively. Over 51.4% of the Saudi population carries one or more risk alleles, potentially at increased risk of developing warfarin-related side effects.
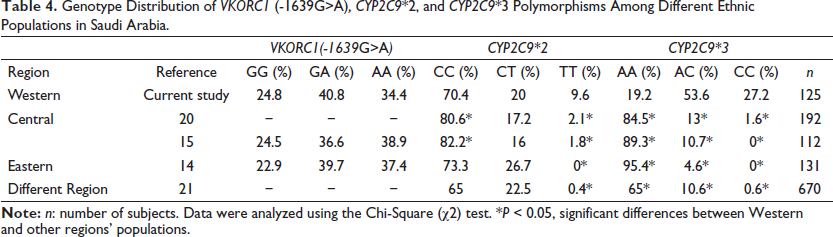
Several studies have previously reported the prevalence of the VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 polymorphisms among different regions in Saudi Arabia, but this is the first study to report them in the Western Province (Table 4).14, 15, 20, 21 Alzahrani et al. and Al-Saikhan reported that the homozygous mutant VKORC1 (-1639G>A) genotype was present in 37.4% and 38.9% of participants in the Eastern and Central Populations of Saudi Arabia, respectively.14, 15 Our data from the Western Province are consistent with previous research conducted in the Central and Eastern provinces (Table 4). Furthermore, we found that the CYP2C9*2 homozygous mutant (TT) and CYP2C9*3 homozygous mutant (CC) alleles were present in 9.6% and 27.2% of the participants, respectively, which are significantly higher than the percentages in other regions in Saudi Arabia (p < 0.05) and even other populations (Table 4). This indicates a notable difference in the prevalence of the CYP2C9*2 and CYP2C9*3 genotypes between the Western and other provinces of Saudi Arabia. These could include differences in genetic composition and ancestry among the populations residing in these regions and variations in environmental and lifestyle factors that can influence gene expression and allele frequencies.
Genotype Distribution of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Polymorphisms Among Different Ethnic Populations in Saudi Arabia.
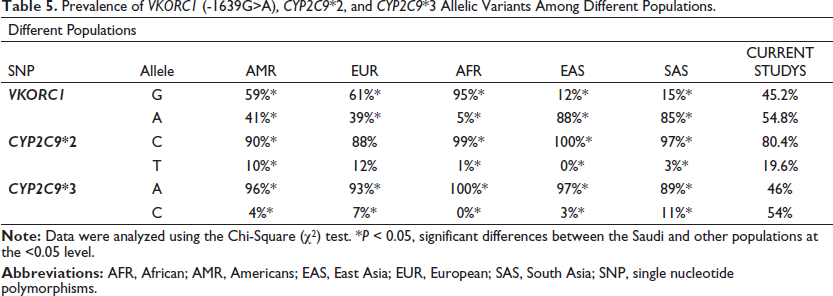
The allele frequency of the VKORC1 (-1639G>A) polymorphism was previously evaluated in different populations. It is well established that the mutant A allele frequencies can vary wildly depending on the population ancestry being studied.11, 22 The lowest A allele frequency observed in any population was 5% in the African populations, according to the 1000 Genome Phase 3 Project (Table 5; Figure 3A). 23 In contrast, the highest A allele frequencies were reported at 88% and 85% in the Eastern and Southern Asian populations, respectively. The current study found that the A allele was present in only 54.8% of the selected Saudi population, which is significantly lower than in other Asian populations. Saudi Arabia is located in Southwestern Asia. Emigration and potential genetic admixture between the Saudi and African or Asian populations could explain our observation. Interestingly, the frequency of the VKORC1 (-1639G>A) allele among Saudis is similar to other Arabian populations, ranging between 52% and 56%.24, 25
Prevalence of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 Allelic Variants Among Different Populations.
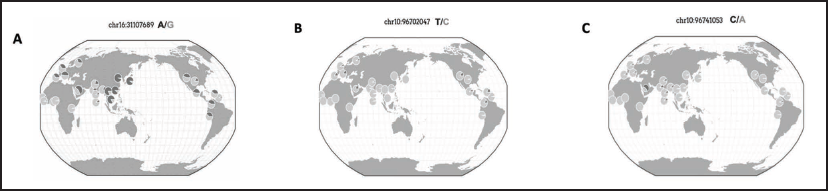
Regarding the CYP2C9*2 polymorphism, previous research found that Americans and Europeans had the highest prevalence of the mutant T allele, at 10% and 12%, respectively. The prevalence was also high across the Middle Eastern population. The highest prevalence was in Lebanon (11.2%), followed by Iran (10%). The CYP2C9*2 allele is absent in East Asian populations, whereas the frequency in the South Asian population was only 3% (Table 5; Figure 3B). We found that the T allele was present in 19.6% of the Saudi population examined, which is significantly higher than in other populations. 23
Considering CYP2C9*3 polymorphism, Europe features high CYP2C9*3 frequencies (11%). However, the allele was rare in East Asia (3%) and absent in Africa. Contrary to CYP2C9*2, the CYP2C9*3 allele was highly prevalent in South Asia, with frequencies as high as 11.9% in Pakistan and 11.6% in Bangladesh. In the current study, the mutant C allele for CYP2C9*3 was observed in 54% of the tested Saudi participants, which is substantially higher than that reported for different populations (ranges from 0% to 11%) (Table 5; Figure 3C). In Arabian populations, interethnic variation in the frequency of the CYP2C9*2 and CYP2C9*3 alleles ranges from 7% to 21% and 3% to 9%, respectively.24, 26, 27 Interestingly, according to a recent study, the United Arab Emirates had very high CYP2C9*3 frequencies (21.3%) compared to other Middle Eastern populations. Data on the genetic variations of CYP2C9 for many populations are still lacking, indicating that future research in these ethnographic groups is still crucial.
The most severe side effect of warfarin is bleeding, which has been linked to several clinical factors, including VKORC1 and CYP2C9 polymorphisms. 28 The Food and Drug Administration authorized warfarin label alterations with dose recommendations based on VKORC1 and CYP2C9 genotypes field (Teva Pharmaceuticals USA) to reduce warfarin-related adverse effects. 29 VKORC1 -1173C>T, CYP2C9*2, and CYP2C9*3 have been reported to be responsible for 35%–50% of the variation in warfarin’s initial and maintenance doses across several populations. 5 Studies showed that genetic-based warfarin dosing upon initiation improves target INR attainment during the initial month. A recent Holail et al. study reported that the warfarin dosage in Saudi patients was associated with polymorphisms in the VKORC1 and CYP2C9 genes. 30 Patients who carry at least one copy of the mutant alleles for VKORC1 -1173C>T, CYP2C9*2, and CYP2C9*3 require a significantly lower warfarin maintenance dose. 30
This is the first study to evaluate allelic variants in the warfarin-related genes VKORC1 and CYP2C9 in a Western Saudi population. However, some limitations must be addressed. A limitation of the study is that the sample size calculation was based only on the data for CYP2C9*3 and did not consider VKORC1 (-1639G>A) when estimating the prevalence of the alleles in the Western Saudi population. This may have led to underestimating or overestimating the true prevalence of the VKORC1 (-1639G>A) allele. Therefore, the study results may not represent the true prevalence of the VKORC1 (-1639G>A) allele in the selected population. Future studies with larger sample sizes are required to address this limitation and provide a more accurate estimate of the prevalence of the allele. Furthermore, to implement warfarin pharmacogenomics in clinical practice in Saudi Arabia, the prevalence of VKORC1 and CYP2C9 genetic variations in other regions of Saudi Arabia must be determined to aid clinical decision-making and enhance patient treatment.
We observed that the genotype distribution for VKORC1 (-1639G>A) and CYP2C9*2 did not adhere to the principle of HWE. When a population is not in HWE, it suggests that there may be non-random mating, genetic drift, selection, mutation, or other factors affecting the allele frequencies. These factors can lead to an excess or deficit of specific genotypes, affecting the statistical power and accuracy of association tests. Furthermore, it is recommended that the frequencies of different CYP2C9 alleles, particularly those variations known to be related to a reduction in the CYP2C9 enzyme activity, such as CYP2C9*5, *8, *11, *13–18, *30, and *33, should be investigated in Western and other Saudi populations.
Given the high prevalence of VKORC1 (-1639G>A), CYP2C9*2, and CYP2C9*3 alleles, these variants might be the most crucial factor of warfarin pharmacogenomics in the Saudi population. Therefore, genetic testing should be conducted to detect any potential allelic variants that may affect the warfarin response. Genetic counseling should also be provided for patients who possess one or more of these polymorphic variants so that they understand their implications and can make informed decisions regarding their anti-coagulant medication options.
Authors Contributions
Conceptualization: AMB and RHH; Methodology: AMB, WSA, and LSG; Writing original draft: AMB, WSA, and LSG; Writing review and editing: AMB, LSB, and RHH; Supervision: AMB and RHH. All authors have critically reviewed and approved the final draft and are responsible for the content and similarity index of the manuscript.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
The study protocol was revised and approved by the KAU Human Research Ethics Committee (076-03-23).
Informed Consent
The participant has consented to the submission of the article to the journal.
Funding
The author received no financial support for the research, authorship and/or publication of this article.
