Abstract
Effective and affordable treatment for rare diseases remains a huge challenge. Only a few drugs have made inroads into the therapeutic arena for rare diseases. The current focus is shifted to different uses for expensive new drugs, referred to as drug repurposing. It is also called drug repositioning or reprofiling, refers to the identification of new therapeutic applications of existing or investigational drugs. This concept and novel approaches have promising role in the treatment of many potentially incurable diseases, particularly certain forms of cancer, neurodegenerative conditions, immunological disorders and uncommon infectious diseases. There are huge expectations filled with optimism that the drug repurposing would offer fresh opportunities for the treatment and management of rare diseases. This is undoubtedly more relevant for low and middle income countries.
Keywords
Introduction
With increasing successes and sophistication in the genomic and multi-OMIC diagnosis of rare diseases, the focus is now on targeted and precision therapy. Tremendous success has been achieved in new drug discovery and drug development for rare or so-called neglected orphan diseases (Drugs for Neglected Diseases initiative;
Selected Examples of Drug Repurposing in Rare Diseases
Most rare diseases have complex molecular mechanisms involving different genes and peptide molecules, collectively referred to as ‘gene-molecular pathway complex’, such as the RAS-MAPK pathway, mTOR signalling pathway, TGF-beta signalling pathway and many more. It is thus likely that a drug or chemical that modifies a specific part of a pathway may also influence the action or expression of other molecules in the same pathway. This is evident from the side effects associated with several new drugs. This concept is capitalised in drug purposing research leading to few successful examples (
Nitisinone for Alkaptonuria
Historically, alkaptonuria (OMIM#203500) laid the foundation of inherited metabolic diseases coined by the pioneer Oxford physician Professor Sir Archibald Gerrard who made simple observation of dark-coloured urine in patients presenting with skin pigmentation and chronic knee pain (ochronosis) and linked it to the deficiency of enzyme required for the breakdown of homogentisic acid. This rare disease is closely linked with disorders of tyrosine and phenylalanine metabolism. Mutations in the homogentisic acid oxidase (HGD) gene result in the accumulation of homogentisic acid with clinical manifestations of alkaptonuria [Figure 1].
Deficient activity of homogentisic acid oxidase (HGD) in alkaptonuria
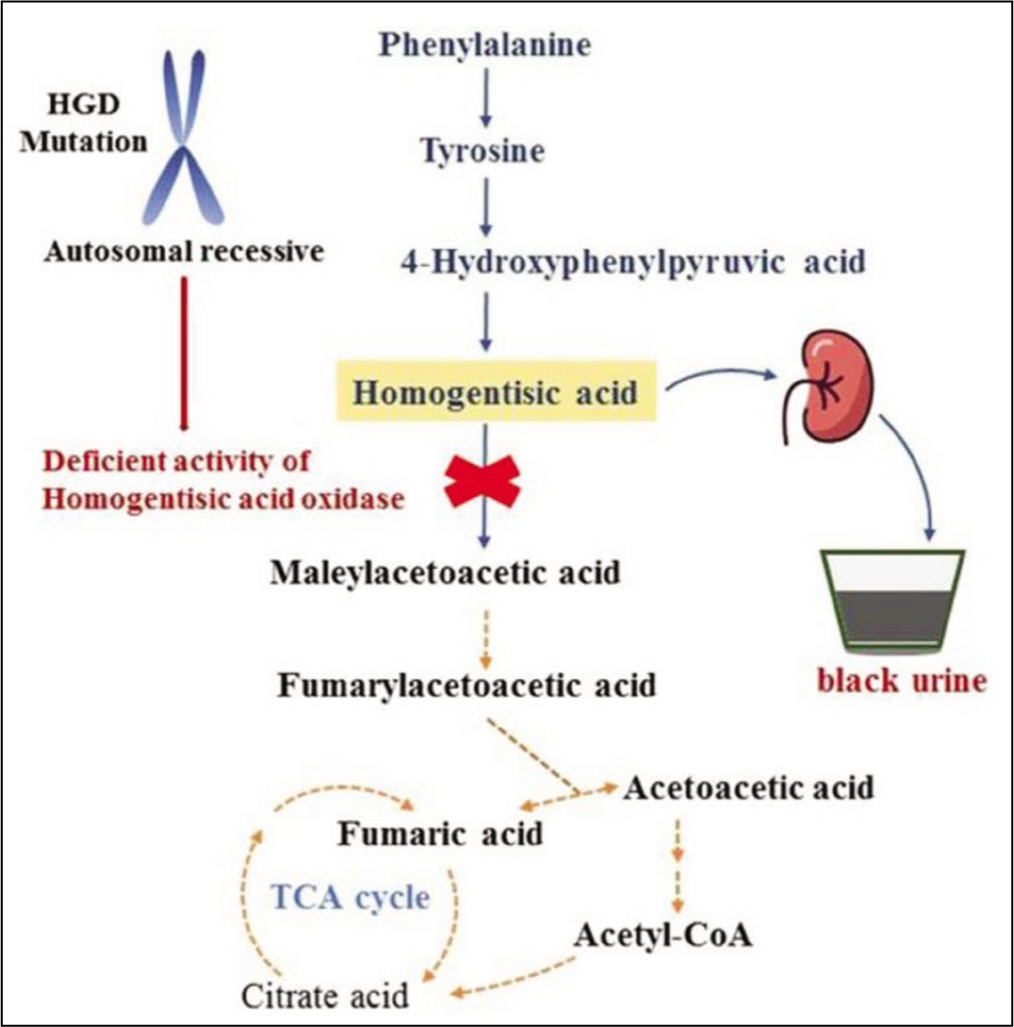
A new drug, nitisinone, a known weed killer, is licensed for the treatment of tyrosinemia type 1. This drug was later used for alkaptonuria. However, clinical trials in the USA were unsuccessful. Later, the UK-based Alkaptonuria Society conducted a large international clinical trial and generated positive data leading to the licensing of nitisinone for the treatment of alkaptonuria. Nitisinone treatment-induced changes in biomarkers of bone and cartilage remodelling (collagen type I and II). These biomarkers can aid patient management and deepen our knowledge of the molecular mechanisms of this rare disease.[5,6]
mTOR Inhibitor (Everolimus) for Tuberous Sclerosis
Many different protein kinase (PK) molecules are associated with a number of clinically heterogeneous rare disorders including cancer family syndromes. Most PKs are involved with cell proliferation, autophagy, and apoptosis through participation in multiple signalling pathways. Molecular mechanisms of different PKs are closely interlinked with the final endpoint of mTOR (Mammalian target of rapamycin) signalling pathway receptors (mTORC1 and mTORC2) [Figure 2]. There is overwhelming evidence for mTOR signalling pathway associated with cancer, arthritis, insulin resistance, osteoporosis and other common/complex diseases.[7] Several clinical studies have focussed search for mTOR inhibitors for anti-tumour therapy. Results of clinical trials with newly discovered mTOR inhibitors (Everolimus and sirolimus) have shown remarkable benefits in the management of Tuberous Sclerosis Complex (TSC).[8] The TSC includes two heterogeneous autosomal dominant disorders (OMIM 191100 TSC1; 191092 TSC2) manifesting with characteristic skin, eye, brain, neurological features and tumour development in the brain, eyes, heart, lungs and kidneys. Everolimus and other related drugs are now approved by FDA and other drug licensing authorities. The multi-disciplinary management for both forms of TSC involves input from several specialists. Further clinical studies are underway for broader usage of RAS-MAPK-mTOR inhibitors in the treatment of RAS-MAPK disorders (Rasopathies).[9]
RAS/MAPK/mTOR signalling pathway
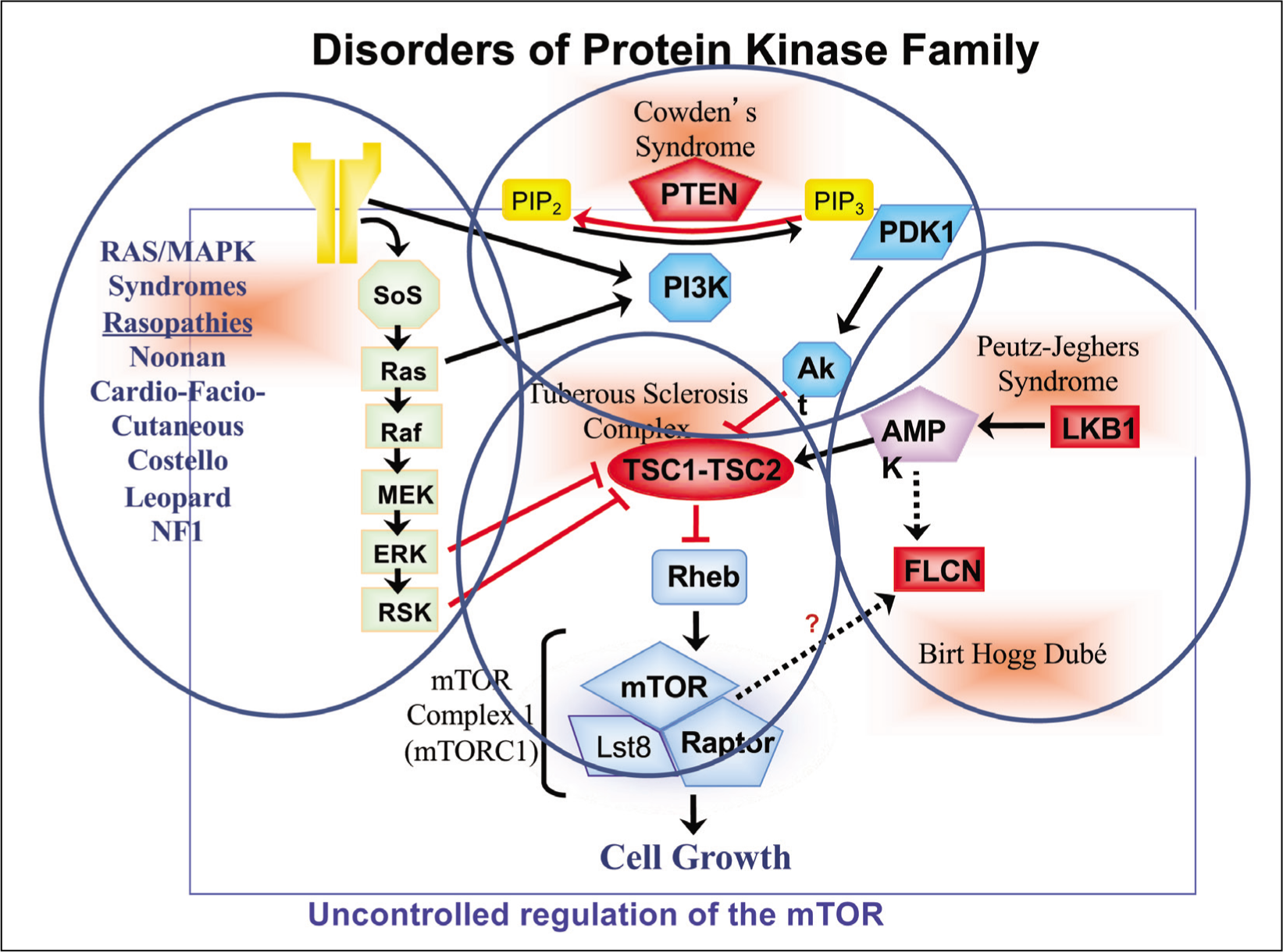
mTOR Inhibitor (Sirolimus) for Autoimmune Lymphoproliferative Syndrome
The autoimmune lymphoproliferative syndrome (ALPS; OMIM#601859), also known as the Canale-Smith syndrome, is one of the rare primary immune deficiency disorders, caused by heterozygous pathogenic variants in the FAS or FAS ligand genes (ALPS-FAS/ALPS-FASLG). Major clinical manifestations include enlarged lymph nodes (lymphadenopathy) and spleen (splenomegaly), as well as decreases in red blood cells, platelets, and, occasionally, neutrophils, as well as an increased lifetime risk of lymphoma.[10,11] It is estimated that around 1,000 individuals are diagnosed with ALPS-FAS. Inheritance is autosomal dominant with incomplete penetrance and wide clinical variation. A rare autosomal recessive form of ALPS-FAS is also described in a girl, born to consanguineous parents, with homozygous 20-bp duplication in the FAS gene.[12]
The current treatment for ALPS-FAS is limited to primary disease characteristics and related clinical complications. Different models of treatment are used solely based on immunosuppression. Few clinical studies have shown positive results in ALPS-FAS patients treated with sirolimus, one of licensed the mTOR inhibitors. A recent meta-analysis of clinical trials on the use of sirolimus in ALPS-FAS reported almost complete responses in around 50% or more patients. It is now considered to be safe with few side effects, thus supporting the use of Sirolimus in the treatment of ALPS.[13]
Angiotensin Receptor Blocker (Losartan/Irbesartan) for TGF-beta Signalling Disorders
Following the identification of pathogenic variants in the microfibrilin gene in Marfan syndrome [OMIM# 154700], it became apparent the fundamental molecular mechanism involved the activation of TGFß. Further studies in patients with Marfan-like features led to distinct phenotypes in children with additional clinical features described as Loeys-Dietz syndromes [LDS1 OMIM# 609192 & LDS2 OMIM#610168]. Molecular studies in these patients led to the identification of heterozygous mutations in the TGFß receptors (TGFBR1 &. TGFBR2).[14,15] In addition to mutations in TGFBR1 and TGFBR2, other genes in the TGFß signalling pathway are also associated with LDS-like phenotypes [Figure 3].
TGFß signalling pathway and cardiovascular phenotypes
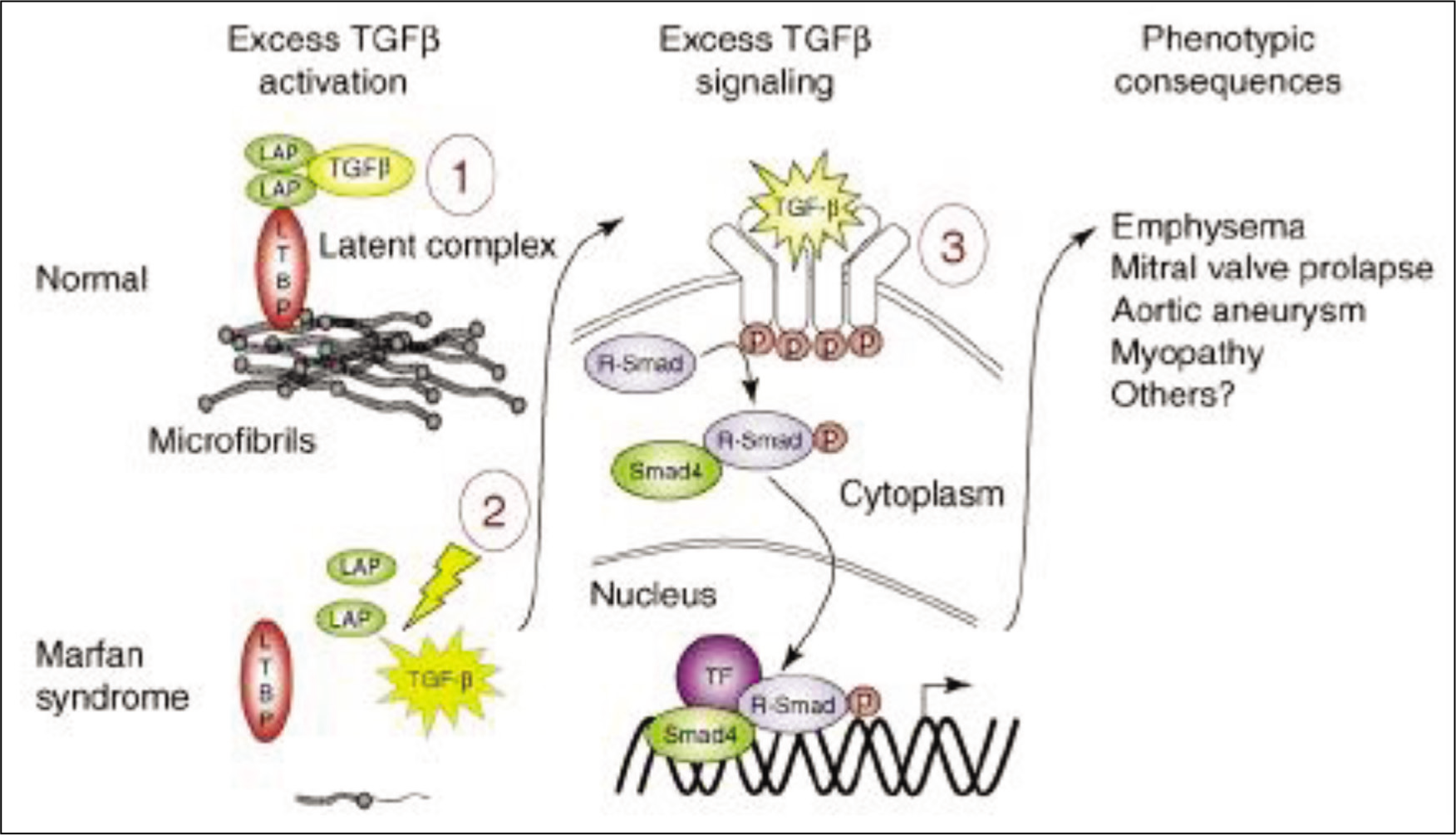
The role of TGFß and other related genes in cardiovascular development and function has been considered for some time.[16] Clinical delineation of Loeys-Dietz syndrome and other clinically similar rare phenotypes have led to firm evidence of the TGFß signalling pathway associated with abnormal aortic and arterial proliferation manifesting with dilatation and/or dissection in both large to medium-sized arteries. Animal studies have demonstrated close links of TGFß signalling with angiotensin II based on the response of losartan administration, one of the angiotensin II receptor blockers, in LDS mice models.[17] Based on the outcomes of different clinical trials, there is a major shift in the therapeutic use of angiotensin-converting enzyme II receptor blockers (losartan, candesartan, irbesartan and others) for the progression and prevention of aortic or arterial dilatation/dissection in Marfan/LDS like disorders.[18] In addition, the use of beta blockers remains in clinical practice supporting the use of combined use of beta blockers and angiotensin II receptor blockers. However, there is no convincing evidence. In conclusion, most angiotensin II blocker drugs offer additional therapeutic advantages in managing aortic and arterial complications and remain potent anti-hypertensive and widely prescribed in most patients with chronic left heart dysfunction.
Machine Learning and Artificial Intelligence for Drug Repurposing
Recent publications on drug repurposing or repositioning include several articles and research articles on the scope and applications of machine learning (ML) and artificial intelligence (AI).[2] Applications of advanced computational methods like ML and AI have facilitated investigators to browse through huge datasets, identify patterns and generate hypothetical models for drug repurposing. The conventional in silico methods are most popular for selecting specific targets, the drug in question and disease phenotypes [Figure 4].[19] Essentially, three fundamental approaches include the network(s), molecular target classification, and matrix factorisation. Details on methodology are outside the scope of this article. By far the best example is the KUALA framework that allows identification of kinase active ligands and prioritising multi-target molecules for drug repurposing.[20] The KUALA project employs 12 different ML methods for classification to assign closely aligned kinase inhibitors in clinical trials in around 80% of cases. This approach has received wide acceptance in the drug repurposing research for drug repositioning on analogous targets.
Selection of specific targets in drug repurposing[19]
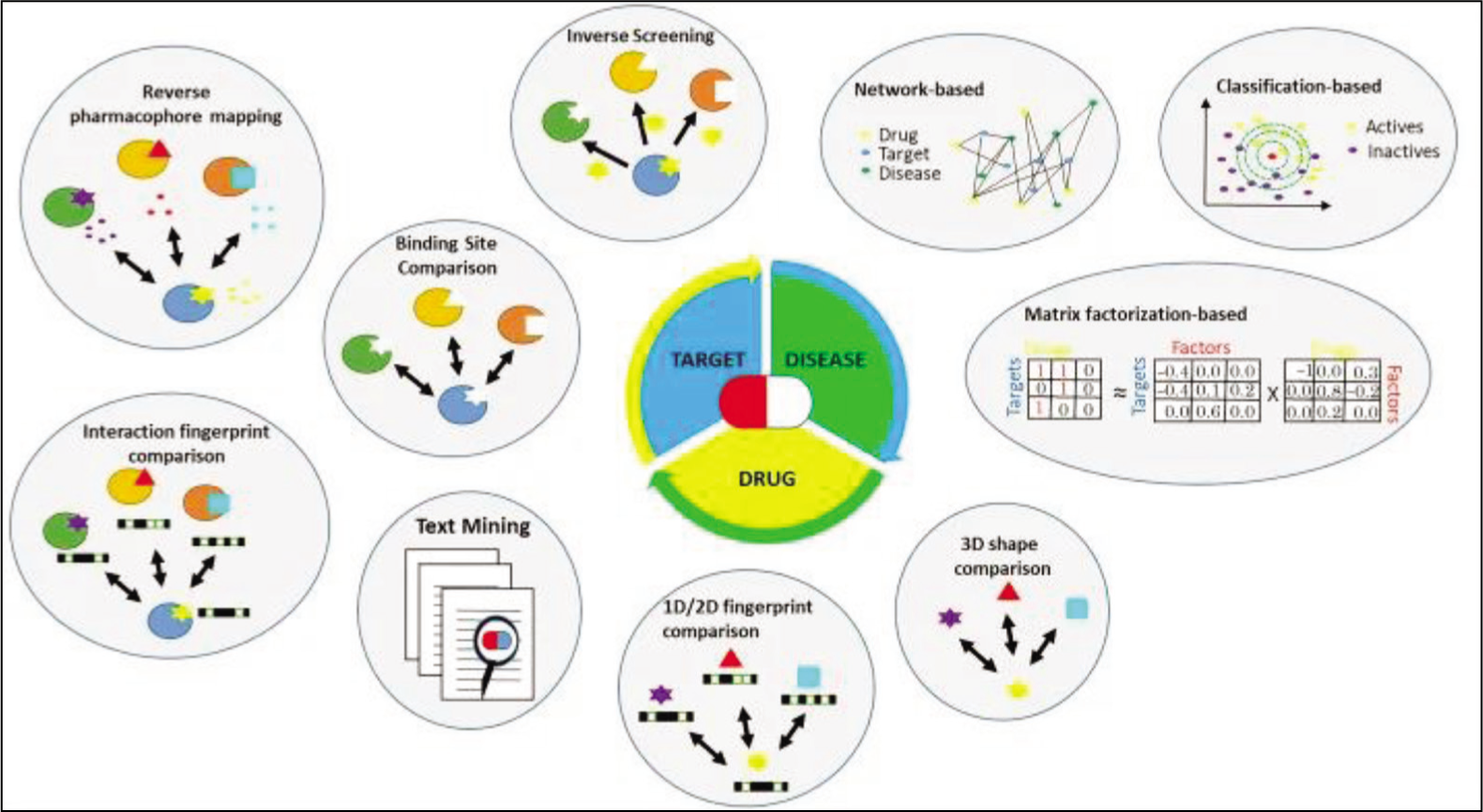
Development of a Drug Repurposing Project
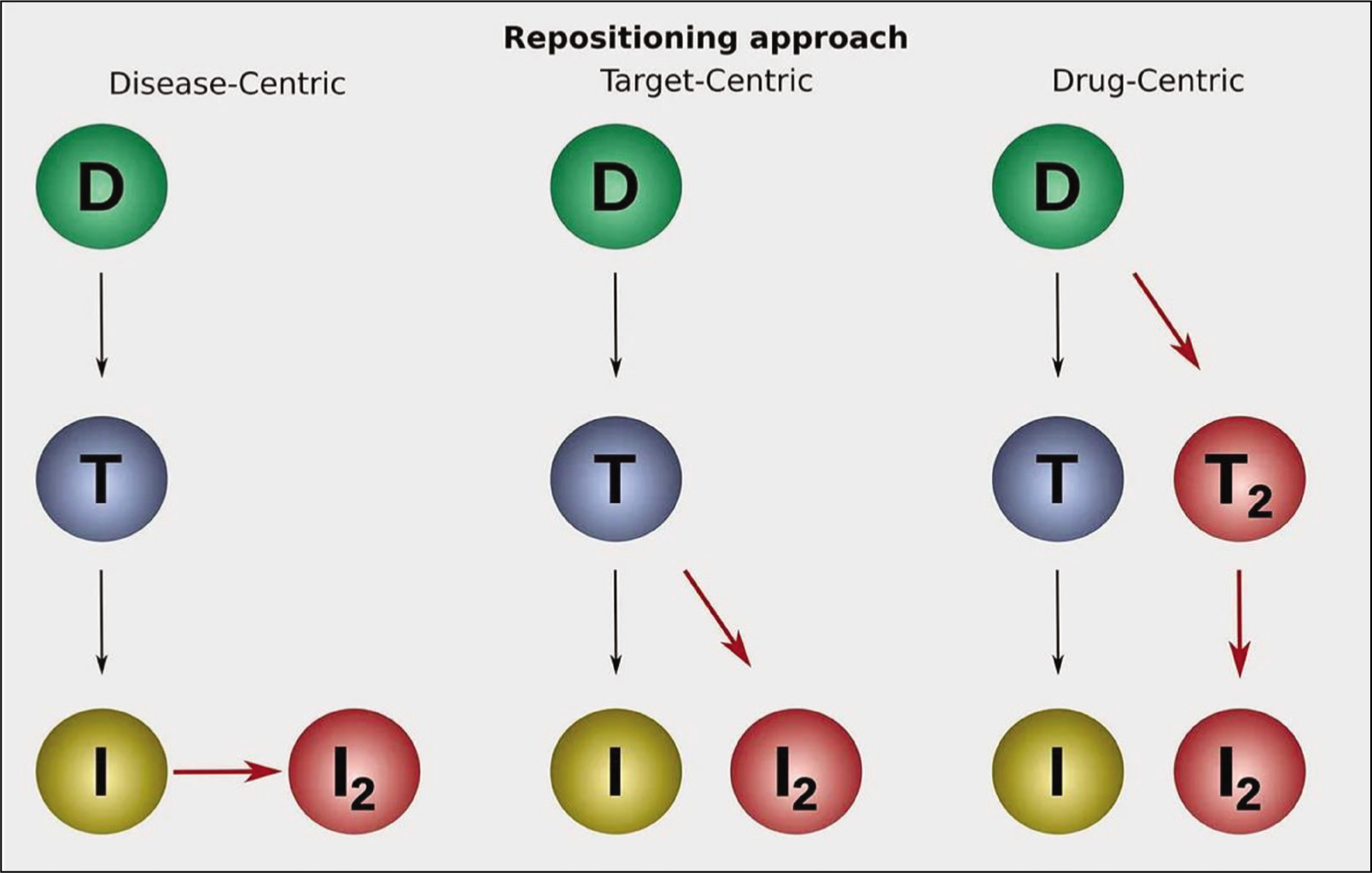
There are three approaches for drug repurposing or repositioning research—drug-centric, disease-centric and target-centric [Figure 5].[21] Most pharmaceutical organisations follow the drug-centric approach for the main reason of achieving best possible efficacy and highest financial gains. On the other hand, a clinician would be more inclined on drug usage in a large number of diseases that share molecular mechanisms, such as inflammation, autoimmune and cancer. All approaches assist in the identification of novel targets and thus setting out new indications.
Recommended requirements for drug repurposing or repositioning:
Identify molecular target or pathway implicated in the disease of interest. Select existing drugs that target or pathway in the disease of interest. Quantify evidence for the drug repurposing candidate for improved efficacy over existing treatment options. Prepare clinical evidence to support the use of the drug in a new context. Probable potential safety and toxicity risks, and side effects and adverse events. Revised dosage, preparation and duration of treatment. Assess sufficient patient population and market size for viable options. Evaluate IP protection or current patents for the selected drug. Understand the regulatory requirements for the candidate drug. Financial requirements for manufacturing and marketing of the candidate drug.
Resources for drug repurposing or repositioning project:
Disease pathology, omics data, scientific literature Clinical trial data, electronic health records Drug compound databases Drug safety data, for example, federal adverse event reporting system (FAERS) Pharmacology data, including pharmacokinetic (PK) data, metropolitan police drug data (MET data), and drug interactions Patents Regulatory documents
Recommended Methodology for Drug Repurposing
Current methods for drug repurposing or drug repositioning are essentially driven by large data and computational analysis. This is employed by most pharmaceutical companies, as well as, academic clinicians or investigators. Essentially, following methods are in use-
Build a disease model pathway to summarise and dissect the underlying biology of the disease. Create a knowledge graph to include different biological concepts (genes, protein molecules, interlinked molecules, molecular mechanisms, and the cell process) for precise understanding and selection of new targets.
A disease biology knowledge graph might include-
Gene expression: expression, promotor binding, miRNA effect. Proteomics/physical interaction: direct regulation, protein modification, binding. Biomarkers: biomarker, genetic change, quantitative change, state change. Metabolomic/molecular transport and modification: molecular synthesis, molecular transport, chemical reaction. Functional association: between disease and a cellular process or another disease. Regulation: a specific type of relationship is used if no more specific information is available.
In addition to the above basic requirements, it would be necessary to build a graph neural network, screen compound libraries, and undertake literature and text mining.[22]
Conclusions
Undoubtedly, drug repurposing or repositioning is the way forward to achieve highest possible efficacy and efficiency of expensive new drugs in the treatment of rare diseases. There are few drugs that are now approved for alternative or additional indications compared to an original licensing agreement. However, there are several barriers in achieving the desired goals of this major initiative.[22] There is an urgent need for multi-partner collaborations, availability and use of compound databases, and necessary financial incentives. Needless to emphasise, affordable and efficacious use of existing drugs repurposing would make a huge impact on the treatment of rare diseases in less-resourced LMICs.
Footnotes
Acknowledgements
This article is based on personal research using the available online resources and recent publications. The content and illustrations are used for educational purposes only. There are no anticipated financial or professional incentives to the author. The manuscript was gratefully reviewed by to Dr. Ambika Gupta, Consultant Clinical Geneticist, Apollo Genomics Institute (Delhi and Ahmedabad), Apollo Group of Hospitals, India.
Declaration of conflicting interests
The author declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The author received no financial support for the research, authorship and/or publication of this article.
Informed consent
Consent was not needed/not applicable, as this is a review article compiled from various research articles and guidelines and not from patients directly.
Credit author statement
All words regarding this article were done by the author.
Data availability
PubMed and Google Scholar.
Use of artificial intelligence
This analysis was conducted entirely by human effort, without incorporating artificial intelligence or automated tools.
