Abstract
Background
Beta-sitosterol, a phytosterol similar to cholesterol, is found in various plants and is recognized for its potential anticancer properties; still, the mechanism of breast cancer (BC) remains elusive.
Objectives
This study investigates the potential targets of beta-sitosterol in BC using network pharmacological analysis and in vitro validation.
Materials and Methods
Targets of beta-sitosterol and BC were identified from online databases, including Swiss Target Prediction, SuperPred, GeneCards, and DisGeNET. Protein–protein interactions of common targets were analyzed using STRING version 11.0, and network construction and core target screening were performed with Cytoscape 3.8.0. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis and gene ontology (GO) annotation were conducted using the ShinyGO web tool. Molecular docking analysis was done with AutoDock Vina and the in vitro validation, including cell viability, apoptosis, and gene expression analysis was done using MTT assay, acridine orange/ethidium bromide staining, and polymerase chain reaction (PCR), respectively.
Results
Sixty-eight common targets for beta-sitosterol and BC were identified from 209 beta-sitosterol targets and 1,350 BC-related genes. Five core targets (HIF1A, ESR1, STAT3, TNF, and MAPK3) were identified. GO analysis linked common targets to biological processes, cellular components, and molecular functions. KEGG pathway analysis showed enrichment in pathways such as central carbon metabolism in cancer, vascular endothelial growth factor signaling, and prolactin signaling. Beta-sitosterol’s impact on MCF7 cell viability was assessed via MTT assay, and its molecular effects were explored through PCR. Results demonstrated beta-sitosterol’s ability to hinder tumor progression by modulating genes involved in ESR1, mitogen-activated protein kinase, and tumor necrosis factor signaling pathways.
Conclusion
These findings highlight beta-sitosterol’s potential as an anticancer agent, offering new perspectives for clinical investigations through the modulation of key signaling pathways in BC.
Introduction
Despite considerable progress in research, cancer continues to be a leading cause of human fatalities globally. According to Global Cancer Observatory (GLOBOCAN) 2020, the anticipated global cancer burden is predicted to reach 28.4 million cases by 2040, reflecting a 47% increase from 2020 (Sung et al., 2021). Among these, breast cancer (BC) stands as the most prevalent malignancy among women. While the majority of BCs are benign and can be cured through surgery, approximately one-quarter exhibit a latent and insidious nature, growing slowly yet metastasizing at an early stage. Due to its elevated malignancy and swift metastasis, BC prognosis remains unfavorable, characterized by comparatively low survival rates (Cowin et al., 2005). Primary risk factors linked to BC include physical inactivity, hormones, inadequate nutrition, hormone replacement therapy, obesity, smoking, and alcohol consumption. Hereditary and genetic factors play a crucial impact in BC’s etiology. Researchers have shown that mutations in the BRCA1 and 2 genes contribute to 5–10% of cases, while promoting breastfeeding and engaging in physical activity can potentially reduce the risk (Schottenfeld & Fraumeni, 2006). The classification, grounded in varied gene expression patterns, identifies at least four primary subtypes of BC: triple-negative subtypes, HER2, luminal A, and luminal B overexpression (Sotiriou et al., 2003).
Understanding the principal actors and molecular mechanisms underlying BC metastasis is crucial due to the high likelihood of recurrence and metastasis to critical organs, including the brain, lungs, bones, and liver, ultimately leading to patient mortality (Ismail et al., 2023). Chemotherapy, in particular, is used as a postsurgical intervention or as an alternative in nonsurgical situations. Despite the implementation of numerous new targeted therapies over the past decades, such as PARP inhibitors (Robson et al., 2017), CDK 4/6 inhibitors (Finn et al., 2016), immunotherapy (Schmid et al., 2018), a phosphoinositide 3 kinase (PI3K) inhibitor (Wu et al., 2023), and HER2-targeted therapies (Swain et al., 2013), the overall improvement in the prognosis of advanced breast cancer (ABC) patients has been relatively modest. However, it is important to note that these chemotherapeutic drugs may still have associated side effects. The increasing burden of cancer highlights the urgent requirement for developing and implementing effective, less toxic, chemoprevention, and chemotherapy strategies that can positively impact diverse populations worldwide. Medicinal plants are documented as a sustainable source of bioactive molecules, particularly phytochemicals. Compounds like polyphenols, resveratrol, curcumin, genistein, sulforaphane, isothiocyanates, and silymarin found in these plants show promise in treating various ailments. Specifically, they exhibit the potential in suppressing or reversing the early stages of cancer, inhibiting the invasiveness of premalignant cells, and modulating cell proliferation and apoptosis signaling pathways in BC cells (Wang et al., 2012).
This study is particularly interested in beta-sitosterol, a medicinal compound derived from a tree, due to its potential medicinal properties, including analgesic (Nirmal et al., 2012), anti-inflammatory (Liao et al., 2018), antipyretic (Gupta et al., 1980), and antidiabetic activities (Babu & Jayaraman, 2020). Beta-sitosterol is noteworthy for its dual functionality as an antioxidant and an inducer for cell death in various cancer cell lines (Halimah et al., 2023). Beta-sitosterol has shown chemo-enhancing effects on tumor cells, particularly in BC. It interferes with numerous cellular signaling pathways associated with cell cycle arrest, cellular proliferation, apoptosis, cell survival, metastasis, invasion, and angiogenesis (Nandi et al., 2024). Various studies have contributed to the discovery of potential targets and target-based therapeutics from plant sources using computational methods. However, the multi-targeting mechanism responsible for the cancer-inhibitory activity of beta-sitosterol in BC requires further clarification. Network pharmacological analysis is a nascent field grounded in the principles of systems biology. It involves the systematic assessment of the pharmacological effects of drugs containing multiple components and targeting multiple pathways. For instance, Mishra et al. (2022) have identified lepitaprocerin D as a potential modulator of BUB1B, which could help control the spread of BC metastasis through in silico studies. This is achieved by establishing connections between drug components, targets, and diseases through the analysis of biological system networks (Zhao et al., 2023). In this study, our objective is to unveil beta-sitosterol’s BC inhibitory potential through an integrated approach that involves the prediction of absorption, distribution, metabolism, and excretion (ADME), along with network pharmacological analysis. Initially, we collected targets associated with BC and beta-sitosterol from various online databases. These targets underwent common target identification, protein–protein interaction (PPI), gene ontology (GO), and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses. Subsequently, we screened hub genes associated with BC and beta-sitosterol. In addition, molecular docking analysis was conducted using the hub genes against beta-sitosterol to assess the molecular impact of beta-sitosterol in BC treatment. The following experiments sought to confirm the mechanism of beta-sitosterol in BC. Consequently, this study aims to anticipate and confirm the target and potential mechanism of beta-sitosterol’s anti-BC properties through network pharmacological analysis and cell-based experiments.
Materials and Methods
ADME Prediction in silico
The structural information of beta-sitosterol (PubChem CID: 222284) was acquired from the NCBI PubChem database. For beta-sitosterol, ADME screening criteria were applied, including assessments of drug-likeness (DL), bioavailability (OB), and other parameters. The SwissADME web tool was utilized for these analyses.
Target Retrieval and Identification of Intersecting Genes
The potential targets of beta-sitosterol were collected from Super Pred and Swiss Target Prediction web tools. In contrast, targets associated with BC were obtained from the DisGeNET online database and Gene Cards, with a threshold value of 20 or above. After removing duplicates, the list of target names and gene IDs was updated through integration via the UniProt database. To identify shared genes between beta-sitosterol and targets associated with BC, a Venn diagram was generated. This visualization helps to elucidate the shared targets between beta-sitosterol and those associated with BC, providing insight into the potential impact of beta-sitosterol in BC treatment.
PPI Network Construction and Hub Gene Prediction
The genes that were identified as common between beta-sitosterol and BC targets were subjected to a PPI study using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) database. The analysis focused on “Homo sapiens” with a set confidence value for the interaction of 0.7, ensuring high confidence in the interactions on the STRING platform. The construction of the PPI network involved generating a connectivity diagram, allowing for an exploration of the interactions between BC and beta-sitosterol-related targets.
Detection of Pivotal Targets
Cytoscape 3.9.0 was utilized to assess the topology parameters of the constructed network for identifying target genes. The degree value was visually represented by color intensity. The CentiScaPe 2.2 plugin was employed to identify primary targets by calculating mean values for closeness, betweenness, and degree. Subsequently, the Maximum Clique Centrality (MCC) method was applied to find the top five hub genes.
GO and KEGG
The core targets that were identified underwent analysis for enrichment in GO and KEGG analysis using the ShinyGO version 0.80 web tool. “Homo sapiens” was selected as the species background for this analysis. The GO annotation encompassed molecular functions (MFs), biological processes (BPs), and cellular components (CCs).
Molecular Docking
The three-dimensional (3D) structures in PDB format for the top five targeted proteins were acquired from the RCSB Protein Data Bank and converted into the PDBQT format, charges were assigned, and water molecules were removed using the AutoDock bioinformatics tool. For ligand preparation, the 3D structure of beta-sitosterol was acquired in Structure Data File (sdf) format from PubChem. The pdbqt file format for beta-sitosterol was generated using AutoDock (version 1.5.7). To evaluate the docking score and binding affinity of beta-sitosterol with the target proteins, AutoDock (version 4.2) was employed. The optimal binding pose of beta-sitosterol with the top targeted proteins was visualized using the DS Visualizer software.
Cell Culture and Cell Viability Assay
MCF7 cells were cultured in Dulbecco’s modified Eagle medium (DMEM) (Gibco, USA) supplemented with 10% fetal bovine serum (FBS) (Gibco, USA), 100 units/ml penicillin, and 100 mg/ml streptomycin under standard conditions (5% CO2, 37°C, 95% humidity) to maintain their growth phase consistently. Beta-sitosterol was administered at varying concentrations (ranging from 0 to 100 µM) for 24 hours. Following treatment, an MTT assay was conducted to determine the IC50 concentration of beta-sitosterol in MCF7 BC cells.
Polymerase Chain Reaction
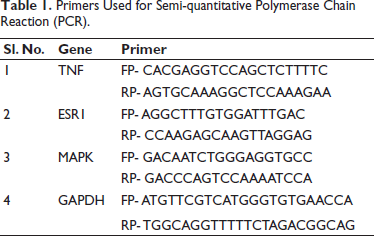
Total RNA from MCF7 cells was extracted following the protocol outlined in the RNA isolation kit (One-step RNA TRIzol Reagent; Biobasic Inc., Markham, Ontario, Canada). First-strand cDNA synthesis was conducted using the iScript cDNA synthesis kit from Bio-Rad Laboratories, Inc., USA, as per the manufacturer’s instructions. Real-time amplification of the cDNA was carried out using SSO Advanced Universal SYBR Green Supermix, following the manufacturer’s protocol (Bio-Rad Laboratories, Inc., USA). Gene expression was assessed using a conventional PCR system, utilizing the primer list provided in the accompanying Table 1.
Primers Used for Semi-quantitative Polymerase Chain Reaction (PCR).
Acridine Orange/Ethidium Bromide (AO/EtBr) Staining
AO/EtBr staining was employed to assess the qualitative level of damage caused by beta-sitosterol on MCF7 cells. MCF7 cells were cultured in six-well plates using serum-free media, with or without beta-sitosterol treatment for 24 hours. Both control and treated MCF7 cells were subjected to 10 µM (AO/EtBr) staining, where acridine orange illuminated live cells and ethidium bromide highlighted damaged cells. Images were captured at a magnification of 20×.
Statistical Analysis
The data is presented as Mean ± Standard Deviation (SD). Group variances were evaluated through one-way analysis of variance (ANOVA) analysis conducted with GraphPad Prism 9.0 software. Subsequent pairwise comparisons between groups were carried out utilizing Tukey’s multiple comparisons test. Statistical significance was established at p < 0.05.
Results
ADME Prediction of Beta-sitosterol
The canonical SMILES format of beta-sitosterol (PubChem ID: 222284) was obtained from the PubChem database. This SMILES notation was then input into the SwissADME web tool to predict DL and physicochemical properties. The results indicated that most of the pharmacokinetic (ADME) parameters were found to be quite agreeable and within the satisfactory range, as shown in Table 2.
Physiological Properties of Beta-sitosterol.

Target Retrieval and Intersecting Genes
A cumulative of 209 targets for beta-sitosterol (Figure 1A) and 1,350 targets associated with BC were curated. The overlapping targets between BC and beta-sitosterol were considered as potential targets. Utilizing this data, the Venn diagram web tool was employed to identify 68 shared therapeutic targets for BC and beta-sitosterol.
(A) The Venn Diagram Illustrating Common Targets in Breast Cancer and Beta-sitosterol. (B) Top 5 Key Targets of Beta-sitosterol in Breast Cancer.
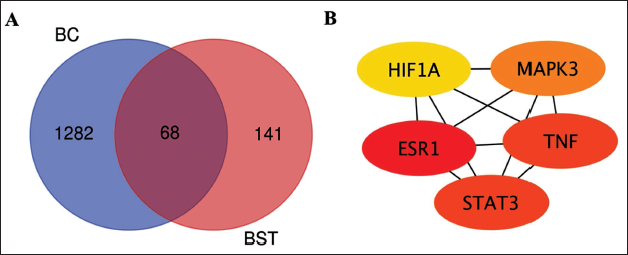
Network of PPI Among Shared Targets
Using the information provided, the Venn diagram Webtool was used to ascertain 68 shared therapeutic targets for BC and beta-sitosterol (Figure 1A). Subsequently, the PPI of these 68 common targets underwent STRING analysis, followed by the construction of a network showcasing genes shared between them in Cytoscape. Topological properties were scrutinized to pinpoint hub targets, revealing hypoxia-inducible factor 1alpha (HIF1A), estrogen receptor 1 (ESR1), signal transducer and activator of transcription 3 (STAT3), tumor necrosis factor (TNF), and mitogen-activated protein kinase (MAPK3) as prominent entities in this complex biological network (Figure 1B). Notably, most of these hub genes are significant in the context of BC. Consequently, the subsequent molecular docking investigation focused on these hub targets, suggesting their potential pivotal role in the treatment of BC with beta-sitosterol.
Gene Ontology
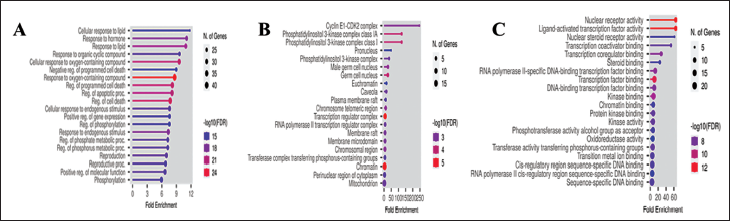
The exploration of biological functions and pathways associated with BC and beta-sitosterol involved the use of GO terms through the ShinyGO web tool. After p value correction, the top 20 most significant GO terms related to BPs, CCs, and MFs were identified (Figure 2A, B, and C). According to the GO analysis, the key GO-BPs contributing to the anticancer properties of beta-sitosterol were associated with the cellular response to lipids and response to hormones. The most significant GO-CC terms included the cyclin E1-CDK2 complex and phosphatidylinositol 3-kinase complex class IA. Within GO-MFs, crucial terms encompassed ligand-activated transcription factor activity and nuclear receptor activity.
Top 20 Gene Ontology Map of Putative Target Genes. (A) Biological Process Categories. (B) Cellular Component Categories. (C) Molecular Function Categories.
KEGG Pathway Enrichment Analysis with Intersected Targets
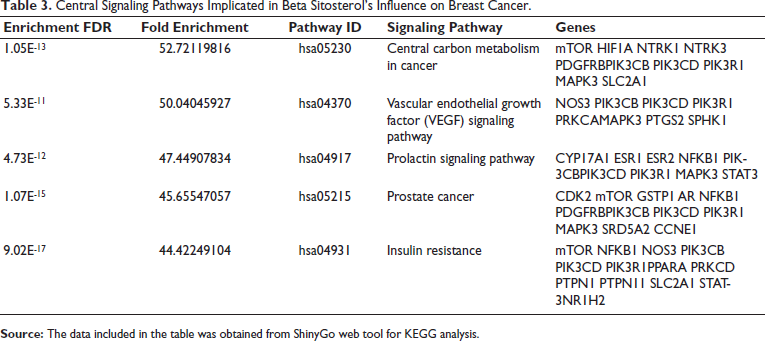
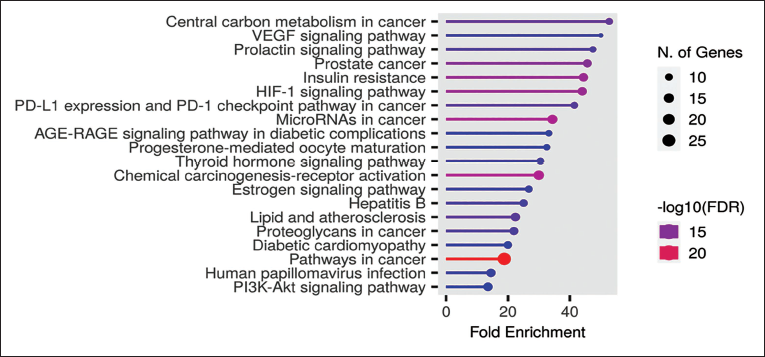
The therapeutic targets underwent enrichment analysis for KEGG pathways using Shiny Go, with a significance threshold set at p < 0.05. This screening identified prominent pathways as depicted in Table 3. Following this, networks specific to beta-sitosterol-targeted pathways were constructed by organizing the genes enriched in each pathway. Notably, three pathways stood out prominently for their significant enrichment: central carbon metabolism in cancer, vascular endothelial growth factor (VEGF) signaling pathway, and the prolactin signaling pathway (Figure 3).
Central Signaling Pathways Implicated in Beta Sitosterol’s Influence on Breast Cancer.
Top 20 Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathway Enrichment Analysis Map.
Confirmation of Binding Interaction of Beta-sitosterol with Hub Genes
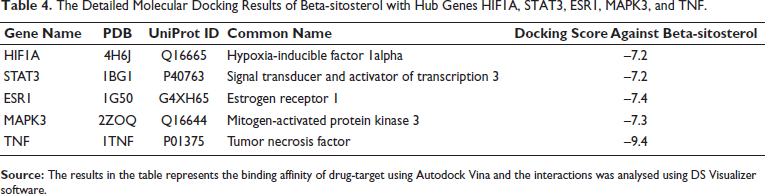
Beta-sitosterol was subjected to molecular docking analysis with five hub genes. Typically, a more robust binding affinity between a drug and its target indicates a higher drug binding capacity. In this investigation, specific focus was given to beta-sitosterol targets that displayed the most robust binding. Proteins forming strong bonds with beta-sitosterol might play a role in reducing BC tumorigenesis. Importantly, the central key targets exhibited a strong binding affinity for beta-sitosterol, as depicted in and detailed in Table 4.
The Detailed Molecular Docking Results of Beta-sitosterol with Hub Genes HIF1A, STAT3, ESR1, MAPK3, and TNF.

Beta-sitosterol Inhibits BC Cell Proliferation
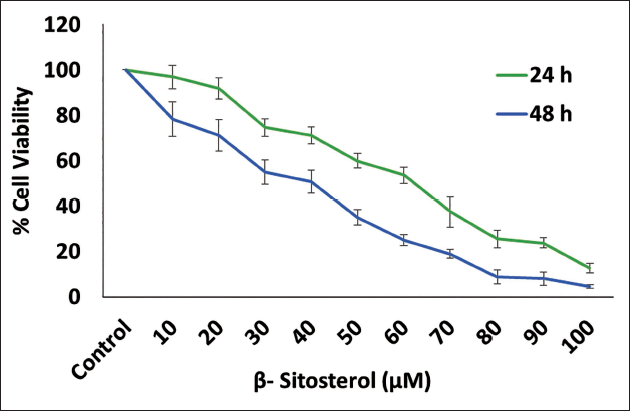
The MTT assay revealed a significant decrease in cell viability in MCF7 cells treated with the increasing concentration of beta-sitosterol for 24 and 48 hours, with an IC50 value of 59 ± 3.2 µM and 36 ± 5.1 µM. This indicates that beta-sitosterol effectively inhibits cell proliferation in MCF7 cells in a dose-dependent manner (Figure 4).
Impact of Beta-sitosterol on Viability of MCF7 Cells Assessed by MTT Assay for 24 and 48 Hours with Different Concentration of Drug Treatment Ranges from 0 to 100 µM.
Beta-sitosterol Induces Apoptosis in BC Cells
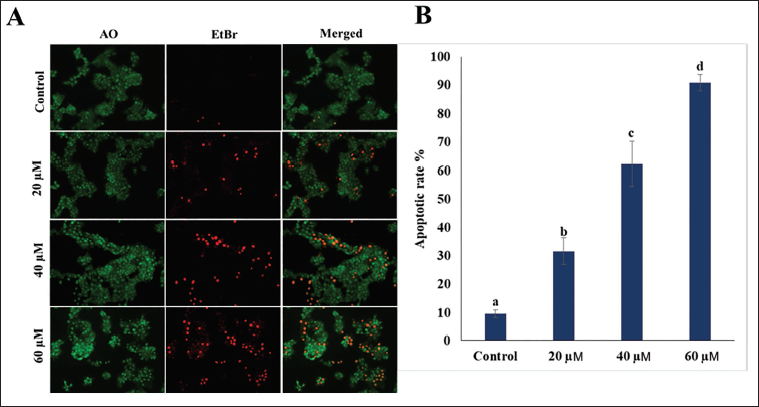
Beta-sitosterol treatment triggers apoptosis-related alterations in MCF7 cells, the cells were fluorescently stained with AO/EtBr and examined under a fluorescent microscope. Following 24 hours of beta-sitosterol treatment, MCF7 cells were stained using AO/EtBr dyes. This staining revealed distinct morphological changes such as condensed chromatin, nuclear condensation, and fragmentation in a dependent manner (Figure 5). Moreover, the apoptotic cell population was found to be 31 ± 4.6%, 62 ± 7.8%, and 90 ± 2.9% for the treatment of 20, 40, and 60 µM beta-sitosterol, respectively.
Acridine Orange/Ethidium Bromide (AO/EtBr) Staining to Assess the Apoptotic Rate of Cells Upon Treatment with Beta-sitosterol with Different Concentrations (0, 40, 60, and 80 µM).
Beta-sitosterol Regulates the Gene Expression of Hub Genes
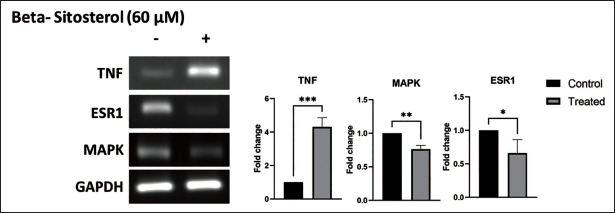
The mRNA expressions of predicted hub genes, namely TNF, ESR1, and MAPK, were investigated via PCR to elucidate their role as potential targets of beta-sitosterol in BC tumor progression. Results revealed significant alterations in the expression levels of these hub genes upon treatment with beta-sitosterol. Notably, beta-sitosterol treatment led to a substantial reduction in the expression of MAPK and ESR1, both of which are known to positively regulate tumorigenesis. Conversely, beta-sitosterol treatment elevates TNF expression, suggesting its involvement in inducing apoptosis (Figure 6).
Discussion
Despite continuous advancements in medical research, BC persists as the second most prevalent and deadly malignancy among females. Indeed, in recent years, there have been notable improvements in both basic research and clinical applications of phytomedicine for treating BC. As a result, researchers have increasingly turned their attention to exploring antitumor drugs derived from phytomedicine. This interest is fueled by the perceived advantages of these drugs, including lower negative effects and greater specificity in targeting crucial proteins involved in the abnormal activation of pathways associated with cancer metastasis. However, it is critical to note that while phytomedicine exerts a synergistic effect with its multicomponent and multitarget nature, it also poses significant challenges in understanding the drug mechanism of action and the development of new drugs (Devan et al., 2023).
Computational studies play an effective role in elucidating the mechanisms of cancer metastasis. For example, research has discovered prognostic biomarkers in various cancers, including BC (Mishra et al., 2023a) and hepatocellular carcinoma (Mishra et al., 2023b) through integrated transcriptome and pathway enrichment analyses using computational data. These computational analyses are highly beneficial for discovering novel, potential, and less toxic drug candidates for cancer treatment.
Beta-sitosterol, a primary dietary phytosterol present in plants, has potential applications in the prevention and therapy of various cancers (Awad et al., 2000; Choi et al., 2003). However, the specific molecular mechanism through which beta-sitosterol exerts its cancer-inhibitory activity by inducing apoptosis remains elusive. Through network pharmacological analysis, the intricate interactions between a drug and its targets, as well as potential mechanisms, can be effectively elucidated at a systemic level. So, this study aims to investigate potential therapeutic molecules for BC, considering beta-sitosterol targets through a network pharmacological analysis approach.
The study sourced beta-sitosterol’s 209 targets and 1,350 targets associated with BC from various online databases. Using a Venn diagram tool, 68 common intersecting/shared genes were identified. These common targets underwent further analysis, including GO classification into BPs, MFs, and CCs, as well as PPI, hub gene identification, network construction, and KEGG pathway enrichment analysis. Based on the GO analysis, the top 20 categories were curated, and the primary GO-BPs contributing to beta-sitosterol’s anticancer properties were associated with cellular response to lipids, response to hormones, response to lipids, response to organic cyclic compound, and cellular response to oxygen-containing compound. The most significant GO-CC terms included the cyclin E1-CDK2 complex, phosphatidylinositol 3-kinase complex class IA, phosphatidylinositol 3-kinase complex class I, and pronucleus. Within GO-MFs, crucial terms encompassed ligand-activated transcription factor activity, nuclear receptor activity, nuclear steroid receptor activity, and transcription coactivator binding.
In the network pharmacological analysis, we identified the top five core genes, HIF1A, ESR1, STAT3, TNF, and MAPK3, as hub targets. These genes are predicted to play a vital role in the effects of beta-sitosterol against BC. Elevated levels of HIF-1α have been linked to increased metastasis and reduced survival rates in BC patients (Wong et al., 2011). Conversely, beta-sitosterol treatment has demonstrated the potential to decrease HIF1A expression in various pathophysiological conditions, such as arthritis (Zhang et al., 2020). Therefore, it is anticipated that beta-sitosterol may exhibit cancer inhibitory activity by suppressing the levels of HIF1A, which are observed to increase in BC. The GO analysis reveals that ligand-activated (nuclear receptor) activity is significantly involved in the progression of BC. ESR1, recognized as a nuclear receptor, is acknowledged as a crucial player in the initiation of BC through mutation and alterations in expression. Furthermore, beta-sitosterol has been predicted to be a target candidate for ESR1 (Wu et al., 2022). TNF-α plays a key role in initiating an immune response and supporting various phases of tumor progression, including initiation, tumor cell proliferation, cell survival, and metastasis (Liu et al., 2020). In this study, it has been reported that the STAT3 signaling pathway is implicated in BC proliferation and apoptosis through the downregulation of the zinc-finger gene DPF3 (a.k.a. CERD4). This downregulation promotes the proliferation and invasion of BC by activating the Janus kinase (JAK) 2/STAT3 pathway, which subsequently upregulates cyclin D-1, c-myc, and B-cell lymphoma 2 (bcl-2), leading to the suppression of apoptosis in BC cells (Lin et al., 2019). The degradation of ULK1, dependent on MAPK1/3 kinase, has been identified as a mechanism that attenuates mitophagy and supports BC bone metastasis (Deng et al., 2021). The KEGG pathway analysis highlights that the VEGF signaling pathway, central carbon metabolism in cancer, and prolactin signaling pathway are pivotal in the progression of BC. Furthermore, the molecular docking analysis unveils the protein relationship between beta-sitosterol and the top hub genes by interacting with high binding affinity. In summary, beta-sitosterol has the potential to influence HIF1A, ESR1, STAT3, TNF, and MAPK3, thereby regulating the VEGF and prolactin signaling pathways for the treatment of BC, with MAPK3 playing a crucial role in this regulatory process.
In silico predictions identified ESR1, MAPK, and TNF as core targets for BC treatment. The MTT assay demonstrated a concentration-dependent inhibition of MCF7 cell proliferation by beta-sitosterol, yielding an observed IC50 concentration of 59 ± 3.2 µM and 36 ± 5.1 µM for 24 and 48 hours of treatment, respectively. Additionally, AO/EtBr staining revealed the induction of apoptosis in treated cells. Subsequent mRNA-level analysis of hub genes under beta-sitosterol treatment conditions showed significant alterations, particularly in TNF, MAPK, and ESR1 expression. These findings substantiate that beta-sitosterol effectively suppresses tumor proliferation and induces apoptosis by targeting pivotal genes such as TNF, MAPK, and ESR1. Nonetheless, further in vivo experiments are essential for confirming their activities. Furthermore, our study suggests that applying network pharmacological analysis to investigate the effects of beta-sitosterol against BC is a scientifically valid and feasible approach.
Conclusion
Through an integrated approach encompassing database target screening, PPI analysis, GO, KEGG pathway exploration, and molecular docking studies, we have unveiled a suite of potential targets and signaling pathways that have thus far received limited attention in studies investigating the efficacy of beta-sitosterol against BC. Our investigation has pinpointed crucial pathways implicated in the action of beta-sitosterol in BC, notably including the VEGF signaling pathway, central carbon metabolism in cancer, and the prolactin signaling pathway. These findings provide a conceptual framework and offer valuable insights for further inquiry into the anti-BC properties of beta-sitosterol. Experimental validation has demonstrated that beta-sitosterol treatment of MCF7 cells for 24 hours exerts an impact on cell viability and triggers apoptosis by modulating key target genes, including TNF, ESR1, and MAPK. These observations collectively suggest that beta-sitosterol treatment presents a promising alternative strategy for BC therapy.
Footnotes
Abbreviations
3D: Three-dimensional; ADME: Absorption, distribution, metabolism, and excretion; BC: Breast cancer; bcl-2: B-cell lymphoma 2; BPs: Biological processes; CCs: Cellular components; GO: Gene ontology; GLOBOCAN: Global Cancer Observatory; HIF1A: Hypoxia-inducible factor 1alpha; JAK: Janus kinase; KEGG: Kyoto Encyclopaedia of Genes and Genomes; MAPK: Mitogen-activated protein kinase; MCC: Maximum Clique Centrality; MFs: Molecular functions; PCR: Polymerase chain reaction; PDB: Protein Data Bank; PI3K: Phosphoinositide 3 kinase; PPARα: Peroxisome proliferator-activated receptor-alpha; PPI: Protein–protein interaction; RCSB: Research Collaboratory for Structural Bioinformatics; SDF: Structure Data File; SMILES: Simplified Molecular Input Line Entry System; STAT3: Signal Transducer and Activator of Transcription 3; STRING: Search Tool for the Retrieval of Interacting Genes/Proteins; TNF: Tumor necrosis factor; VEGF: Vascular endothelial growth factor.
Acknowledgments
None.
Authors’ Contributions
JY and SC—Manuscript writing and experimental design; MZ and XZ—Experimental analysis; XZ—Manuscript review.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent Statement
Not applicable.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work is funded by Tianjin Key Medical Discipline (Specialty) Construction Project (TJYXZDXK-009A) and Jiangsu Province elderly health research project (LD2022009).
