Abstract
Background:
Coronaviruses (CoVs) are single-stranded, polyadenylated, enveloped RNA of positive polarity with a unique potential to alter host tropism. This has been exceptionally demonstrated by the emergence of deadly virus outbreaks of the past: Severe Acute Respiratory Syndrome (SARS-CoV) in 2003 and Middle East Respiratory Syndrome (MERS-CoV) in 2012.
Summary:
The 2019 outbreak by the new cross-species transmission of SARS-CoV-2 has put the world on alert. CoV infection is triggered by receptor recognition, membrane fusion, and successive viral entry mediated by the surface Spike (S) glycoprotein. S protein is one of the major antigenic determinants and the target for neutralizing antibodies. It is a valuable target in antiviral therapies because of its central role in cell-cell fusion, viral antigen spread, and host immune responses leading to immunopathogenesis. The receptor-binding domain of S protein has received greater attention as it initiates host attachment and contains major antigenic determinants. However, investigating the therapeutic potential of fusion peptide as a part of the fusion core complex assembled by the heptad repeats 1 and 2 (HR1 and HR2) is also warranted. Along with receptor attachment and entry, fusion mechanisms should also be explored for designing inhibitors as a therapeutic intervention.
Key message:
In this article, we review the S protein function and its role in mediating membrane fusion, spread, tropism, and its associated pathogenesis with notable therapeutic strategies focusing on results obtained from studies on a murine β-Coronavirus (m-CoV) and its associated disease process.
Keywords
Introduction
The new coronavirus disease 2019 (COVID-19) is caused by a novel coronavirus, SARS-CoV-2. It is about 80% genetically identical to SARS-CoV and displays clinical symptoms similar to those reported in previous outbreaks of Coronaviruses (CoVs), namely, SARS-CoV and MERS-CoV.1–8 The severity of CoV infections depends on virus-mediated tissue damage as well as the antiviral immune inflammation, which are influenced by viral tropism, infectivity, virus spread, and specificity of host responses, all of which are majorly regulated by the CoV Spike (S) protein. S protein is a multidomain Class I viral fusion protein that exerts its effect stepwise, by a concerted action of individual domains.9–11 Considering the highly infectious nature of CoVs and their associated lethality and ability to emerge from zoonotic hosts, SARS-CoV-2 has once again conjured the entire scientific community in an urgency to develop effective therapeutics against CoVs.
The vast reservoir of research contributed by Corona virologists over the years where Mouse Hepatitis Virus (MHV), a group 2-β-CoV, has been an epicenter sheds light on the importance of S protein in controlling the pathogenic properties of CoV. Murine-CoV (m-CoV) infection in mice is the best-explored experimental animal model for studying respiratory, enteric, hepatic, as well as neurological illness because of their differential organ tropism like the distinct tropism showed by the strains of human-CoVs (H-CoVs).10, 12–21 However, it is difficult to explore the disease mechanisms in humans, and the generation of an animal model is necessary where the function of individual genes can be studied. In this context, discontinuous transcription process producing subgenomic mRNA in CoV, despite their enormous genome size, has proved advantageous for developing a reverse genetic system for CoVs wherein each protein could be targeted separately to understand their contribution to pathogenic properties.22–25 A battery of recombinant strains created by adopting a reverse genetics tool has largely demonstrated that the S protein is a major determinant of pathogenic properties associated with m-CoV.
S protein of SARS-CoV-2 and SARS-CoV share about 74% to 76% amino acid identity in their receptor-binding domain (RBD). Still, they show significant differences in their ability to infect and transmit in humans.2, 4, 26, 27 Both the SARS strains identify ACE2 receptor, SARS-CoV-2 binds to it with 10 to 20 folds more affinity.8,11,27–30 An interesting question that stems from this observation is, can S protein alone contribute to the higher virulence of SARS-CoV-2? A lack of a suitable animal model to study H-CoV pathology makes it challenging to answer this question. However, owing to the high similarity in recombination potential and replication kinetics of a fellow β -CoV, MHV, we can draw parallels between H-CoV and m-CoV S proteins to gain a better understanding of the unanswered questions related to virus entry, replication kinetics, pathogenesis, host immune responses, and virus persistence.5, 31–33 This review presents insights into the contribution of S protein in the resultant immune inflammation and pathogenesis based on the notable contributions over the last three decades with particular emphasis on the presence of two consecutive prolines in the fusion peptide of m-CoV Spike. Coronavirus induced cell-cell fusion mechanism offers a potential target for antiviral therapy. This review will discuss how knowledge from m-CoV studies holds lessons to understand SARS-CoV pathogenesis.
CoV Virions and Genome
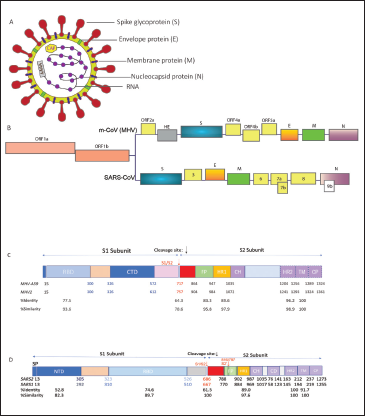
The family name Corona is because of the outward radial projection of S protein, which gives the virion a crown-like structure. CoV virions can be spherical or pleiomorphic, with typical sizes ranging from 60 to 200 nm.20, 34, 35 Their genome is an internal 5' capped single strand of helical RNA with a length of 26 to 32 kb, complexed with Nucleocapsid (N) protein encoded by gene seven in MHV and nine in SARS. The genome comprises six to ten open reading frames (ORFs) where the ORF1 forms 2/3rd of the genome coding for nonstructural polyproteins with helicase, polymerase, and replicase functions. The latter one-third of the genome encodes structural proteins in the order Hemagglutinin Esterase (HE), S, the envelope protein (E), transmembrane glycoprotein (M), and nucleocapsid phosphoprotein (N). ORF2 may or may not be functional in all strains of CoVs; in certain MHV strains it encodes a 65 kD HE protein that forms smaller spikes on the surface, but their function in the life cycle is not well understood. The genome is packaged within a lipid envelope containing S, E, and M encoded by genes 3, 5, and 6, respectively.20, 34, 35 The M and E proteins are essential for virus assembly and S protein functions in virus entry. Some CoV strains also have an Internal protein (I) of unknown function. 36 Figure 1A and B depict an overview of the virion structure and genome organization.
Spike Function in Attachment, Fusion, and Entry into the Host Cell
S protein is a large, multifunctional, highly glycosylated type I transmembrane protein incorporating 21 to 35 N-glycosylation sites.37–40 They assemble into trimers on the viral surface to form the characteristic “corona” and mediate virus-host attachment through the ectodomain. The ectodomain of all CoV S proteins share the same two domains: An N-terminal S1 domain responsible for binding the receptor on the host surface and a C-terminal S2 domain responsible for fusion of their envelope with the host cell membrane to deliver their nucleocapsid. 41 As a Class-I fusion protein, S protein has the typical α-helical secondary structure and contains characteristic two heptad repeats comprising of repetitive heptapeptide with hydrophobic residues in the S2 subunit that aid the formation of coiled-coil structure to participate in the fusion process. 9 The characteristic postfusion structures of the HR have been solved for SARS-CoV and m-CoV, they form the characteristic six-helix bundle. Their functional roles in MHV and SARS-CoV were confirmed by mutating key residues and by inhibition experiments using HR2 peptides.42–45
The S1 contains two subdomains, an N-terminal domain (NTD) and a C-terminal domain (CTD), both of which can function as receptor binding domains (RBDs) in different strains of CoVs.4,8,46–53 For example, MHV uses the S1 NTD as the RBD to bind to the carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1) in contrast to its close relatives SARS-CoVs and MERS-CoV who have the RBD in their S1 CTDs.8, 54–57 After the binding of the spike with the receptor, S2 triggers the fusion process either by entering the host cell directly or after internalization through endocytosis. 58 Additionally, S protein is known to become fusion competent by cleavage.58, 59 Some β- and all γ-CoVs spike have a furin cleavage site between the S1 and S2 domains, which is typically recognized and cleaved by a Golgi-resident host protease, Furin.7, 60, 61 Interestingly, within the β-genera, two closely related m-CoV species, MHV-2 and MHV-A59, display different cleavage requirements for infectivity. 62 MHV-A59 strain shows pH-dependent and independent fusion processes. 63 Whereas, in MHV-2, the entry is shown to be dependent on the endosomal proteases, Cathepsin B, and L that are activated at low pH. 64 In SARS-CoV also, the fusion process is dependent on Cathepsin L and the proteolytic processing of the spike in sequential cleavage events. 65 The requirement of endosomal proteases for efficient entry by MHV-2 and SARS-CoV could be attributed to their uncleaved spike.
Kathryn Holmes and Larry Sturman, 66 in very early studies, showed that MHV-A59 S protein is proteolytically cleaved by furin into S1 and S2 subunits at RRAHR (Arginine-Arginine-Alanine-Histidine-Arginine) during the virus attachment to the host membrane. This cleavage property was later shown to be essential for cell-cell fusion in MHV-A59 infection. Recent studies have shown that SARS-CoV-2 S protein also contains a furin recognition site between S1 and S2 subunits, which was absent in SARS-CoV. 11 However, SARS-CoV shows trypsin mediated sequential cleavage of Spike at two discrete locations. 67 The first cleavage (R667) takes place at the S1/S2 boundary by Cathepsin.64, 67 Later, another cleavage site (S2’) was recognized within the S2 domain as a second cleavage event at R797 that exposed the fusion peptide required for entry. Reports also suggest that a cleavage mediated by Elastase at T795, next to the fusion peptide, can significantly alter the cleavage activation in SARS-CoV. 68 Additionally, the S proteins of SARS-CoV use the transmembrane protease/serine subfamily member 2 (TMPRSS2) colocalized with ACE2 on the cell surface to enter the host cell.69, 70 TMPRSS2 family proteases are majorly found in the respiratory tract and lungs which can explain their tropism in the lung tissues. It is recently shown that SARS-CoV-2 entry also employs the TMPRSS2 activity and hence the similarity in tropism. 4 The two cleavage events, one by furin and second by TMPRSS2 might increase the infectivity of SARS-CoV-2. 26
Overview of m-CoVs and H-CoVs Belonging to the Genera β-Coronavirus
m-CoV, SARS-CoV, MERS-CoV, and SARS-CoV-2 belong to the family Coronaviridae, order Nidovirales, and genus β-coronavirus. The other three genera in the family Coronaviridae are α-coronavirus, γ-coronavirus, and δ-coronavirus. Among the four genera, α and β-coronaviruses have majorly emerged to cause several gastrointestinal, respiratory, hepatic, and CNS illness in mammals. Whereas γ-coronaviruses are known to infect only avian species, and δ-coronaviruses infect both mammalian and avian species. 10
Five naturally occurring strains of m-CoV have been identified and thoroughly studied. MHV infects many vertebrate hosts and induces several diseases oscillating in severity. MHV-1 causes respiratory disease. MHV-2 is purely hepatotropic and causes severe hepatitis and lethality, whereas MHV-3 is a neurotropic strain that results in hepatitis, vasculitis, initial ependymitis, meningitis, and encephalitis but not white matter lesions.15, 19, 71 Temperature-sensitive JHM strain causes severe encephalitis and demyelination but is unable to infect the liver and is highly lethal. It infects nonneuronal cells and has a special affinity toward astrocytes.21, 72 MHV-A59 is dual hepato-neurotropic.12, 19, 73
So far, seven humans CoVs have been identified. NL63 and 229E (α-CoVs) and OC43, HKU1 (β-CoVs) cause mild common cold symptoms whereas, SARS-CoV, MERS-CoV(β-CoVs) are known to cause acute respiratory syndrome and the newly emerged SARS-CoV-2 (β-CoV) with its high zoonotic potential causes severe acute respiratory syndrome associated with heightened lethality.1, 10 The periodical emergence of the new CoVs among humans can be attributed to their great genetic diversity, high pervasiveness, and recurrent genetic recombination. The risk of cross-species transmission increases further because of the increase in the human-animal interaction, thus enabling the viruses to select the most favorable receptors in the host. While m-CoVs are known to infect cells in the hepatic and central nervous system primarily, SARS-CoV2, SARS-CoV, and MERS-CoV primarily target the respiratory system and present with diffuse alveolar damage.3, 35, 74 Major clinical manifestations of SARS-CoV-2 causing COVID-19 are like the previous outbreaks, including cough, pneumonia, RNAemia, respiratory distress, cardiac injury, and the serious ground-glass opacities in both lungs preceding death.75, 76 Interestingly, SARS-CoV-2 rapidly translocates to the lower respiratory tract and other organs, including the liver, gastrointestinal tract, kidney, CNS, and cardiac muscles leading to multiorgan failure, all of which can be attributed to the appropriate choice of vastly expressed host receptor by the virus.4, 8, 30, 77
Comparison of S Protein Between m-CoV; MHV-A59 and MHV-2 and H-CoV; SARS-CoV and SARS-CoV-2
MHV-A59 (gene bank accession number 9629812, RefSeq: NC_001846.1) and MHV-2 (gene bank accession number: AF201929) show 94% to 98% sequence homology in their replicase genes, 83% to 95% sequence homology of genes 2a, 3, 5b, 6, and 7. However, considering the significant differences in their cleavage and fusion properties, both of which rely on S protein, the S gene identities were compared (Figure 1). As discussed above, the major difference between the two strains is cleavability and fusogenicty. MHV-2 lacks the furin cleavage site, unlike MHV-A59, and their cleavage depends on Cathepsins. Both strains’ cleavage signals showed a 64.3% sequence identity. Their fusion peptides are 83.3% identical. Like MHV-2, SARS-CoV (NCBI Accession: NP_828851.1) also depends on Cathepsins for their spike cleavage into S1 and S2. When compared with SARS-CoV-2 (NCBI Accession: QIK02964.1, RefSeq: YP_009724390.1), which shares a furin cleavage site similar to MHV-A59, they were 61.5% identical in their cleavage signals and 78.9% identical in the fusion peptide. We also compared the RBD, HR1, HR2, and TM domains; their respective identities and sequence similarities are denoted in Figure 1.
Corollary Between SARS-CoV-2 and m-CoV Disease Kinetics
Intracranial (i.c.) inoculation of MHV-A59 and RSA59 in mice is used as extensively studied prototypic animal model to study virus-induced inflammation of the brain, spinal cord, optic nerve, and retinal ganglionic cells of the eye. Acute disease symptoms during three to seven days postinfection (p.i.) are characterized by robust host immune responses accompanied by pathological consequences including meningitis, encephalitis, and myelitis with or without hepatitis. Infectious viral particles are cleared within the first 10 to 14 days; however, at this time, mice begin to develop demyelination, either clinical or accompanied by chronic hind limb paralysis. Demyelination and consecutive axonal loss reach its peak at day 30 p.i. when the infectious particle is completely cleared, and innate immune inflammation is majorly resolved, this represents the second or chronic disease phase. On day 30 p.i., only viral RNA persists at a low level in the spinal cord.18, 19, 78 An intermediate stage may be present, which is governed by the continuous infiltrating T cells into the CNS and helps connect the acute host-inflammatory responses with the chronic anti-inflammatory responses. This phase, however, is less explored.
Likewise, SARS-CoV-2 infection is divided into three phases. 79 The first phase is represented by the initial infection of the mucosal membranes and respiratory tract and dissemination of virus into the peripheral blood. During the second pneumonia phase, the virus replicates profusely and causes an upregulation of host-inflammatory responses. If the antiviral immune response is proficiently active, and the virus replication is reduced, the patient enters the final recovery phase; otherwise, the uncontrolled cytokine storm results in lethality.28, 80 Results from case reports have shown that the timely detection of virus RNA is crucial as the viral titer lies within the detection limit only for a short duration of time, i.e., in between the first two phases and drops down drastically by the second week.79, 80 Figure 2 depicts the characteristic neuroinflammatory pathology induced by m-CoV MHV-A59, and a comparative schematic of viral replication with disease kinetics of m-CoV and SARS-CoV-2.
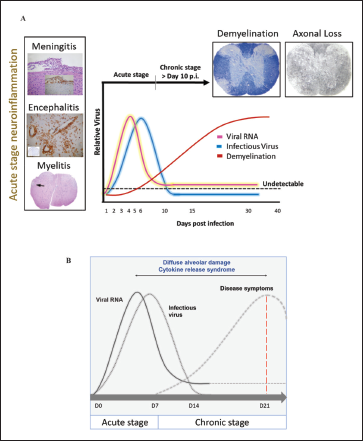
The studies we have at our disposal about SARS-CoV-2 infection mostly tell us about the disease kinetics extending up to 21 days. However, m-CoV has significantly shed light on the fact that major disabling chronic pathology of demyelination and axonal loss associated with MHV-A59 and RSA59 reaches its peak rather silently when the infectious viral particles go below the detection limit, acute inflammation is resolved, and only viral RNA persists at a low level. 81 So, the question is, do the same phenomena exist in the patients recovered from COVID-19? Though it is too early to comment on cases of COVID-19 patients, mounting evidence indicates a possibility. A study showed that patients affected with SARS-CoV-2 presented with neurological symptoms like headache (8%) and confusion (9%). 82 Also, several studies have shown that SARS-CoV and MERS-CoV can infect and replicate in the CNS of both humans and animal models, especially the brainstem.3,74,83–86 While many animal models of SARS-CoV were tested, none were able to mimic the disease symptoms like humans, but MHV-1 produced a SARS-CoV like lethal disease in mouse and has provided few insights into the host cytokine responses against the virus. 87 Still, there is a shortage of studies directly correlating the replication of the virus with the resultant pathology as the disease caused by both SARS-CoV in humans and MHV-1 in mice are milder than SARS-CoV-2. Moreover, the neurological manifestations of SARS-CoV-2 infection is finding more relevance by the day and therefore the consequences of SARS-CoV-2 infection can be analyzed from the eye of m-CoV, MHV-A59 to uncover aspects of disease pathology that may be instrumental in designing therapeutic strategies.
Spike Controls m-CoV Pathogenesis
While a vast range of studies was identifying and emphasizing on the spike function in infection and associated pathology, less was clear about the role of S protein in CoVviral antigen spread, cell-cell spread, and associated pathogenesis. The study on natural recombinants obtained by co-infection of MHV-2 and LA-7 strains opened avenues for RNA recombination, where distinct mutations could be linked to altered pathologies, if any, in a measure to understand the role of distinct spike domains in the disease outcome.
88
Paul Masters and group developed the first reverse genetic system for MHV using RNA recombination.25, 89 This has been extensively used to exchange specifically the structural and nonstructural genes among the MHV strains and to introduce single amino acid substitutions.
90
The large genome size of the CoVs made it difficult to use the reverse genetics approach as a tool to design the whole genome infectious cDNA clone. Still, in the last few years, full-length infectious clones for many coronaviruses, including MHV, TGEV (transmissible gastroenteritis coronavirus
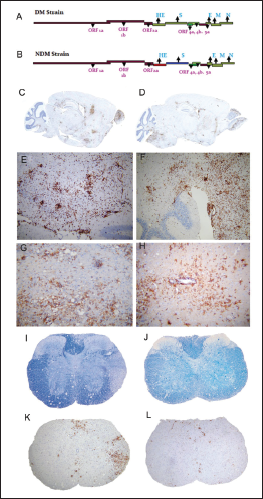
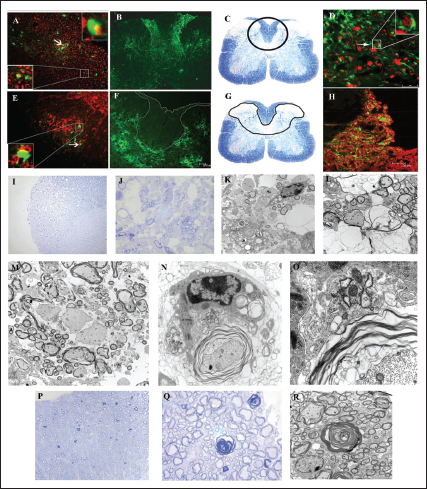
Using targeted RNA recombination, two isogenic recombinants of MHV, RSA59, and RSMHV2 (background is from demyelinating strain MHV-A59) that differed only in the S gene were generated. RSA59 had the spike from MHV-A59, and RSMHV2 had the spike from the parental nondemyelinating MHV-2 strain.96, 97 The replication, virulence, and spread of the recombinants were like the parental spike strains (Figure 3).81, 96, 98 These recombinant viruses have served as suitable tools to study the role of Spike mediated pathogenesis which has been thoroughly reviewed previously.18, 19 From the perspective of neuropathogenesis, the major highlights from the comparative studies are that spike gene mediated entry in the brain or the induction of encephalitis may not be the restricting factors for causing demyelination as both strains induced acute stage neuroinflammation but only RSA59 caused chronic stage demyelination. Other functions of spike like cell tropism, virus transport along the axons associated with virus spread, fusion, and persistence may attribute to the differential chronic stage pathologies between the two strains. RSA59 takes advantage of the synaptically linked neuronal and glial cells circuits to travel from the neuronal body centripetally along the axons to spread from brain to the brainstem during day 3 p.i. It reaches the spinal cord grey matter by day 5 upon releasing at the nerve endings and quickly translocate to the white matter microglia and oligodendrocytes by day 7 p.i. as a function of its cell-cell fusion (Figure 4).99, 100, 101, 102 RSA59 was also capable of retrograde axonal transport from the optic nerve into the retinal ganglionic cells and caused optic nerve inflammation and demyelination. Additionally, RSA59 induces a persistent activation of CD11b+ microglia/macrophages in the demyelinating regions which showed a mechanism of macrophage-mediated myelin stripping in RSA59 infection (Figure 4).99, 19 RSMHV2 and RSA59 also differ in their cleavage signal site. As mentioned before, RSA59 S is cleaved posttranslationally into S1 and S2 subunits, but RSMHV2 S protein does not and is unable to cause fusion in vitro.103, 104 Penn 97-1, a recombinant strain with S1 from demyelinating strain MHV-A59 and S2 domain from MHV-2 was nonfusogenic and did not induce demyelination, demonstrating the role of S2 domain in the fusogenic property of MHV-A59.16, 105 Thus, variable regions of the S2 domain or differences in the cleavage signal site and fusion domain between RSA59 and RSMHV2 could be candidates to explain the differential axonal transport and demyelination potential.
Delineating the Minimum Essential Motif in the S2 Domain of Spike Protein Responsible for Fusogenicity and Demyelination
The strategy for inhibiting the fusion of CoV S2 domain is based on the current understanding of the conformational transition that takes place during the fusion event. 34 The basis for this transition is the formation of the six-helix bundle viral fusion core driven by hydrophobic interactions between HR1 and HR2. 9 Thus, all efforts of researchers have been directed in disrupting the formation of the fusion core, and this has been facilitated by the fact that HR regions are the better-conserved regions of the S protein and more amenable for peptide design. The identification of the double-proline site opens a new possibility of mimetic peptide design with the potential to disrupt Spike fusogenicity.
In 1990, for the first time, it was indicated that MHV-A59 contains an internal FP (929–944) that could be the fusion domain considering its hydrophobicity and location adjacent to the heptad repeat domains. 106 Previous studies had shown that substitution of 936 methionine residue with lysine (M936K) or leucine (M936L) in the fusion domain did not affect fusion, but the substitution of 938 proline with lysine (P938K) partially impaired the fusogenicity. 107 The fusion domain of MHV-A59 contains two consecutive central proline (938, 939), but that of the MHV-2 (nonfusogenic and nondemyelinating) strain contains only one proline (976). A series of 3-dimensional in silico and biophysical nuclear magnetic resonance (NMR) studies were carried out on MHV-2 and MHV-A59 Spike fusion domains in their wild-type and mutant states which revealed that the fusion peptide might contribute in a significant way toward the fusogenicity of MHV-A59(Figure 5). 62 The double proline fusion peptides were found to be markedly rigid than single proline. The presence of two proline in the Spike of MERS-CoV, SARS-CoV and HCoV-HKU1 were also shown to stabilize the S protein in the prefusion state. 108 Recently, as a proof of concept for the use of S-2P as a vaccine for SARS-CoV-2 it was shown that an mRNA vaccine expressing the MERS S-2P elicits a protective response in mice. Similarly, SARS-CoV-2 S-2P expressing mRNA served as a potent immunogen inducing both neutralizing activity and CD8 T cell responses in mice. It protects mice from both upper and lower airway SARS-CoV-2 infections. The mRNA vaccine has shown promising results in phase II clinical trials and is soon expected to be in the phase III efficacy evaluation phase. 109 The S-2P design has been used in several vaccine strategies including the
95% effective, BNT162b2 by BioNTech/Pfizerand 94% effective mRNA1273 by Moderna/NIAID, the first two vaccines approved by the FDA upon exercising their emergency authority protocols.110–112
In earlier studies, the role of two Proline infusogenicity was demonstrated by generating a mutant by targeted RNA recombination, RSA59 (P), which had one deleted proline. RSA59 (PP), i.e., the wild-type recombinant MHV-A59 strain and RSA59 (P), were compared in a series of in vitro and in vivo experiments. RSA59(P) had significantly reduced fusion and syncytia formation. 62 Both the PP ad P mutants have the same cytoplasmic tail which has the retention signal for controlling protein trafficking, therefore, RSA59 (P) slower replication and syncytia formation could not be only linked to the movement of S Protein to the cell surface but a combinatorial outcome of low replication, destabilized S protein structure and inefficient trafficking (Figure 5).62, 113 RSA59 (P) showed limited ability to spread across the brain parenchyma; viral antigen staining was mostly restricted to the site of infection and in the meninges and it induced significantly low demyelination pathology compared to the wild-type. Additionally, RSA59(P) showed significantly less inflammation of the optic nerve and induced no damage to the RGC. The virus did transport retrogradely, but its reduced ability toward cell-cell fusion did not allow it to move until the retina and infect RGCs. 114 The proline thus plays an essential role in fusion, which further affects the replication, spread across the brain and spinal cord, retrograde transport to the optic nerve and RGC, and the ability to cause demyelination.
These reports, for the first-time, unraveled the role of two proline in the backbone of S protein fusion peptide in m-CoV pathogenesis. M-CoV fusion mechanism offers a potential therapeutic target. Dissecting the minimum essential required for fusogenicity is a significant advancement for designing a mimetic peptide to restrict CoV infection.
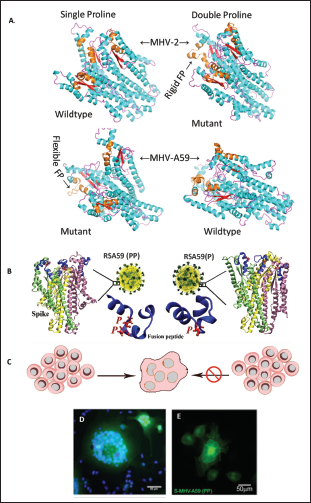
Two Consecutive Prolines in the Centre of the Fusion Peptide of m-CoV Regulate Fusogenicity. (A) Diagram Showing the Illustration of MHV-2 and MHV-A59 Spike Fusion Domain Structures in the Trimeric Quaternary State. Frames From a 1µS Long Molecular Dynamics Simulation Trajectory were Randomly Drawn to Depict the Shown Structures. The Fusion Peptides are Marked in Brown. The Single Proline Containing Fusion Peptides Showed more Flexibility than the Double Proline Cases in the Trajectory. (B) Shows the Illustration of Spike of RSA59 (PP) Wildtype Recombinant and RSA59 (P) Single Proline Mutant of RSA59. Only RSA59(PP) can Form Syncytia (C, D). Spike Alone can Traffic to the Cell Surface and Cause Syncytia Formation was Shown by the Transient Overexpression of Spike of MHV-A59 (S-MHV-A59 [PP]) in HeLa Cells (E) by YFP.
S2 and Not S1 Domain of Spike Can Serve as Better Pan-CoV Drug Target
Despite considerable progress in the understanding of spike protein functions, it may still be limited in our ability to predict the effect of mutations that Spike, and the virus genome might acquire. Therefore, insights from decade-long studies on other CoVs and understandings from them hold a compelling case for improving our understanding of the current COVID-19 pandemic and assess its pathology and course of evolution.
Given the rapid expected evolution of the SARS-CoV-2, it is imperative to ask which of its two domains, S1 or S2, is likely to evolve faster. This can be answered from sequence comparisons with other existing members of the β-coronaviruses. The receptor-binding domain is most variable across the CoVs, and this has also been corroborated by the limited success we have achieved in drugs’ discovery against the receptor-binding domain as target.10, 11 Antibodies capable of recognizing the SARS-CoV receptor binding domain have failed to recognize the SARS-CoV-2 receptor binding domain, although they share >74% identity, thereby showing low cross-reactivity. 30 In comparison, the fusion domain (S2) has more conserved regions, and studies in m-CoV have demonstrated the importance of specific residues, including our illustration of the high significance of two prolines in the fusion peptide and Spike associated disease pathology.62, 114–116 The fusion domain thus promises to be a better cross-functional success as the potential pan-coronavirus therapeutic target. However, the receptor-binding domain contains most antigenic determinants because of its larger surface exposure in the prefusion state. Another aspect of Spike that strengthens our premise for S2 as a therapeutic target is based on the profound structural and functional similarities it shares within all class I viral membrane fusion proteins. In all cases, S2 is in a metastable state that exists in prefusion and postfusion conformations. The prefusion structures are similarly triggered for state change through comparable conformational rearrangement, folding to a highly similar six-helix bundle post-fusion structure with exposed fusion peptides. The complexity and intricacy of the fusion mechanism appear to be specialized and inherited, although its independent evolution in viruses cannot be completely ruled out. Spike evolution is justifiable; therefore, an intriguing question because a Spike from neurotropic strain MHV-JHM exists that can mediate receptor-independent virus entry into cells that do not express its receptor. 117 This suggests that receptor binding need not be essential for a virus to fuse, infect, and replicate, and its sole purpose is cell-recognition. However, such a Spike with S2 alone may be inefficient for the fusion.
It has been shown that the HR1/HR2 regions of class I viral fusion proteins of enveloped viruses like HIV, respiratory syncytial virus (RSV), 21 Ebola virus, 100 paramyxoviruses SV5, 101 Nipah virus, 22 and murine hepatitis virus (MHV) 23 serve as efficient drug targets. Likewise, the heptad region in the spike S2 domain is highly conserved and peptide sequences based on these segments have been most tested for inhibiting CoVs. During SARS-CoV 2003 outbreak, three independently developed peptides based on the heptad repeat two region were found to inhibit CoV fusion.118–120 Later another three peptides based on the same region were specifically found to block the six-helix bundle formation. 121 SARS-CoV-2 not only binds to the ACE2 receptor with higher affinity, but it also displays a significantly higher capability of membrane fusion compared to SARS-CoV which is elucidated by the formation of syncytium in culture. Thus, suggesting that the fusion machinery of SARS-CoV-2 can serve as an important drug target. This could be partly attributed to the presence of few alterations in the SARS-CoV-2 HR1 region. The HR2 peptide sequence is identical in both SARS-CoV and SARS-CoV-2 and the mutations in HR1 might have resulted in rendering the virus fusogenic. Recent study showed that peptides targeted against the HR1 domain were effective against SARS-CoV-2 and other H-CoVs. 29 Nonheptad repeats regions-based peptides have also been developed for fusion inhibition study, albeit to a limited degree. Two designed peptides upstream of the S1/S2 and S2’ cleavage sites have elicited anti-fusogenic activity likely because of interference with S1/S2 cleavage and conformational restriction for fusion. 121 Our discovery of surface-exposed double proline containing rigid segments in the S2 domain could offer a new premise for structure-based peptide design for CoV fusion inhibition. These segments could also be presented as antigens to raise antibodies and, therefore, also suitable for vaccine design. There is a need to do clinical tests using few peptides discovered – for proof of pan-COVID utility.
m-CoV Spike in Inflammation and Immunity
The integration of reverse genetics, molecular biology, and pathology has significantly contributed toward the understanding of S protein function in CoV biology, to the extent of deciphering a minimum essential motif required for entry and fusogenicity. However, Spike function is not limited to mediating entry into the cell, and it is known to regulate immunopathogenesis, which is the critical regulator of virus infection.18, 19, 26 In fact, the interaction between the virus and the innate immune system determines the outcome of the disease. While the early control of virus replication elicits a strong pro-inflammatory, Type I interferon response, it also helps promote the development of an anti-inflammatory response.122–124 The balance between these two contrasting yet complementary states of immunity is central for reinstating tissue homeostasis.125, 126 A shift toward either side can be detrimental and result in immune pathology and tissue damage.
MHV infection causes neuroinflammation as a result of pronounced activation of CNS resident immune cells, microglia, and astrocytes.72, 127 Microglia, upon activation, take the characteristic activated phenotype and start expressing microglia/macrophage-specific protein Iba1(ionized calcium-binding adaptor molecule 1), which promotes ruffling and phagocytosis.99, 19, 128, 129 Both RSA59 and RSMHV2 trigger innate immune responses predominated by chemokines like CXCL10, CXCL9, CCL5, and CCL12 and CD molecules during the acute stage. Antiviral host responses are associated with perforins and genes involved in interferon (IFN) gamma signaling. The inflammatory responses gradually decline in RSMHV2 infection after virus clearance. However, RSA59 chronic disease is represented with persistent CD11b+ microglia within the demyelinating lesions, and the production of microglia-associated inflammatory mediators. These differences could be attributed to the ability of RSA59 to translocate through the axons and evade the robust immune responses targeted to clear the virus completely. However, parallelly, regulatory, and anti-inflammatory cytokine IL-10 significantly controls the enlargement of tissue lesions in an attempt to reduce the chronic pathology.130–133 IFN responses can promote phagolysosomes maturation and autophagy in the persistently activated microglia/macrophages, which can engulf the myelin sheath, leading to demyelination. 99
Further, virus-specific T cell effector functions are essential to eliminate the infectious virus load during most acute infections.134–136 Control of m-CoV spread requires the functioning of both CD4 and CD8 T cells, were CD8 T cells are the primary effectors but require support functions from CD4 T cells.20, 137 A recent study in CD4-/- mice showed impaired virus clearance, despite the presence of functional CD8 T cells, demonstrating the importance of CD4 T cells for the efficient functioning of CD8 T responses. 138 While T cells are the helpers for the development of intact adaptive responses, their functions are acutely regulated by CNS resident immune cells to reduce their cytolytic effects. 139 However, a deviation from the optimum control of CNS cells can significantly dampen the T cell functions that may sometimes result in the establishment of persistent virus infection. 139 In a recent study, we have shown a critical role of CD4 T cells in RSA59 clearance and resultant pathology. The demyelination pathology is significantly more severe in the CD4-/- mice where phagocytic M2 phenotypic microglia/macrophages are found in abundance in the demyelinating plaques.
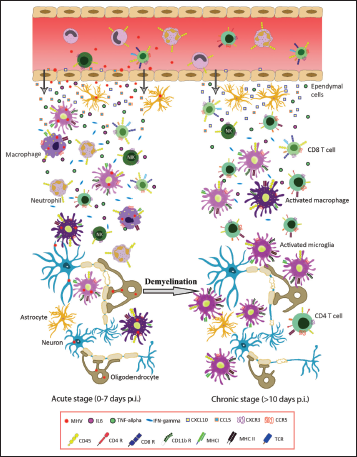
The studies on Spike mutants have served as a valuable tool in the elucidation of the Spike mediated mechanisms of intracellular spread and the processes that specifically allow CoVs to evade the immune system to establish successful disease pathologies. Figure 6 presents an illustration of the kinetics of immune cell infiltration and pathogenesis upon RSA59 infection.
Insights From m-CoV Help to Understand the Underlying Mechanisms of Immune Inflammation in SARS-CoV-2
As the infection kinetics of MHV-A59 and SARS-CoV-2 are quite similar, it is not surprising to note that the immune induction is also similar in the two strains based on the current literature.140, 141 SARS-CoV-2 infection initially replicates in the airway epithelium and induces virus-associated pyroptosis in the infected cells.1, 140–142 This highly inflammatory type of cell death program triggers a whole chain of pro-inflammatory responses like m-CoV, starting with IL1β and other chemokines secretion, which is followed by the infiltration of activated macrophages and establishment of a local inflammatory niche.28, 77, 141 Many cytokines and chemokines, mostly of the TH1 response, are identified in the plasma of the patients suffering COVID-19 during the acute infection, including, but not limited to, IL6, IFNγ, MCP1, and IP-10.141, 143 In most individuals, these innate immune responses clear the virus, and patients recover. But dysregulation of these tightly controlled host responses can trigger a cytokine release storm (CRS) which is represented by increasing levels of IL-6, IL-2, IL-7, IL-10, granulocyte colony-stimulating factor (G-CSF), IP-10, MCP1, macrophage inflammatory protein 1α (MIP1α), and tumor necrosis factor (TNF). 28 Inflammatory monocytes derived activated macrophages persist in the lung tissues like the activated glial cell persistence in RSA59 infection that secrete significant levels of cytokines like MCP1, IP-10, and MIP1α. 144
Interestingly, the adaptive immune system, including CD4 T cells and B cell responses, occurs concomitantly after the first stage of the infection, i.e., after the first week in SARS-CoV-2, as is also seen in m-CoV.145, 146 While CD8 T cells are required for direct killing of the virus, CD4 T cells are essential for activating both CD8 and B cells as well as cytokine production. Not much is known about the role of CD4 T cells in the COVID-19 pathology. Still, there are pieces of evidence from case reports which show the presence of T cells in the lungs and lymphopenia, indicating their putative role in the attempt to prevent tissue damage.141, 147–149
The main reason behind the extreme morbidity associated with COVID-19 is the imbalance between this see-saw where proinflammation is so high it outweighs the protective anti-inflammatory response. It has been demonstrated in SARS-CoV that virus infection downregulates the expression of ACE2, which has shown to play essential roles in lung injury and the regulation of the renin-angiotensin system (RAS).75, 150 RAS dysregulation disturbs the blood pressure and fluid/electrolyte balance and induces a heightened inflammatory response along with increased vascular permeability. Now, SARS-CoV-2's binding to the receptor is 10 to 20 folds higher, and it is not unlikely to hypothesize that it causes a much more significant downregulation in ACE2, which results in the inefficiency of RAS and dysregulation between the pro and anti-inflammatory responses which results in a cytokine storm.27–30 This highly pro-inflammatory microenvironment and permeable vasculature invite an ever-increasing number of inflammatory cells into the lungs, which ultimately clogs the airways resulting in lethality. SARS-CoV emerged, caused a few hundred deaths, and disappeared, but it seems that SARS-CoV-2 is here to stay. The efficiency of human-human transmission and an enormous virus load is still beyond the control and scope of the current database of knowledge that has been generated in years. JHMV, a highly neurovirulent CoV, shows similar uncontrolled acute inflammation state, whereas less as 10PFU of virus creates havoc in the CNS of the mice so much so that results in the death of mice at ten days p.i.5, 151, 152 Though this virus and its recombinant strains have generated enormous amounts of results to understand CoV biology, it is considered unsuitable for the study of virus persistence. MHV-2, on the other hand, triggers a robust innate immunity in the acute stage, which helps clear the virus and bring back homeostasis. 129 It does not cause any kind of progressive illness. MHV-A59, because of its mild nature has proven to be an appropriate model to study the development of a protective host response that efficiently balances the innate antiviral responses at the same time, the virus persists in the spinal cord, silently causing chronic pathology without any visible symptoms. SARS-CoV was reported to infect the brains of patients, especially the neurons, and upon intranasal inoculation in animal models could also infect the thalamus and the brain stem regions of the brain.3, 153–156 In fact, another study showed MERS-CoV tropism only in the brainstem upon low dose intranasal inoculation, indicating that the infection in the brain and not lungs led to significant morbidity in mice. 157 The mechanism of the CNS invasion is not clear, but they probably take the synaptic route, as shown in other CoVs.101, 86 A neurotropic propensity is common to many CoVs.155, 158, 159 Furthermore, some CoVs can spread trans-synaptically to the brain from the mechanoreceptors and chemoreceptors in the lung and lower respiratory airways. 160 As SARS-CoV and SARS-CoV-2, are quite similar and specific neurological symptoms have been demonstrated in COVID-19 patients, the potential invasion of SARS-CoV2 in the brain might probably lead to the acute respiratory failure of patients.160–165
COVID induced by m-CoV and H-CoV share similarities in disease, including acute immune responses, virus persistence, and the development of adaptive protective immunity. Their comparative study to investigate therapeutics is an open forum.
Discussion and Conclusions
SARS-CoV-2 has infected millions of people worldwide to date, implying that it has presented itself with a golden opportunity to diversify further. 166 With a low fidelity of the virally encoded RNA-dependent-RNA-polymerase, 167 its large size of genome and replication strategy is a boon for frequent homologous recombination. This process enables the exchange of genetic material during co-infection. Persistent infection can further lead to the accumulation of adaptive mutations. Therefore, there is a high possibility of the virus becoming more lethal and jump to more species and cause more devastation. In such a scenario arresting Spike mediated fusion, either during virus-host attachment or cell-cell spread, can be the key to early containment of infection. Else because of the significant role of the spike in tissue tropism, its modification through evolution would mean altered cell and tissue tropism, and additional association with other viral and host factors may accelerate the alteration of virus pathogenicity. Such rapid change may overwhelm our capacity for surveillance. Recent studies showed that SARS-CoV-2 is actively evolving. D614 G mutation within the viral spike protein (S1 CTD) has rendered the virus more infective than the wild-type virus, though virus’s response to antibodies was unaltered.168–171 While this ensures that vaccines currently under development can be effective against the new strain, it also raises serious concern to address the COVID-19 pandemic and future coronaviruses, by investigating their entry and fusion mechanisms in consort, which can guide new therapeutic efforts. Many other aspects of the viral fusion reaction are also potential targets for modulation. These include the inhibition of the proteases that cleave the Spike, small molecules, or lipids that alter the lipid composition or ionic environment and antibodies that recognize the S domains. The research community is poised to see what novel therapeutics can address CoV infections.
Several treatment options for COVID-19 are emerging at an accelerated rate. Many vaccines candidates have been developed. Some have even cleared the trials and approved by the FDA. The first two mRNA vaccines delivered by nanoparticle showed exceptional results so far. 171 A third adenovirus-based vaccine 172 from United Kingdom also made to the market and is showing 91% effectivity. While vaccine development is the need of the hour to control COVID-19 pandemic and return to the prepandemic normal state, our long-term goal should be addressing the pan-COVID question and that demands the development of effective broad-spectrum therapeutics which cannot be achieved without considering in detail the host immune response. It is becoming increasingly evident from the current data on SARS-CoV-2 that host responses contribute a great deal to the morbidity. Therefore, we need to consider the host responses alongside Spike functions to be fully prepared for future outbreaks. Cellular factors like the production of reactive oxygen species have been associated with many virus infections, including CoVs. An imbalance between reactive oxygen species and antioxidants’ generation leads to a dysregulation of oxidative stress, endoplasmic reticulum stress and unfolded protein response pathways. 173 Deprivation of antioxidant mechanisms and oxidative damage to the tissues is relevant to the aging process, which could be one of the reasons for making the aged more susceptible to SARS-CoV-2 infection.2,4,77,160,174,175 Also, nonstructural proteins have shown to affect tropism and pathogenesis by regulating the rate of virus replication either by interacting with cell type-specific factors or with components of the immune response.176–181 But Spike alone can as well alter fusion, entry, cell-cell fusion, pathogenesis, host immune response, and virus persistence. This was corroborated from an excellent mouse model system of RSA59/RSMHV2 infection, demonstrating demyelination pathology because of either pathogenic immune outcome, direct virus-induced cytopathic effects, or their combined outcome for RSA59, and weak effect for RSMHV2, the strains differing only in their Spike. When immunity clears the virus and prevents subsequent pathology, it also supports a paradigm where cell-mediated immunity affects pathology indicated by microglia-mediated myelin stripping and the inability of T cell effector functions to completely clear the virus which results in persistently active immune responses.18, 19, 138, 182 This establishes that inflammation is a double-edged sword and provides useful clues for hypothesizing SAR-CoV-2 immune response. The unique nature of S glycoprotein suggests a need to focus on identifying the domains of Spike that interact with the specific host cellular pathways and use the information for therapeutic intervention. Health threats from CoVs are constant and understanding the antiviral host responses targeted against Spike is essential for controlling the virus with a specific goal to preserve global health and establish economic stability.
Footnotes
Acknowledgments
m-CoV research in an experimental animal model has provided a timely and appropriate platform to take a step toward writing a comprehensive review of the importance of S protein in COVID biology. This work has been a collaborative effort for two decades between the Indian Institute of Science Education and Research Kolkata (IISER-K), India; Indian Institute of Sciences (IISc), India; University of Pennsylvania, USA; Thomas Jefferson University, USA and University of Colorado Denver, USA. The joint endeavor has been supported and nurtured by the Indo-U.S. Science and Technology Forum (IUSSTF) on several occasions. M-CoV research was generously funded by the Department of Biotechnology (DBT), India, Council for Scientific and Industrial Research (CSIR), India; National Multiple Sclerosis Society, USA; M. E. Groff Surgical Medical Research and Education Charitable Trust; and Lindback Foundation Career Enhancement Award, USA. The authors would like to convey their special gratitude to Dr Manmeet Singh, Mr Saurav Saswat Rout, Ms Debanjana Chakravarty, Ms Lucky Sarkar, and Dr Rahul Basu for their significant contribution toward understanding the role of Spike fusogenicity in the progression of neuropathogenesis. The authors would also like to express their gratefulness to Dr Dhriti Chatterjee, Dr Kaushiki Biswas, Dr Abhinoy Kishore, Dr Subhajit Das Sarma, Dr Afaq Hussain, Mr Soumya Kundu, Mr Abhishek Bose, Mr Sourodip Sengupta, Ms Mithila Kamble, Ms Vaishali Mulchandani, Mr Saurav, Mr Safiriyu Abass A, Dr Mahua Maulik, and Ms Soma Nag who have contributed immensely to the generation of a large volume of work on m-CoV research over the years. Authors would like to thank Dr Kenneth S Shindler, Dr Lawrence C Kenyon, and Dr Randall J Cohrs for their long-standing collaboration on m-CoV research. The success of this work has been primarily dependent on the collaborative efforts of the national and international m-COVID team and many researchers whose names are listed in the references.
Authors’ Contribution
This review was conceptualized by JDS. Literature survey was conducted by FS in consultation with JDS. FS and JDS wrote the original manuscript where DP contributed significantly to reviewing the in silico part of the review.
Statement of Ethics
Most of the m-CoV work discussed in this review either adopted from our published work or from other studies adhered to the experimental procedures and animal care and use in accordance with good animal ethics approved by the Institutional Animal Care and use Committee.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
