Abstract
Background: Exposure to adverse environmental conditions such as toxic chemicals, viral infections, and even stress during pregnancy or early life may disrupt the development of normal brain and its functioning leading to incidence of neurodevelopmental disorders at later stages of life. Recently, we reported that poly (I:C) exposure altered synaptic plasticity protein level and impaired memory through activation of microglia cells.
Purpose: As epigenetic modifications are involved in memory formation, we have studied methylation of DNA and acetylation of histone at promoters of synaptic plasticity genes in the brain of rats exposed to poly (I:C) during early life.
Methods: One dose of poly (I:C) (5 mg/kg bw) was intraperitoneally injected to rat pups on postnatal seventh day. A set of pups exposed to vehicle was included as control. In order to assess methylation of DNA and acetylation of histone at synaptic plasticity gene promoter, we performed qPCR after methylated DNA immunoprecipitation and chromatin immunoprecipitation.
Results: Poly (I:C) exposure reduced the level of 5-methylcytosine (5mC) at synaptic plasticity gene (
Conclusion: Poly (I:C) exposure altered 5mC and H3K9Ac at synaptic plasticity gene promoters resulting in memory impairment of rats at later life.
Keywords
Abbreviation
AD, Alzheimer’s disease; ARC, activity-regulated cytoskeleton-associated protein; BDNF, brain derived neurotrophic factor; ChIP, chromatin immunoprecipitation; DNMTs, DNA methyltransferases; EGR1, early growth response 1; FC, frontal cortex; HATs, histone acetyl transferases; HDACs, histone deacetylases; HP, hippocampus 5-mC 5-methyl cytosine; MeDIP, methylated DNA immunoprecipitation; NARP, neuro activity regulated pentaxin; PND, postnatal day; Poly (I:C), polyinosinic-polycytidylic acid; qPCR, quantitative polymerase chain reaction
Introduction
Brain development is a tightly regulated programmed process, which includes cellular events like proliferation, migration, differentiation, and synapse formation. 1 Prenatal exposure to adverse environmental conditions like toxic chemicals, viral infections, and even maternal stress during pregnancy negatively affects brain development and its function leading to neurodevelopmental disorders such as autism and schizophrenia in late life. 2 Our previous study showed impairment of learning and memory through activation of microglia and alteration in the level of expression of synaptic plasticity related proteins (BDNF, Arc, EGR1) in the frontal cortex (FC) and hippocampus (HP) of rats exposed to poly (I:C). 3 Highly activated microglia produce inflammatory cytokines and induce neuronal apoptosis. 4 In addition, poly (I:C) exposure triggers the apoptosis signaling leading to degeneration of neurons and subsequently deficits in cognitive function including memory. 5 On the other hand, it is reported that viral infection alters 5mC and histone acetylation, which in turn affects gene expression leading to neurodevelopmental disorders.6–9 Singh and Thakur 10 have also shown that variation in epigenetic modifications changes the synaptic plasticity gene expression in cerebral cortex as well as HP of aged mice.
Methylation of DNA and acetylation of histone play a major role in regulation of chromatin remodeling, gene expression, and memory formation.11–13 They have also important functions in normal aging and associated neurodegenerative diseases. 14 For DNA methylation, DNA methyltransferases (DNMTs) transfer methyl groups to fifth carbon of cytosine at CpG sites. In case of histone acetylation, histone transferases (HATs) transfer acetyl groups to histone tails and histone deacetylases (HDACs) remove them. 15 These epigenetic modification machineries are sensitive to environmental stimulus. 16 Earlier reports have shown that poly (I:C) alters methylation of DNA and acetylation of histones in brain as well as immune cells.17, 18
These evidences advocate that poly (I:C) exposure during early life might alter the expression of synaptic genes through modulation of epigenetic modifications and lead to memory deficits. This has prompted us to analyze methylation of DNA and acetylation of histone at synaptic plasticity gene promoters in the FC as well as HP of 3-, 6-, and 12-week rats.
Methods
Animals and Poly (I:C) Injection
Wistar rats were maintained in animal colony at School of Studies in Neuroscience, Jiwaji University, Gwalior. Poly (I:C) was administered to rat pups on postnatal day (PND) seven as described previously. 3 All the experiments were performed according to the instructions of Committee for the Purpose of Control and Supervision of Experiments on Animals (CPCSEA), and approval of the Institutional Animal Ethical Committee, Jiwaji University, Gwalior.
The schematic timeline for drug administration and other molecular experiments is presented in Figure 1.
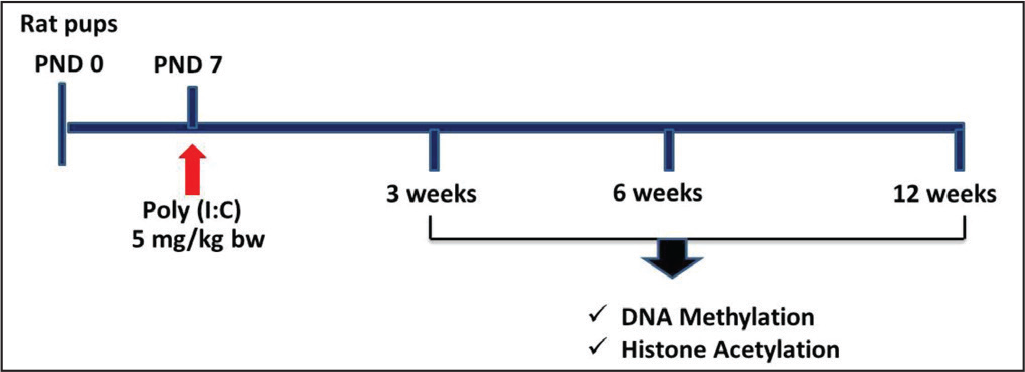
Analysis of Epigenetic Modifications
Following poly (I:C) administration, rats were sacrificed at different age of 3-, 6-, and 12-week and FC and HP were isolated and used for the analysis of epigenetic modifications.
Methylated DNA Immunoprecipitation (MeDIP)
The levels of 5mC at
Chromatin Immunoprecipitation (ChIP)
ChIP was performed to analyze H3K9 acetylation at histone bound to gene promoter sequence. Briefly, the FC and HP tissues were removed from the brain of experimental and control rats. Then tissues were chopped into small pieces in ice cold 1 × PBS and crosslinked with 1% formaldehyde at 25°C for 15 min. Thereafter, crosslinked tissues were homogenized in lysis buffer (5 mM KOH pH 8.0, 85 mM KCl, 0.5% NP-40 with 1 mM protease inhibitors), kept at 4°C for 5 min, and spun at 1,000 ×
Statistical Analysis
The data obtained from MeDIP and ChIP qPCR assay were statistically analyzed by SPSS 16.0 (Statistical Products and Service Solutions, IBM Corporation, Armonk, NY, USA) to check the significant mean difference between control and experimental groups. The
Results
Effect on DNA Methylation
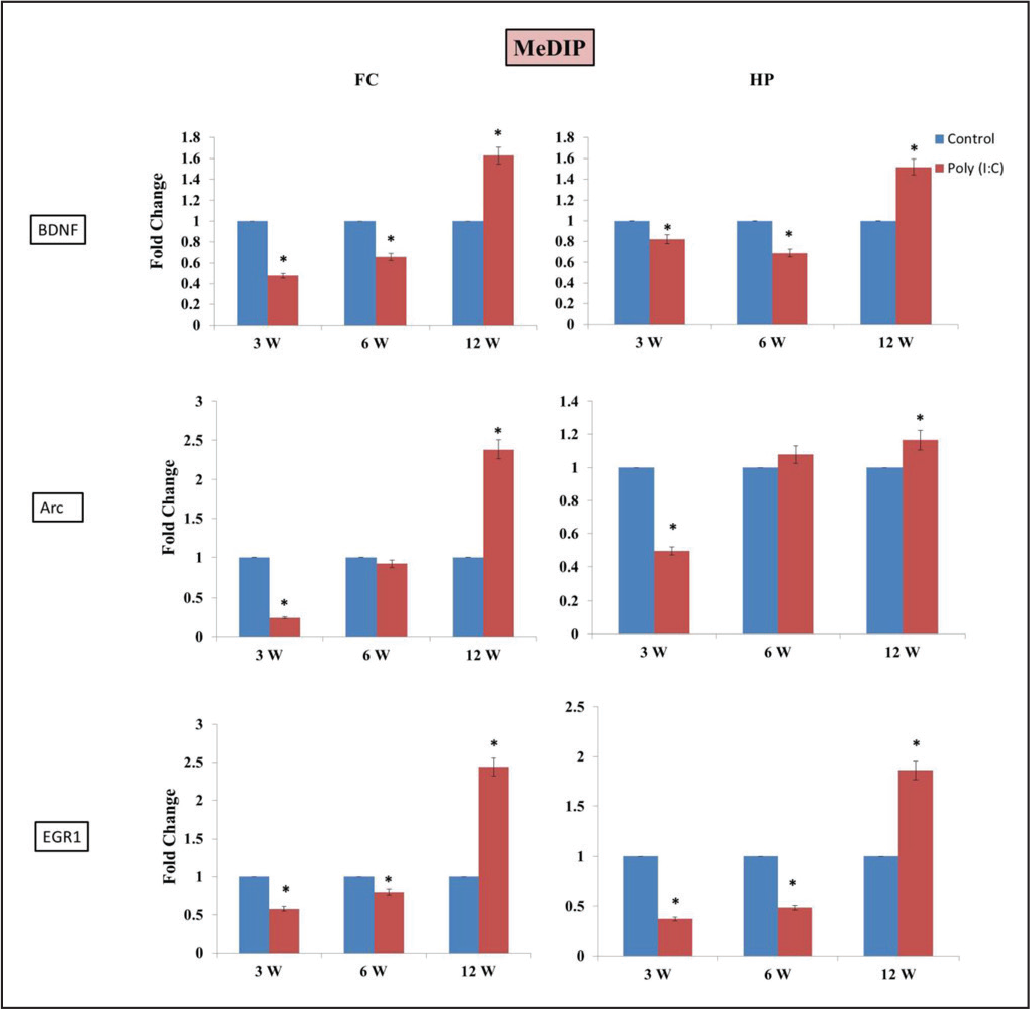
Alteration in 5mC level at gene promoter region tightly regulates its expression. To check this, we performed MeDIP-PCR of promoter sequence of
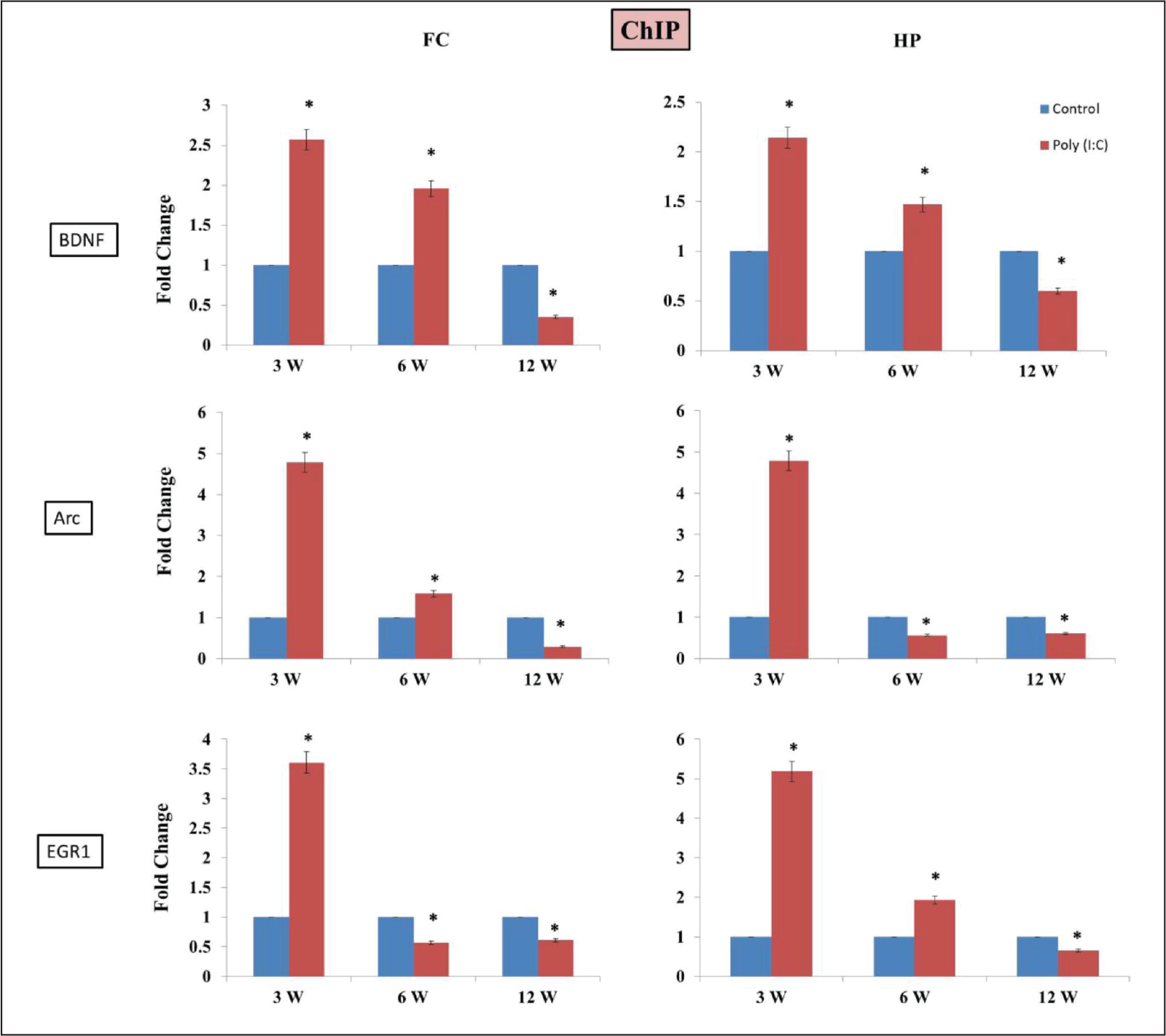
Effect on Histone Acetylation
We have also analyzed histone H3K9Ac status at the promoter of these genes. The exposure to poly (I:C) significantly increased the H3K9Ac level at
Discussion
In the current investigation, we have analyzed epigenetic regulation of differentially expressed synaptic plasticity genes during postnatal exposure of poly (I:C) to developing rats. Poly (I:C) has been reported to cause cognitive deficits and alterations in epigenetic modifications leading to neurological disorders.6, 8, 9 Therefore, it was likely that poly (I:C) affects the epigenetic regulation of synaptic plasticity genes and consequently impairs memory. In epigenetic machinery, DNA methylation is involved in the development of neural system, cognitive functions, neurodevelopmental disorders and neurodegenerative diseases.23–25 Alteration in 5mC level is one of the causal factors for aging and related neurodegenerative diseases like AD. 26
We have found differential methylation pattern at the promoter of
Methylation pattern also corroborates with the expression pattern of these genes.
3
Similarly, alterations in DNA methylation and histone acetylation at
Methylated DNA recruits HDAC2 repressor complex, which removes acetyl group and regulates H3K9Ac level.
35
Histone acetylation is crucial for memory acquisition and consolidation. Therefore, we have analyzed H3K9 acetylation level and found that in the promoter of
Conclusion
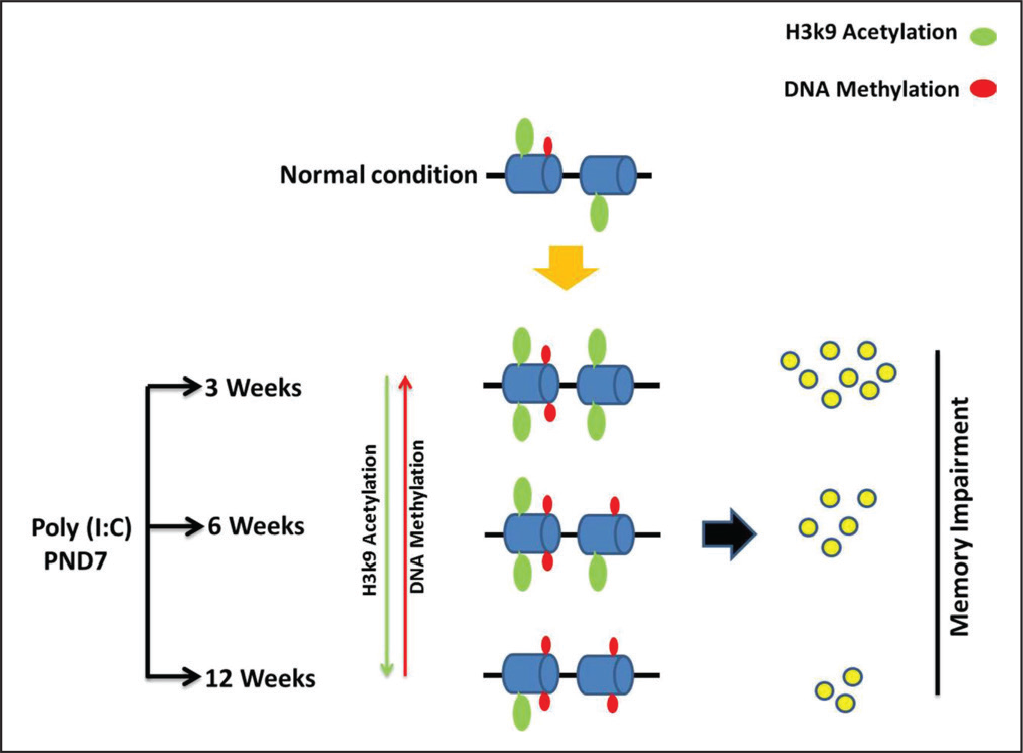
Our findings show that the influence of poly (I:C) exposure during early life also persists in later stages. The poly (I:C) exposure alters methylation of DNA and acetylation of histones at synaptic plasticity gene promoters and subsequently affects gene expression in FC and HP regions of rats even at later ages leading to impaired memory (Figure 4).
Footnotes
Acknowledgements
This work was conducted by three research centers under a collaborative project supported by the Department of Biotechnology (DBT), Ministry of Science and Technology, Government of India (BT/PR14311/MED/30/470/2010). We acknowledge UGC-UPE FA-II, and Interdisciplinary School of Life Sciences for qPCR facility; UGC-CAS in Zoology, Banaras Hindu University, Varanasi for SRF to MSB, and DBT-BET JRF program for DBT-SRF to BS.
Author Contributions
MS Baghel, N Patro, VK Khanna, IK Patro, and MK Thakur: planned and designed the experiments.
B Singh: Conducted poly (I:C) treatment and collected tissue samples.
MS Baghel: Carried out experiments, analysed data and drafted the manuscript.
VK Khanna, IK Patro, and MK Thakur: Edited the manuscript and corresponded with the journal.
Ethical Statement
The maintenance and experimental protocols were approved by the Institutional Animal Ethical Committee, Jiwaji University, Gwalior (Ref. no. IAEC/JU/2010/02) and carried out according to the instructions of Committee for the Purpose of Control and Supervision of Experiments on Animals (CPCSEA).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for
the research, authorship and/or publication of this article: Department of Biotechnology (DBT), Ministry of Science and Technology, Government of India (BT/PR14311/MED/30/470/2010).
