Abstract
Autoimmune rheumatic diseases are often characterised by heterogeneity in presentation. The traditional approach to diseases guided by their phenotype may be suboptimal with the advent of precision medicine. Precision medicine is the integration and application of multiomics to predict the best-performing drug and its toxicity profile to derive optimal benefits. With novel drug discoveries and an expanding therapeutic armamentarium, it potentially aids in clinical and therapeutic decision-making, while saving time and averting adverse events. However, multiomics comes with ‘big data’, and owing to the costs, the sample size is usually small. Machine learning (ML) plays an important role in these scenarios where conventional statistics fall short. So, by integrating clinical data with the data from -omics, ML models can be built, which can accurately predict the clinical factors or even novel biomarkers that predict response. This approach has a potential for great benefit as valuable time or the ‘therapeutic window of opportunity’ would be saved, with fewer adverse events, eventually translating to lower damage accrual and better outcomes. Most of the evidence for the use of ML in precision rheumatology comes from rheumatoid arthritis and the factors predicting response to various drugs, including tumour necrosis factor inhibitors. This approach also has its limitations such as the lack of generalizability and the current scarcity of longitudinal data. These models must be tested in larger cohorts and population-based studies for validation, failing which there is a risk of apparent identification of multiple ‘novel’ biomarkers that may or may not be mechanistic.
Introduction
The traditional approach to diagnosis and treatment of autoimmune rheumatic diseases (AIRDs) has been phenotype-based. With the advent of multiomics, a paradigm shift has been anticipated. Precision medicine stems from this integrated ‘systems biology’ approach, where the -omics data of the patients is integrated to predict the best-performing, tailored therapeutics and in addition, anticipate toxicity, to achieve optimal outcomes. 1 However, the big data from multiomics present a challenge of their own, stretching the limits of conventional statistics. Thus, there is a perfect niche for machine learning (ML) to contribute to precision medicine.
There is a plethora of symptoms for various AIRDs. Though standard guidelines exist, at many places the choice of drugs is left to the treating physician. Owing to the heterogeneity of the disease manifestations and the presence of comorbidities, the response to a particular treatment may not always be predictable. 2 This leads to a ‘trial-and-error’ approach in clinical practice, during which the therapeutic window of opportunity may be missed, eventually leading to damage accrual owing to suboptimal disease control. 3 The therapeutic armamentarium is gradually expanding with addition of newer drugs with novel targets. Another issue may be a low therapeutic index of some drugs, where the clinician has to decide the individual risks before prescribing.
In the background of these issues, precision medicine has gained traction in the last decade. Though the inception of precision medicine has been mostly in the field of oncology, there is growing evidence in immunology and rheumatology.
Artificial intelligence has immense potential in healthcare and one of the major tools, the ML models, has been widely applied in many scenarios such as prediction of disease outcomes, treatment responses, prediction of disease inception, and analysis of big data, among others. ML, in simple terms, gives a system the ability to learn from experience, without being systematically programmed to do so. There are four major types of ML models – supervised, semi-supervised, unsupervised, and reinforcement learning models, with supervised models having found most use in healthcare. 4 While traditional statistical analyses merely infer relationships between variables, ML models aim to make accurate predictions. 5 So, by integrating the systems biology approach, combining different –omics data with supported bioinformatics analysis from ML models, we can predict, treat and monitor these diseases, translating to better clinical outcomes and lower overall damage accrual.
Cracking the ‘code’ of the right biomarker, the right genome, and the right drug that will have the best effect on a patient, will refute any ambiguity in choosing treatment, and also avoid the multiple trials needed to find the ‘right fit’ to obtain optimal response. Traditional statistical analysis may fall short in analysing the big data required for the robust implementation of precision or personalised medicine. ML is a great leap for science in this area, as it can accurately predict the response to treatment while integrating the other available data. This would also lead to significantly conserving resources and avoiding the hassle of frequent testing for adverse events of a particular medication while trying it out on a patient.
In this review, we aim to explore the applications of ML models in the prediction of response to treatment in rheumatic diseases, which may potentially aid in identifying novel biomarkers and traditional risk factors. In addition, it can possibly be applied in conceptualising and building more robust algorithms and formulating guidelines for better application and integration of technological prowess to bridge the gap in achieving complete remission.
Search Strategy
We conducted a thorough search of PubMed/MEDLINE, WebOfScience and Scopus with the medical search heading (MeSH) terms ‘machine learning’ AND ‘arthritis, rheumatoid’ OR ‘lupus erythematosus, systemic’ OR ‘spondylarthritis’ OR ‘vasculitis’ OR ‘Sjogren’s syndrome’ OR ‘scleroderma, systemic’ OR ‘reactive arthritis’ OR ‘vasculitis’ in various combinations. We also included the relevant articles that were cross-referenced from these. We did not specify a time period for article inclusion. We included only the papers published in English. Conference abstracts were excluded. The final review was written adhering to the guidelines and the standard framework to write a narrative review. 6
Discussion
ML is emerging as an apparently accurate tool for an expansive range of applications in healthcare and rheumatology. Most of the research and evidence for the applications of ML models in precision medicine have emerged from rheumatoid arthritis (RA), with recent evidence surfacing from systemic lupus erythematosus (SLE) and spondyloarthritis (SpA) too. Though ML models abrogate the issues with analysis faced due to small sample sizes in the traditional statistical methods to some extent, they are plagued by other problems like dimensionality. This is of value in precision medicine while exploring big data from -omics in the presence of a large number of variables, which also results in higher redundancy. 7 But owing to the costs and inadequate funding, especially in developing countries like India, obtaining larger samples may not be feasible. In such scenarios, multiple variables may be checked to build a model with the best-performing set of variables. 8 While most treatment is governed by standard guidelines, it may not be the most optimal. With precision medicine and integration of ML to identify the right variables or the biomarkers to predict response to a particular drug, there is improved potential to optimise therapy, minimise toxicity and translate to better clinical outcomes.
While supervised ML models have found the most use in the prediction of response to therapy and in precision medicine, some studies have also explored semi-supervised models in treatment decisions in RA.9,10 Depending on clinical utility, baseline data availability and its heterogeneity, and the outcome expected the decision to use either can be made. One study by Morid et al. demonstrated that semi-supervised models fared better than supervised models in predicting the group of patients that would eventually need step-up therapy in RA. 11
The different ML models used in precision medicine and the different outcome variables and the best model adopted for prediction of response to treatment are summarised in the tables below.
Rheumatoid Arthritis
Most evidence for the use of ML models in precision rheumatology stems from RA. Tumour necrosis factor-α (TNF-α) is the prime disease-driving cytokine in RA. Toll-like receptors (TLR) by inducing TNF-α have a role in the pathogenesis, and their polymorphisms, by virtue of their influence on TNF production serve as potential markers to predict response to anti-TNF and have been attractive targets to study in precision medicine, having demonstrated a role in the pathogenesis.12–14 While studying TLR polymorphisms is a novel step in itself in precision, integrating it with ML would further strengthen the predictive accuracy for remission. 15 RETN gene (coding for resistin) polymorphisms is another attractive target studied in the field of precision medicine in RA. ML models integrating RETN gene polymorphisms with the sex of the patient and other clinical factors created a robust model to predict remission in patients on anti-TNF, with the male sex favouring remission. 16
Though the practice of rheumatology may be governed, rather guided by treatment guidelines issued by different bodies such as the American College of Rheumatology (ACR), European League Against Rheumatism (EULAR), Asia Pacific League Against Rheumatism (APLAR) among the notable few, real-world practice may be far from ideal.17,18 The debunking of the apparent notion of one-size-fits-all with the deeper research into precision medicine has led to trial of therapeutics that deviate from these ‘guidelines’. However, robust evidence may not support this as randomised control trials take time to be formulated and most supportive data is from observational real-life cohorts. ML models have proven useful in such settings, for instance, to integrate variables and real-world data on tocilizumab monotherapy to provide robust evidence and build and validate a prediction model for remission with tocilizumab monotherapy in RA.19,20
Most evidence on the implementation of ML in guiding therapeutics in RA has been with anti-TNF (summarised in Table 1). While most of the studies aimed at building models integrating clinical, real-world, and biochemical data for predictive accuracy, there were some that integrated multiomics, generating more robust prediction tools. A Swedish group integrated variables from gene expression that predicted response to anti-TNF with transcriptomics, which made the predictive accuracy more robust, integrating this data with the clinical data and the transcriptome, proteome and the metabolome predicting response or lack thereof, to anti-TNF with a higher accuracy as compared to the gene transcriptions alone. The advantage of this particular model was the benefit obtained from it to predict unresponsiveness before initiating therapy that would have a huge cost benefit and reduced wastage. Predictively, precision medicine had the upper hand here, with the models integrating transcriptomic data having a higher predictive accuracy as compared to the ones integrating clinical data. 21
Summary of Studies Exploring Machine-learning Models Predicting Response to Different Therapies and Outcome in Rheumatoid Arthritis.
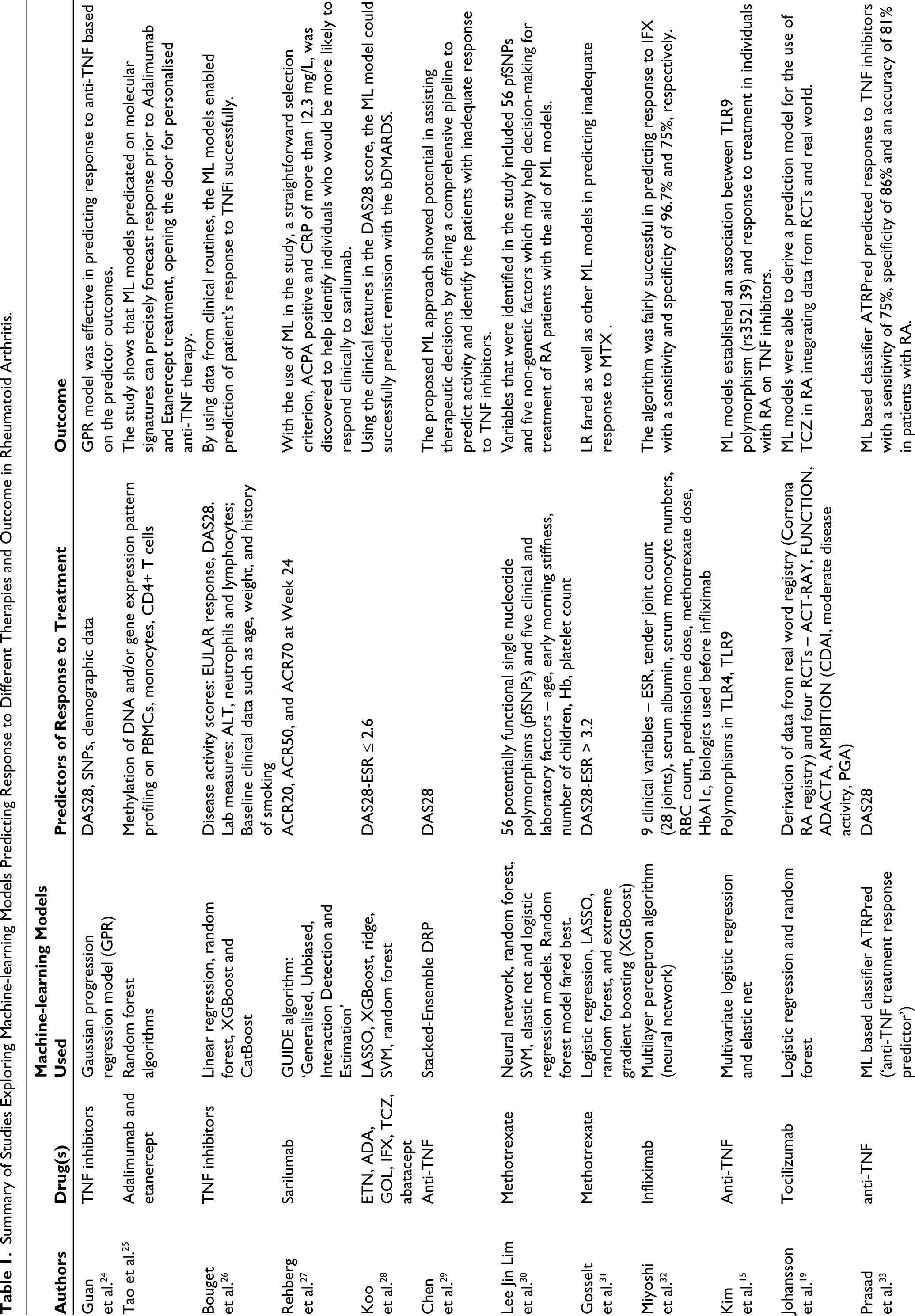

Beyond integrating genetic and multiomics to predict therapeutic response, ML models have also been used to integrate and consolidate data from various randomised controlled trials (RCTs), where meta-analysis is not available. Though methotrexate is the most used drug in RA, the response to therapy may not be homogenous. ML models can integrate and consolidate easily available data such as routine clinical information, rheumatoid factor, anti-cyclic citrullinated peptide antibody (anti-CCP), disease activity scoring, quality of health assessment, which can greatly aid the treating physician in clinical decision-making at baseline regarding methotrexate monotherapy. 22
However, the limitation with most of the current studies is the small sample size and they need to be validated in larger cohorts before implementing in real-world practice.
Beyond monitoring response to treatment and predictions, ML models have also been used to predict commonly associated complications with RA like osteoporosis, which is more common with late-onset RA. 23
Spondyloarthritis
Though most evidence for ML in precision medicine is derived from RA, there is some emerging evidence in spondyloarthritis spectrum of disorders, mainly psoriatic arthritis (PsA). In contrast to studies in RA, these studies mostly consolidated clinical data to devise models for prediction, but the genetic basis for response to treatment is still left unexplored. Compared to RA, the disease drivers and the pathogenetic factors in PsA are more multifaceted, and targeting a single cytokine or gene editing may not result in robust disease control. Precision medicine and personalised medicine have a greater role to play in such diseases, where determining the exact pathway that is dominant, has a great translational relevance. Also in such scenarios, ML has an advantage as the pathogenesis here is a complex interplay of multiple pathways and networks of cytokines.
Evidence in PsA is mostly limited to secukinumab where one group has explored the factors predicting remission to the drug, while the other tried to determine the set of patients that would respond to a starting dose of 150mg versus 300mg as this is a common dilemma encountered in the clinic. 40 While most of these decisions are left to the discretion of the treating physician, employment of ML models provides evidence with high predictive accuracy. PsA is a disease with heterogenous presentations ranging from predominantly cutaneous psoriatic phenotype or peripheral deforming arthritis or a predominant axial disease. 41 Owing to the non-homogeneity in presentation, the first choice of DMARD may not always be the right one. Additionally, the options of biological therapy also include anti-TNF and IL-17 and at baseline, and predicting the response to a particular drug is not foolproof. ML models can greatly aid in this treatment decision and result in saving time and finances by avoiding the ‘trial-and-error’. In patients with inadequate response to anti-TNF, those with early PsA and that had enthesitis at baseline were predicted to have remission with secukinumab and also 300mg fared better than 150mg in those treated without concomitant methotrexate, and with PsO. 42
One conflicting evidence in this regard comes from a study in ankylosing spondylitis (AS), where Lee et al. found no benefit of implementing ML models over traditional logistic regression (LR) model in predicting response to bDMARD in AS. In the same study, they found benefit in a random forest model in RA patients; however, ML failed to fare better than LR in AS. 43 The same group subsequently presented an artificial neural network model that integrated demographic and lab data to predict the patients that would require TNFi within six months of diagnosis in AS. 44
The studies exploring ML models in therapeutic response in SpA are summarised in Table 2.
Machine-learning Models Predicting the Response to Treatment and Outcome in Spondyloarthritis.
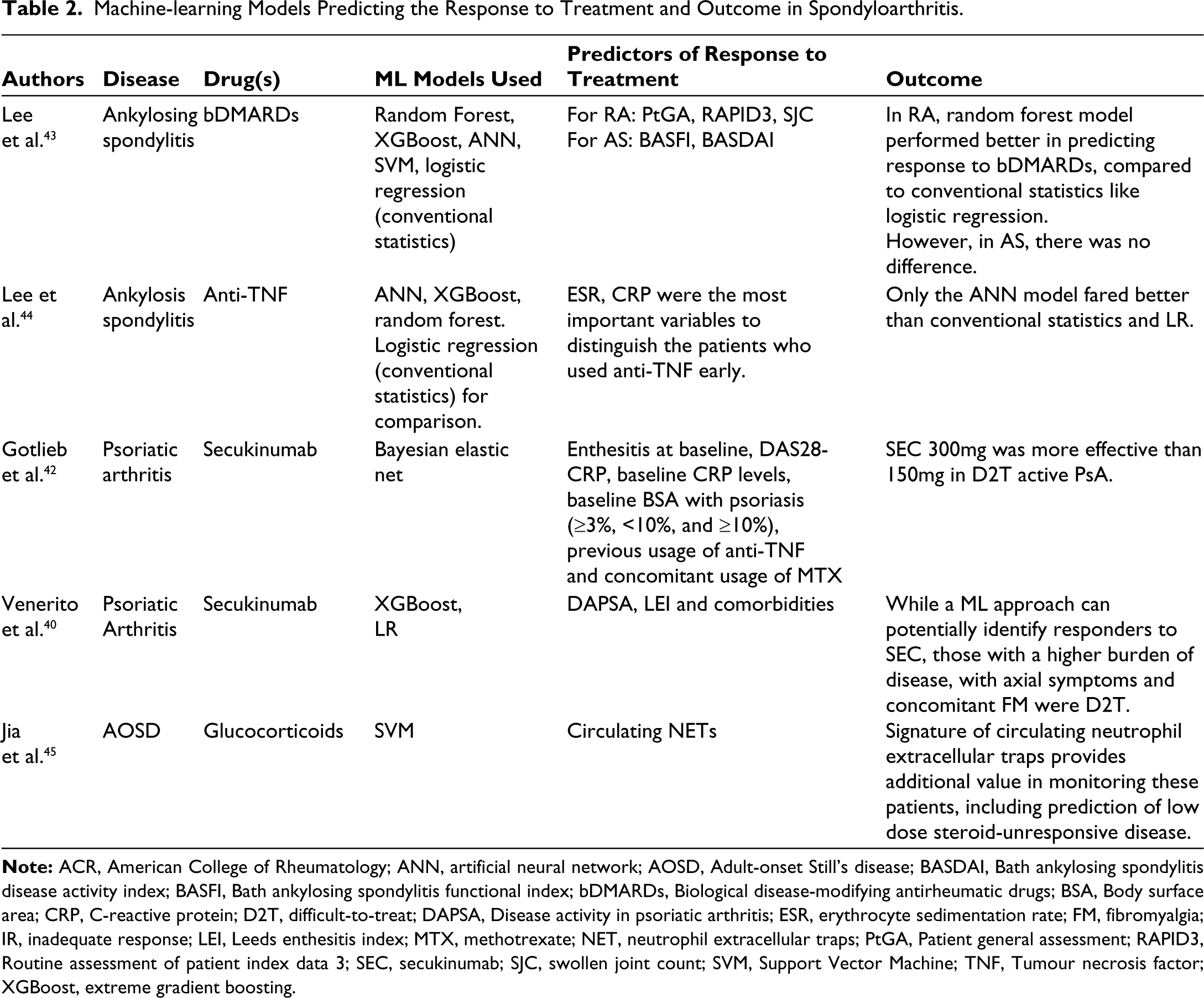
Connective Tissue Diseases
A novel insight into precision medicine and ML in SSc was provided by BK Mehta et al., where they hypothesised that patients of early diffuse SSc with an inflammatory phenotype would have the best response to abatacept and this would depend on the CD28 reactome. 46 Integrating data from the molecular signature patterns of the skin, that is, the inflammatory pattern and the CD28 pathway, which is the one directly affected by abatacept, it was demonstrated that the patients with early disease and an inflammatory phenotype had the best cutaneous response and improvement in mRSS to abatacept.
The researchers from Japan in their landmark DesiReS trial, demonstrated benefit of rituximab on skin fibrosis in systemic sclerosis. 47 However, the therapeutic armamentarium of SSc-related skin fibrosis is wide, and choosing the right agent avoids the hassle of failures and adverse events. The causal tree ML model was implemented in a post-hoc analysis of DesiReS that aided in accurately predicting the set of patients who would have the best response to RTX, by combining clinical and immunological markers to find the optimal predictors 48 (Table 3). Clinical decisions guided by these factors can greatly influence therapeutics and avoid polypharmacy and unnecessary trial to multiple immunosuppression and potentially prevent adverse events and infections.
Machine-learning Models Predicting Response to Treatment and Outcome in Other Connective Tissue Diseases and Vasculitis.
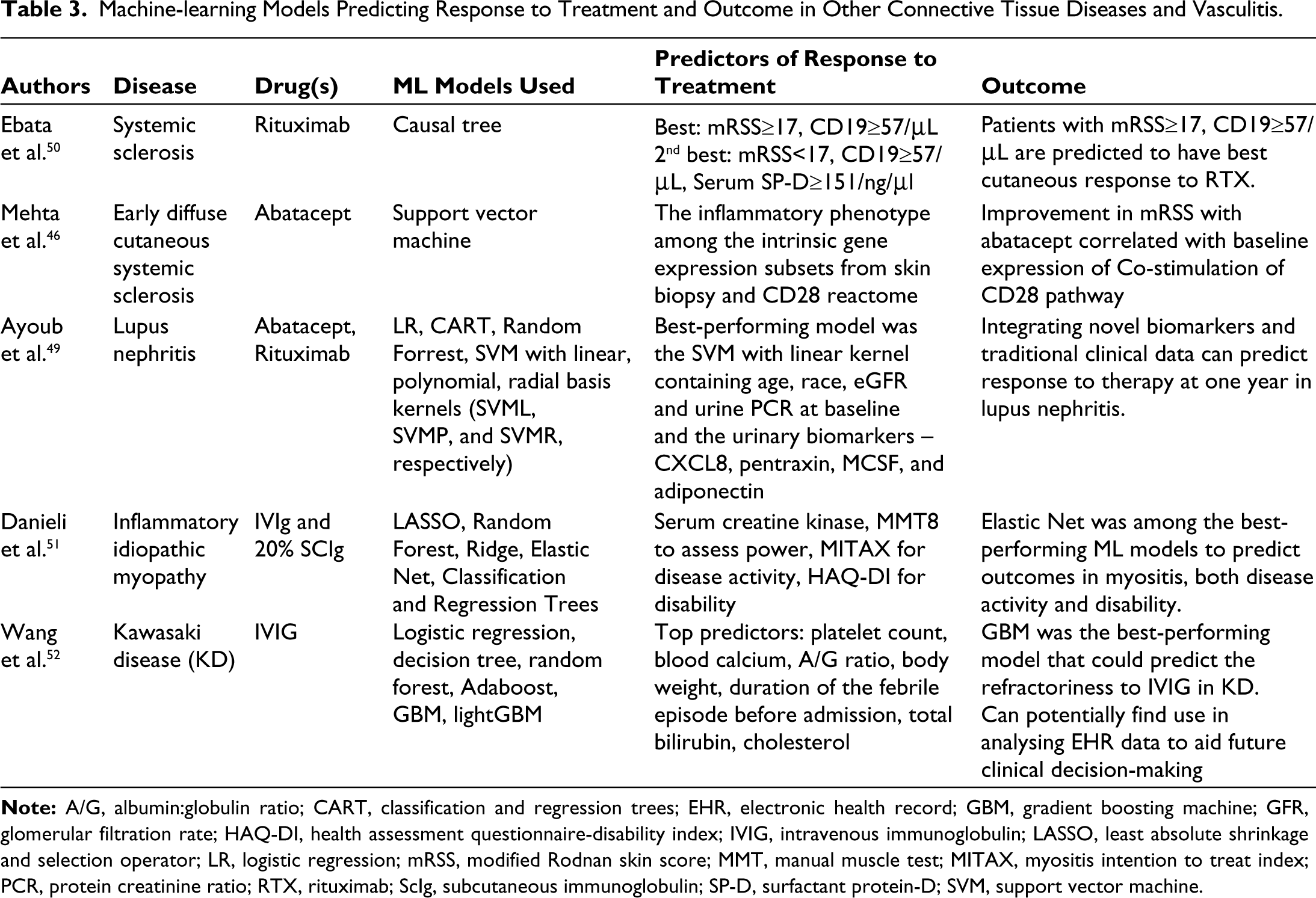

Application of ML in precision medicine in lupus has been surprisingly scarce. We could find one study analysing data from existing cohorts to train and a longitudinal cohort of flare of SLE to validate the model integrating some novel biomarkers and clinical features to build a model to predict response to therapy, mostly abatacept and rituximab (summarised in Table 3). 49 However, the possible fallacy of implementing ML models in lupus may be the sheer number of markers, in routine clinical use or in research, that are available to monitor disease or treatment. It is imperative and ideal to check a combination of different markers to find the best-performing model for predictive accuracy, and these associations may not be known, or easily predictable in retrospect.
The other models used in isolated disease scenarios are summarised in Table 3.
Limitation of Current Evidence
Though there are many advantages of implementation of ML in health care and precision medicine such as robust analysis of big data, better predictive accuracy with a relative smaller sample size, identification of novel biomarkers for prediction and prognosis, these models are not without their limitations. Most of the cohorts where these models have been devised are small and they need to be validated in larger, preferably population-based studies for more robust evidence. While individual studies claim different biomarkers as predictors of response, there is clear lack of generalizability, which may lead to spurious claims of discovery of novel biomarkers. 7
Moreover, there are limited data on the precision of these models, and if the findings of a model can be replicated in other independent models, then there needs to be a validation in real-life scenarios where the utilisation of ML leads to better patient outcomes. However, these can be done only once the models are established based on high-quality, longitudinal data.
While countries such as India have high patient loads, the health-care workers-to-patient ratios are not conducive to allow collection of comprehensive, high-quality data. 53
Conclusion
Attaining the goals of precision medicine will be difficult without the application of ML. Understanding the ‘systems biology’ of each disease and identifying the right set of biomarkers and clinical variables to predict the course and response can potentially save time, and reduce damage accrual by bypassing the trial of traditional/conventional treatment that may not be precise in the particular patient, have cost benefits and eventually translate to better outcomes. However, issues with lack of generalizability exist, leaving researchers and clinicians to ponder if every novel biomarker or predictor is to be taken at face value.
The current challenge lies in collecting quality longitudinal data and the application of robust ML models that can be replicated and validated in population-based studies, beyond the small, isolated cohorts.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Ethical Approval
Not applicable.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
Informed Consent
Not applicable.
