Abstract
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) is a positive sense, single-stranded RNA virus, responsible for the ‘coronavirus disease-2019’ (COVID-19) pandemic. SARS-CoV-2 belongs to the sarbecovirus (lineage-B) sub-genus within the betacoronavirus genus of the coronaviridae family. SARS-CoV-2 shares similarities with SARS-CoV, which was responsible for the 2003 SARS epidemic. SARS-CoV-2 binds with great affinity to the Angiotensin Converting Enzyme-2 receptors on human cells, and its ~30 kilobases long RNA genome hijacks the host machinery and compromises the host immune system with the help of accessory proteins, such as non-structural proteins, resulting in a widespread infection. The spike protein is responsible for the contagious nature of SARS-CoV-2 and, together with the nucleocapsid protein, elicits the host inflammatory response. Several real-time reverse transcriptase polymerase chain reaction (rRT-PCR) tests have been developed to confirm SARS-CoV-2 infection in suspected cases of COVID-19. Furthermore, rapid tests based on SARS-CoV-2-specific antigens and antibodies have been developed to conduct epidemiological surveillance of the hotspot regions that are worst affected by the COVID-19 pandemic. Until effective measures to prevent the occurrence or spread of COVID-19 pandemic are developed, containment measures are being taken, such as isolation of confirmed COVID-19 patients, quarantine of individuals who may have come in contact with a SARS-CoV-2 infected individual, community-wide social distancing, state/nation-wide lockdown, etc. Several vaccines and drugs are being tested that could target the viral structural proteins, non-structural proteins or associated SARS-CoV-2 sub-genomic RNA regions.
Introduction
Coronaviruses are known to infect humans since 1965, when it was extracted from the respiratory tract of an adult with common cold (Tyrrell et al., 1966). Certain types of coronaviruses may cause respiratory and gastrointestinal illnesses in humans that may range from mild symptoms (e.g., common cold) to more severe and potentially life-threatening forms, such as ‘severe acute respiratory syndrome’ (SARS) and ‘Middle East respiratory syndrome’ (MERS) (Y. R. Guo et al., 2020). A novel coronavirus, officially called ‘Severe acute respiratory syndrome coronavirus-2’ (SARS-CoV-2) was detected in Wuhan, Hubei Province, China, in December 2019, and the outbreak has quickly become a pandemic, affecting more than 200 countries and regions.
SARS-CoV-2 is a positive sense, single-strand RNA virus, responsible for the ‘coronavirus disease-2019’ (COVID-19) pandemic (C. Wang et al., 2020). SARS-CoV-2 shares similarities with SARS-CoV, the coronavirus responsible for the previous SARS epidemic that originated in China in 2003. Similar to SARS-CoV, the SARS-CoV-2 binds with Angiotensin Converting Enzyme-2 (ACE2) enzyme receptors on the surface of human cells (A. Wu et al., 2020). Upon gaining entry, the SARS-CoV-2 genome utilises the host machinery to replicate and infect other host cells (Chen et al., 2020). Symptoms of COVID-19 range from asymptomatic or sub-clinical cases to fever and cough, and severe cases like pneumonia-like symptoms that may rapidly progress, deteriorate patient’s condition and may lead to death within weeks of contracting the SARS-CoV-2 infection (Guan et al., 2020). Investigations are ongoing to understand the genomic and molecular structure and function of SARS-CoV-2, to identify vaccine and drug targets. Several real-time reverse transcriptase polymerase chain reaction (rRT-PCR) tests have been developed to confirm SARS-CoV-2 infection in suspected cases of COVID-19 (https://icmr.nic.in/content/covid-19). Furthermore, rapid tests based on SARS-CoV-2-specific antigens and antibodies have been developed to conduct epidemiological surveillance of the hotspot regions that are worst affected by the COVID-19 pandemic. This article provides an overview of SARS-CoV-2 taxonomic classification, genomic and molecular structure, function, and currently utilised diagnostic and surveillance measures.
Taxonomic Classification
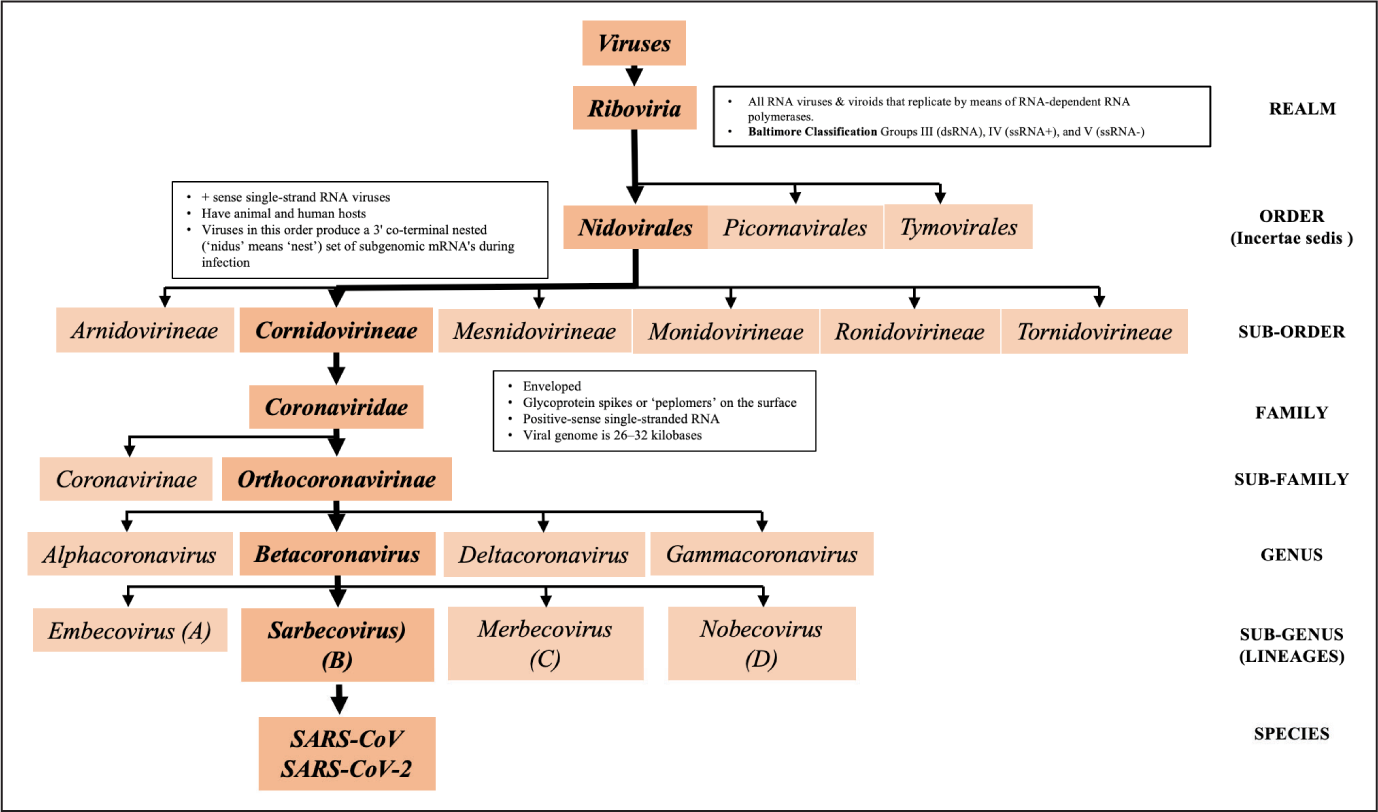
Coronaviruses belong to the Coronaviridae family of viruses in the order Nidovirales. Coronaviridae family of viruses may cause respiratory, gastrointestinal and central nervous system–associated illnesses in humans and animals (Perlman & Netland, 2009).
Based on the genetic architecture, within the Coronaviridae family, the sub-family Orthocoronavirinae is divided into four genera, namely Alphacoronavirus, Betacoronavirus, Deltacoronavirus and Gammacoronavirus (Li, 2016). Coronaviruses that are known to infect humans have been identified in alphacoronavirus (Human CoV-229E and Human CoV-NL63) and betacoronavirus (Human CoV-HKU1, Human CoV-OC43, SARS-CoV, and MERS-CoV) genera. The latest pandemic, Coronavirus Disease-19 (COVID-19) is caused by a betacoronavirus, SARS-CoV-2 (Figure 1).

Genomic Architecture
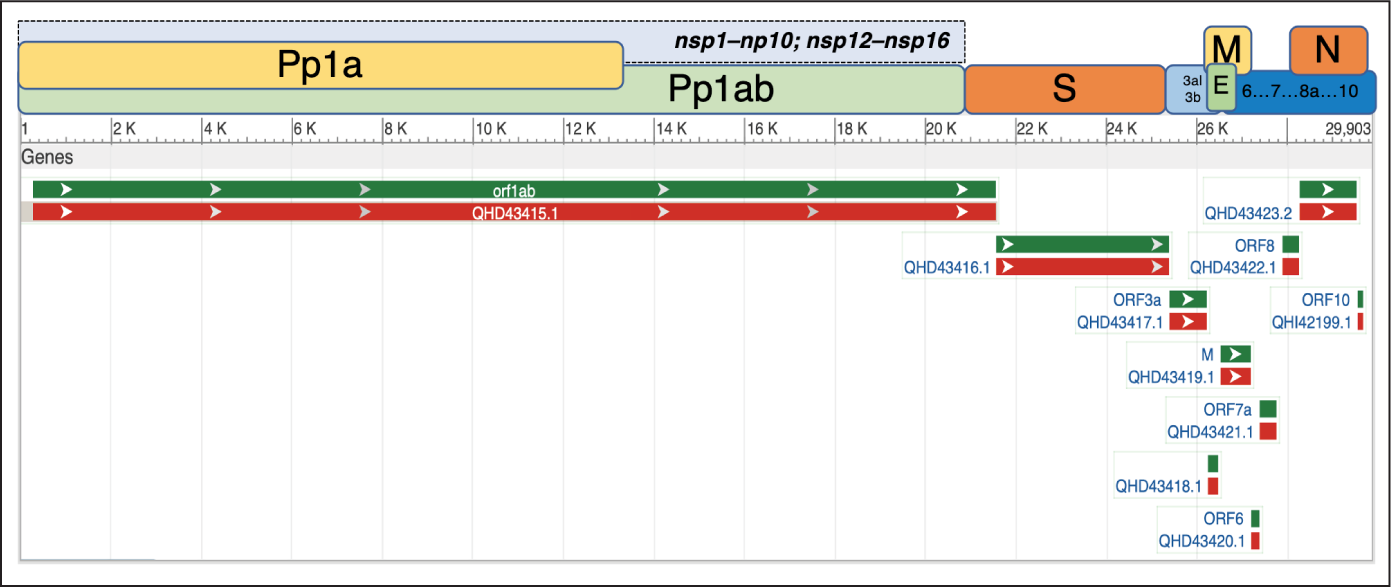
Coronaviruses are encapsulated, positive-sense single-stranded RNA viruses with the largest genome among all RNA viruses (Brian & Baric, 2005; Gorbalenya et al., 2006), with the size ranging from 26–32 kilobases (Lei et al., 2018). SARS-CoV-2 genome size is ~30 kb. In the host cell, the positive-sense RNA genome of SARS-CoV-2 can be directly translated to proteins. SARS-CoV-2 genome contains 14 open reading frames (ORFs) that encode for 27 structural and non-structural proteins (nsps). Starting from the 5′-terminus, the first ORF (orf1ab) is 21,290 bases long, making up 71 per cent of the entire SARS-CoV-2 genome (including untranslated regions [UTRs]) and encodes for pp1ab or orf1ab polyprotein. The spike glycoprotein (S) is encoded by gene S (3,822 bases) and helps SARS-CoV-2 to bind with the ACE2 receptor expressed mostly on certain types of human cells, including type II pneumocytes, endothelial, myocardial and gut mucosal cells (Zhang et al., 2020). The other three structural proteins are the envelope protein (E), membrane glycoprotein (M) and nucleocapsid (N) protein (Table 1 and Figure 2). The receptor-binding domain (RBD) of the spike protein is thought to have a particularly strong affinity for ACE2 receptors, conferring upon the virus its highly contagious nature. Both the S and N proteins are highly immunogenic structural proteins (Qiu et al., 2005), and the resulting antibodies play a crucial role in blocking viral entry into host cells and opsonisation, leading to effective phagocytosis (Walls et al., 2020). Gaining an understanding of these proteins is therefore the key to developing effective vaccines and therapeutic modules.
Key Genes and Encoded Proteins Based on the SARS-CoV-2 RNA, Based on Severe Acute Respiratory Syndrome Coronavirus 2 Isolate Wuhan-Hu-1 (GenBank ID: MN908947.3)

The orf1ab gene of the viral genome encodes for several NSPs that play a role in replication of SARS-CoV-2 inside the host cell. The role includes formation of the replication-transcription complex (RTC), which aids in the replication of the viral genome into genomic RNA as well as sub-genomic RNAs (sgRNA) through discontinuous transcription that encode for the structural proteins. An investigation was done using X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy of the multi-domain nsp3 protein; the largest NSP in coronaviruses is an essential component of the RTC (Lei et al., 2018). Nsp1 protein binds to the 40S subunit of the human ribosomes and potently inhibits or shuts off the translation function in the host cell and causes degradation of host mRNAs in the infected cell (Huang et al., 2011; Tanaka et al., 2012). Nsp12 is a key protein in the virus-encoded RNA-dependent RNA polymerase (RdRp) or RNA replicase (Ahn et al., 2012). A detailed review of nsps has been published previously (Chen et al., 2020).
Host Cell Entry and Replication
The spike protein of SARS-CoV-2 recognises the ACE2 receptor on the human cell surface, binds with it and gains entry into the host cell through membrane fusion or endocytosis (Chen et al., 2020). Once inside the host cell, SARS-CoV-2 RNA genome is released in the host cytoplasm. The viral RNA genome along with the help of nsps hijacks the host ribosomes and nutrients to form RdRp or RNA replicase, as well as other accessory proteins that are needed for subsequent steps of viral replication (Chen et al., 2020). Nsps combine with RdRp to make the RTC. The RTC creates several copies of genomic and sub-genomic (sgRNA) viral RNA sequences (Sola et al., 2015). The sgRNA sequences are formed through discontinuous transcription (Sola et al., 2015; Wu et al., 2010) and encode for SARS-CoV-2 structural proteins. The structural proteins combine and encapsulate the genomic RNA and form the mature virion. The mature copies of SARS-CoV-2 are transported to the infected cell’s surface and released through exocytosis (Chen et al., 2020).
Diagnosis
Clinical Symptoms
The most common symptoms reported in COVID-19 patients include fever, cough, shortness of breath or difficulty breathing; repeated shaking with chills; muscle pain; headache; sore throat; and new loss of taste or smell (Guan et al., 2020). The range of symptoms may vary, and a COVID-19 patient may experience some, and not all, of these symptoms. Also, sub-clinical or asymptomatic patients have also been reported (Y. Bai et al., 2020). For each individual suspected of having COVID-19, detailed travel and clinical history should be collected. Nasopharyngeal swab should be collected for genetic testing of the infection (described later in the article). The individual should be isolated until he or she tests negative on two consecutive tests and is free of clinical symptoms of the disease. In addition, all individuals who have come in contact with the suspected case should be quarantined for at least 2 weeks, in order to prevent further spread of the infection.
rRT-PCR
A variety of tests have been proposed for the detection of COVID-19, and rRT-PCR is currently considered the ‘gold standard’ method (Corman et al., 2020). This in vitro confirmatory test is used to detect SARS-CoV-2 in the samples collected from a patient suspected to have COVID-19. Nasopharyngeal swabs are the sample of choice (Marty et al., 2020). Other acceptable samples include oropharyngeal swabs, nasal aspirate, anterior nares and mid-turbinate (Zou et al., 2020). Samples taken from sputum, endotracheal aspirates, bronchoalveolar lavage or early-morning posterior oropharyngeal saliva samples may be sent directly to the microbiology laboratory for processing and may be more sensitive than upper respiratory tract specimens, although more hazardous to collect (To et al., 2020; W. Wang et al., 2020).
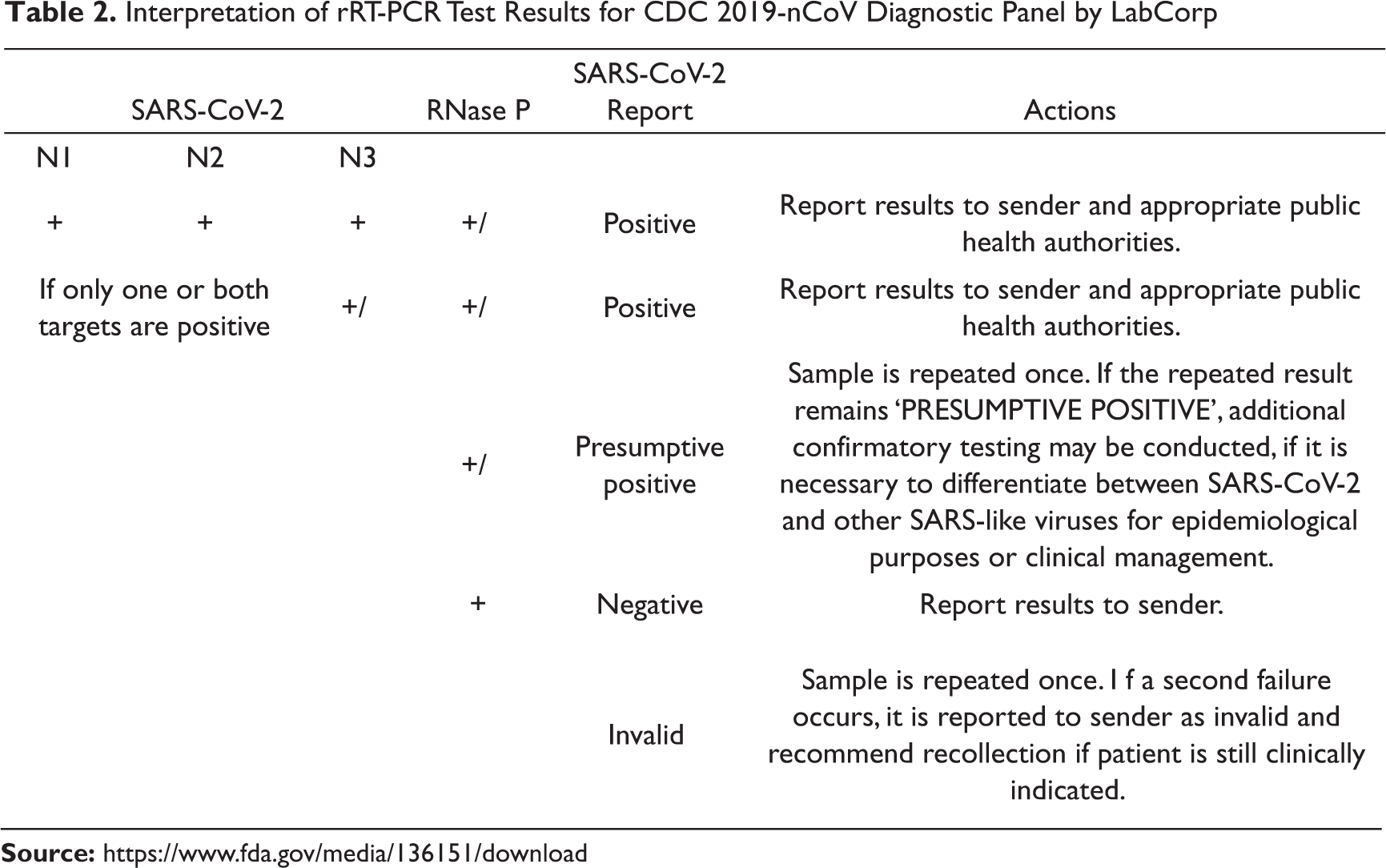
In order to perform rRT-PCR assay-based detection of SARS-CoV-2, a nasopharyngeal swab specimen is collected using standard safety protocol and methods (Marty et al., 2020). Regions of a viral genome are selectively amplified to detect the presence of SARS-CoV-2 genome. The rRT-PCR tests designed for SARS-CoV-2 usually detect RNA regions lying in Orf1ab gene, S gene, E gene and N gene (Zou et al., 2020). The viral RNA is extracted from the infected cells, isolated and purified. The viral nucleic acid needs to be reverse transcribed to a synthetic DNA called complementary DNA or cDNA. The cDNA is then amplified using one of the rRT-PCR assay instruments. The rRT-PCR reports the presence of the target sequence through a fluorescent signal. With each amplification cycle of the cDNA, the fluorescence signal becomes stronger. As soon as the fluorescence level crosses the predetermined fluorescence threshold, presence of SARS-CoV-2 is confirmed. The viral load in the sample determines the cycle threshold (Ct) of a test. Ct is the number of rRT-PCR amplification cycles needed to reach the fluorescence threshold. A lower Ct number indicates higher viral load in the tested sample. The fluorescent signal from each test site of the RNA genome, for example, N1, N2 and N3 sites, are done according to the test protocol put forth by the Centers for Disease Control (CDC). There are predetermined criteria to interpret the rRT-PCR test results as described by the manufacturers like LabCorp (Table 2) and Thermo Fisher Scientific (Table 3).
Interpretation of rRT-PCR Test Results for CDC 2019-nCoV Diagnostic Panel by LabCorp
rRT-PCR result interpretation for TaqPath™ COVID-19 Combo Kit

A recent meta-analysis identified the sensitivity of rRT-PCR to be 89 per cent (95% confidence interval [81%,94%]) (Kim et al., 2020). However, the sensitivity is highly dependent on the quality of sample taken and may be as low as 50 per cent and as high as 100 per cent (Kim et al., 2020). Additionally, the typical turnaround time of 24–48 h warrants the need for more rapid assays that can be used at the point of care. It is important to note that a positive test result does not eliminate the probability of a bacterial co-infection or another co-morbidity. Similarly, a negative result does not categorically rule out the presence of COVID-19 infection, as the quality of the collected sample, concentration of the collected RNA samples, number of amplification cycles and mutations in the tested regions of the RNA genome may affect the sensitivity of the test. Clinical findings should be used by the physician to conclude the final diagnosis.
Rapid Diagnostic Tests
Epidemiological sero-surveillance of the regions most affected by COVID-19, or hotspot regions, needs to be performed. Individuals who are exposed to SARS-CoV-2 develop antigen-specific antibodies in a few days. These antibodies are detectable in patient’s blood/serum/plasma samples within 7–10 days of contracting infection. As of 16 April 2020, the National Institute of Virology, Pune, India, had validated 23 rapid tests, including nine kits that are manufactured in India (Table 4).
Antibody-based Rapid Tests Validated by the National Institute of Virology, Pune, India (as of 16 April 2020)

Serological tests that identify antibodies (such as IgA, IgM and IgG) to SARS-CoV-2 from clinical specimens (such as blood or saliva), such as enzyme-linked immunosorbent assays also have low complexity and can be used at point of care. However, since the body can take days to weeks to mount an antibody response, there is a high probability of false-negative results during the early/asymptomatic stage of disease (L. Guo et al., 2020). Moreover, the test cannot differentiate between past and present infection. There may be cross-reactivity of the antibody to non-SARS-CoV-2 coronavirus proteins in case of coinfection, giving rise to false-positive results (Patrick et al., 2006). At present, serologic assays may be useful in confirming a diagnosis of COVID-19 in patients with complications who test negative for RT-PCR, due to reduced viral load over time (To et al., 2020). Improved test accuracy can make immunoassays an ideal tool for epidemiological surveillance.
Antigen detection tests for COVID-19 are in development, based on recently generated monoclonal antibodies against the nucleocapsid protein of SARS-CoV-2 (Sheridan, 2020). These low-complexity tests that detect antigens by immunoassay directly from clinical specimens already exist for respiratory syncytial virus and influenza virus, and have a fast turnaround time (minutes), although their sensitivity is suboptimal (Hogan et al., 2018; Prendergast & Papenburg, 2013). If sensitivity can be improved for COVID-19, antigen detection tests can be of great value in low-resource situations such as home testing. The purpose of these tests would not be for treatment planning, but to identify persons that need quarantine, including asymptomatic cases and their contacts (Cheng et al., 2020).
Given the limited sensitivity of commonly used detection methods (RT-PCR and serological tests), ancillary tests can be a vital supplement to aid in the clinical diagnosis of suspected cases. Computed tomography of the chest (chest CT scan) has been shown to have even better sensitivity than RT-PCR. Ground-glass opacities, the most commonly detected feature, appear in 78–100 per cent of COVID-19 patients. Peripheral distribution, fine reticular opacities and vascular thickening have also been observed (H. X. Bai et al., 2020). However, these findings have limited specificity for COVID-19, and it may indicate other lung conditions (X. Wu et al., 2020). Since COVID-19 elicits an intense host inflammatory response, some commonly observed laboratory biomarkers include decreased albumin, elevated C-reactive protein, elevated lactate dehydrogenase levels and lymphopenia (Rodriguez-Morales et al., 2020). These may be early markers of organ failure, but their utility as a prognostic tool is as yet unconfirmed.
Cutting-edge diagnostic techniques, such as clustered regularly interspersed short palindromic repeats (CRISPR)-based methods, are now being used to develop rapid, inexpensive, portable, temperature-stable assays for use outside of clinical settings (such as airports and border crossings). A recently developed CRISPR–Cas12 test was shown to have 95 per cent positive predictive power and 100 per cent negative predictive power. Also, it overcame the limitation of cross-reactivity with other coronavirus proteins (Broughton et al., 2020). In India, the Institute of Genomic and Integrative Biology (IGIB) in New Delhi has developed a paper-strip CRISPR-Cas9-based test and can be used at ‘Mohalla clinics’. It is in the validation and approval stage.
In conclusion, there is no effective prevention method currently available to contain the spread of the SARS-CoV-2 infection other than extreme measures of widespread lockdowns. In the first 4 months of this pandemic, more than 2.5 million confirmed COVID-19 cases, including more than 170,000 deaths, have been reported. Rapid and reliable testing methods, early isolation, and effective vaccines and therapeutic approaches are urgently needed to limit the damage caused by this global catastrophe.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
