Abstract
Short tandem repeat (STR) DNA, one of which is the FGA locus, can be used as a profiling tool for personal identification analysis in the forensic field because it has many allelic combinations based on the complex repetitive pattern of DNA sequences. The FGA locus was reported to have the highest discrimination power in the Indonesian population. The aim of this study was to understand the ability of personal discrimination from the FGA locus sequences in a DNA forensics testing scenario of Indonesian subjects. The specimens were used towels (T), toothbrushes (TB), and hair ties (HT) from certain people and buccal swabs (BS) from the same people and two others as controls whose identities were analyzed. DNA extraction using the Phenol CIAA (chloroform isoamyl-alcohol) method followed amplification of the DNA of FGA loci that targeted repetitive sequences, agarose gel electrophoresis, and Sanger sequencing of FGA loci. The amplification showed the DNA bands from the FGA loci were around 350 bp. The sequencing results showed that the number of repetitions of the FGA allele from samples T, TB, and HT was identical to BS1, which was 19 repetitions, while BS2 and BS3 were 22 and 21 repetitions, respectively. Thus, the sample items T, TB, and HT are owned by individual BS1. The FGA gene sequenced by the Sanger method can be used in a limited way, only detecting repetitive sequences on the longest allele as a differentiator between one individual and another.
Introduction
In the human genome, there are many sequences with varying repetitive patterns, one of which is short tandem repeat (STR) DNA. Recently, the standard method of forensic DNA profiling has been STR analysis. 1 STR DNA can be used as a profiling tool for personal identification analysis in the forensic field because it has many allelic combinations based on the length of the complex repetitive DNA sequences being analyzed. 2 STR DNA identification from biological specimens using Polymerase Chain Reaction (PCR) has been proven to be a simple method, but very accurate and sensitive in analyzing or differentiating each individual. 3
DNA profiling is used to determine an individual’s unique genetic characteristics. It involves analyzing specific regions of the DNA that exhibit variations among individuals. These regions, known as genetic markers (biomarkers), include STR loci like FGA. 4 The combined DNA index system (CODIS) is a DNA database system used in the United States for the storage and comparison of DNA profiles obtained from crime scene evidence, convicted offenders, and missing persons. It was developed by the Federal Bureau of Investigation (FBI) and is widely utilized by federal, state, and local forensic laboratories. From the beginning of its appearance in October 1998 until December 31, 2016, there were 13 STR locus cores, then updated effective January 1, 2017, to 20 STR locus cores. The most recent version of CODIS as of now is CODIS 20, which refers to the number of 20 core STR loci included in the system.5, 6
The STR locus on the combined DNA index system 13 (CODIS13), which has the highest discrimination power in the Indonesian population, was the FGA locus. 7 The FGA locus contains mixed tetranucleotide repeats found in the third intron of the Human Alpha Fibrinogen gene on the long arm of chromosome 4 (4q28). FGA, also known as FIBRA or HUMFIBRA, has a high mean exclusion chance compared to other common STR loci, making it an effective paternity test and forensic identification tool. 4
The FGA locus has a complex tetranucleotide repeat motif called core repeat and total repeat; the consensus sequences are [TTTC]3[TTTT][TTCT][CTTT]n[CTCC][TTCC]2. 8 Total repeats range from 13 to 29 times, with a core repeat pattern CTTT of 5–21 times. 2 FGA loci have a size range of 308-465bp. 9 According to the size total repeat pattern, FGA alleles can be divided into two groups, those in the allele size ranges of 16–34.2 and 42.2–51.2. Although we have not yet found any alleles between 34.2 and 42.2, it is possible that some people will carry these alleles. 10 Repetition allele ranged between 16–31.2 and 40.2–44.2 reported in the South African population. 11
STR analysis involves amplifying and analyzing the number of repetition sequence units at certain loci. By comparing the number of repeats at a certain locus between a known sample (e.g., crime scene evidence) and a reference sample (e.g., DNA from a suspect or a DNA database), it is possible to determine if the samples originate from the same individual. This research, therefore, aims to find out how effective FGA locus sequences are for personal identification in DNA forensics testing of Indonesian subjects.
Material and Methods
Study Design
The design of this study was an experimental design. Personal items used as a source of DNA were used towel (T), a toothbrush (TB), and a hair tie (HT) from one female student of the Universitas Muhammadiyah Semarang, while buccal swabs (BS) were taken from BS from the same female student and two other students as a comparison control. This research was conducted at the Molecular Biology Laboratory at Universitas Muhammadiyah Semarang.
Sample Collection
Sample collection was done on a used towel without being washed by cutting the fibers suspected of having attached skin epithelium. TB and HT were cut at the suspected epithelial spots. 12 The buccal area was swabbed by rotating and rubbing the buccal area 20 times.
DNA Extraction
DNA extraction using the Phenol CIAA (Chloroform Isoamyl-Alcohol) method. Each specimen was put into a 1,5 ml tube and added 1:1 with lysis buffer (100 mM Tris HCl, pH 8; 100 mM NaCl; 50 mM EDTA; 2% SDS). 20 µl of Proteinase-K (10 mg/ml) was added and incubated in a dry bath at 55°C for 5 hours. Cotton swabs and towel fibers in the microtube were taken using sterile tweezers. Add Phenol CIAA (25:24:1) 1:1 and vortex for 10 seconds. The suspension was centrifuged at 8.000 rpm for 5 minutes. The aqueous phase supernatant was taken and transferred to a new 1,5 microtube. Cold ethanol (96%) was added 1:2 to the microtube and centrifuged at 13.000 rpm for 5 minutes at 4°C. Remove the supernatant. 70% ethanol was added as much as 500 µl into the microtube containing the DNA and then centrifuged at 13.000 rpm for 5 minutes at 4°C. The supernatant was discarded, and this process was repeated three times. After that, the supernatant was discarded, and the pellet was air-dried. The final step was dissolving the DNA isolate with 25 µl of Tris-EDTA (TE buffer) and storing it at −20°C. The concentration and purity of DNA were measured using the MN-913A MaestroNano Pro (MAESTROGEN) spectrophotometer.
FGA Locus Amplification
DNA was amplified by PCR using the T-Personal Thermocycler (BIOMETRA) using a pair of primers at the FGA locus (forward 5’-GGCTGCAGGGCATAACATTA-3’ and reverse 5’-ATTCTATGACTTTGCGCTTCAGGA-3’). 13 Amplification of all samples used FGA loci by PCR T-Personal Thermocycler (BIOMETRA). The setting was as follows: 95ºC for 3 min, then 95ºC for 30 sec, 58ºC for 30 sec, 72ºC for 30 sec, for 35 cycles. DNA polymerized template continued with typing while stored at 4ºC.
Electrophoretic Reaction
The result shows that PCR products were electrophoresed on MUPID-exU submarine electrophoresis under 3% agarose with DNA ladder DM1100 ExcelBand™ 50bp (SMOBIO) to analyze the size of DNA bands.
DNA Sequencing and Analysis
PCR products from samples that show well-amplified band results will be carried out using the DNA sequencing method of Sanger Sequencing by 1st Base through PT. Genetics Science Indonesia. DNA sequence results were analyzed using assembly contig sequence with DNA Baser Assembler version 5.20 software and nucleotide similarity alignment using BLAST from
Results
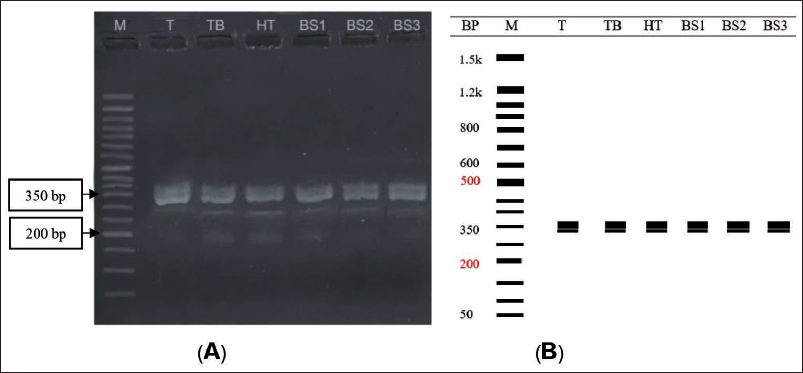
The PCR product, as a result of FGA amplification, was visualized on a 3% agarose gel, with visible DNA bands measuring around 350 bp (Figure 1). The result shows clearly visible DNA bands, which have a size of around 350 bp. The gel also shows double bands from the sample.
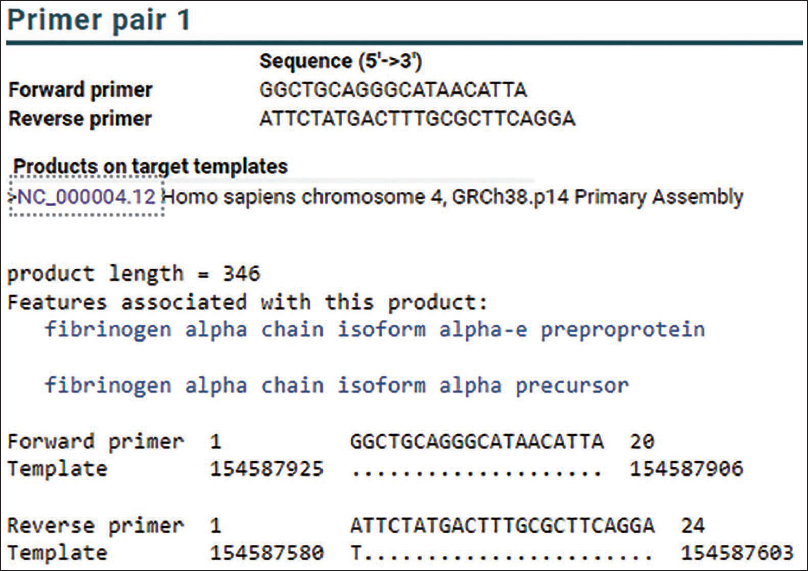
Simulation results from running Primer-BLAST NCBI using the FGA primer obtained the target product with a size of 346 bp (Figure 2) and used it as a reference. In the reference sequence, we can see that there are 14 repetitions of the core sequence [CTTT]14 (Figure 3). The target DNA sequence is on the negative strand, based on the Genbank database.
Modified Figure of Running Primer-BLAST Using FGA Primers.
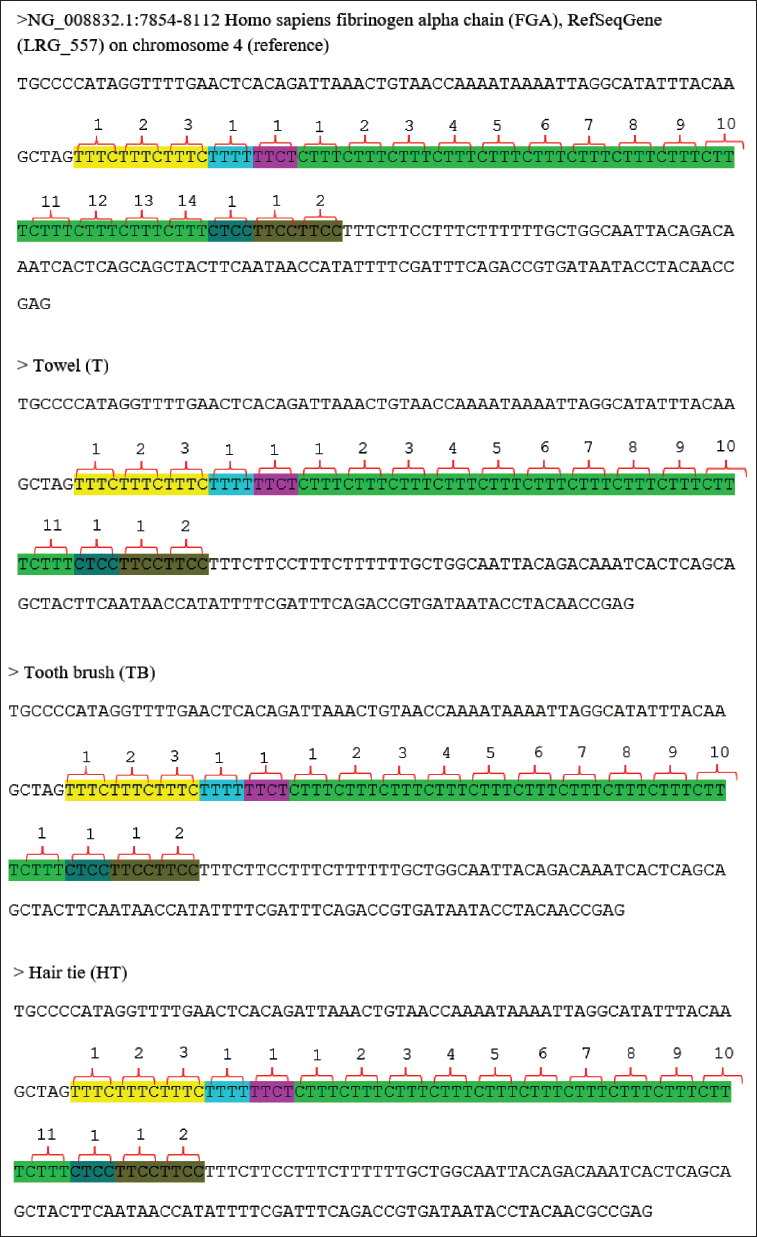
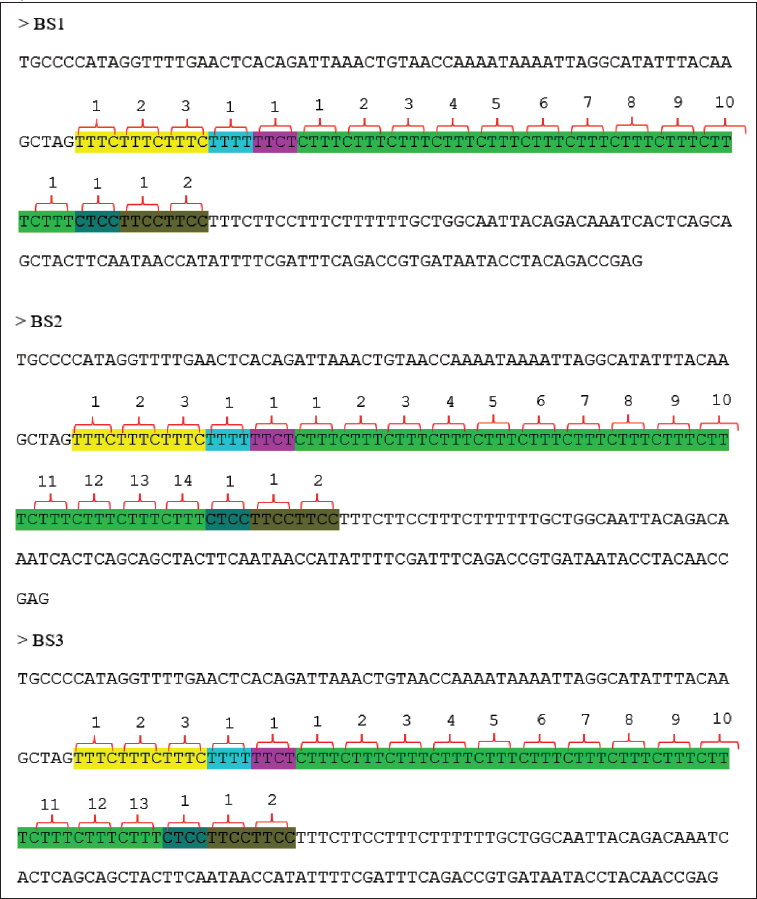
FGA Sequences Result in the Form of Complex Repetition Motives from the Database Reference with Accession Number NG_008832.1 and the Samples: Towel, Toothbrush, Hair Tie, Buccal Swab 1 (BS1), Buccal Swab 2 (BS2), and Buccal Swab 3 (BS3).
During the assembly process, the DNA sequences from Sanger sequencing were trimmed at the beginning and at the ends of the DNA segments so that the contig DNA sequences were shortened. The contig was then subjected to alignment on BLAST and showed the highest similarity to Homo sapiens fibrinogen alpha chain (FGA), RefSeqGene (LRG_557) on chromosome 4 with accession number NG_008832.1 and will be used as a reference gene.
Multiple sequence alignment analysis from the DNA sequencing results was performed using Clustal Omega software (
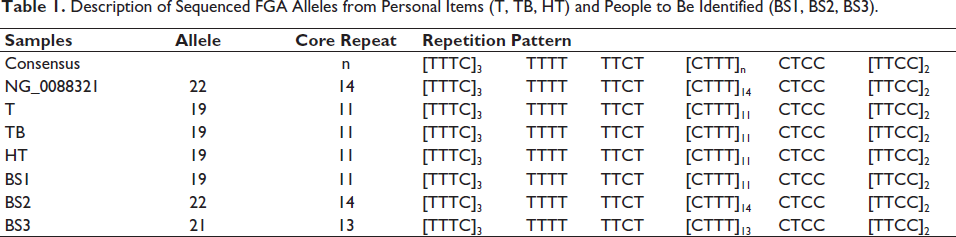
The STR allele polymorphism of the FGA locus in the samples was the number of repetitions of the [CTTT]n core pattern, which was different for each human in this research scenario. This can be used as a difference between one individual and another. The total repetition pattern [CTTT]n in the samples showed variation results (Table 1).
Description of Sequenced FGA Alleles from Personal Items (T, TB, HT) and People to Be Identified (BS1, BS2, BS3).
Discussion
STR loci have various types based on repetitive patterns, including simple, compound, and complex types. Simple types have repetition units that are identical in repetitive sequence, along with the length. Compound repetition is made up of two or more contiguous simple repeats, while complex repetition has several repeat units with varied unit lengths as well as variable intervening sequences and a large number of variant alleles. FGA is a complex type of STR. 2 The basis for determining alleles from a STR locus is based on the length of DNA fragments, especially those that have certain repeating patterns in them. The standard method for analyzing alleles in STR DNA is capillary electrophoresis. 14 In this study, we try to perform the Sanger sequencing method to analyze STR DNA in the FGA locus. The results of this study were a DNA band from a 3% agarose gel and a Sanger sequencing result.
Electrophoresis band results may show the size of the target DNA fragment (Figure 1). However, the agarose gel electrophoresis method cannot show accurate measurement results; therefore, we followed it with the Sanger sequencing method, which can analyze the sequences of the target DNA. The result from a 3% agarose gel was around 350 bp and showed double bands. It means that the DNA fragment has a different length for each allele in diploid chromosomes; the more repetitive units, the longer the DNA band detected in an allele, and this finding is in line with. 15
The sequencing results at the FGA target locus clearly show the repetition pattern sequence (Figure 3) and the number of repetition units (Table 1). However, the repetition unit that was read in the sequence was only the longest allele, so this is a weakness of STR analysis by Sanger sequencing analysis; this finding is supported by. 16 Sanger STR sequencing cannot be performed routinely because loci cannot be multiplexed and heterozygous alleles must be separated manually before sequencing. 17 This method can be used as an alternative to STR DNA analysis for personal testing, although it has limitations. This limitation can be reduced by using more than one STR locus to strengthen the analytical results.
Based on the scenario of our research, using non-living material as a sample to determine the identity of the owner of the sample, DNA extraction has been successfully carried out. We matched the longest repetition alleles so we could get a match with the owner of the DNA in the sample. The sample used towels, tooth brushes, and HT was identical to BS1, which was 19 total repetitions with 11 core repetitions [CTTT] of allele variations. BS2 has a total of 22 repetitions with 14 core repetitions [CTTT] of allele variations. BS3 has a total of 21 repetitions, with 13 core repetitions [CTTT] of allele variations (Table 1). The results of the used towels, toothbrush, and HT sample match BS1, an indication that the three samples have the same number of repetitions as BS1. It can be concluded that the sample belongs to BS1.
Conclusions
A personal identification test using the FGA locus can be done, but with limitations, only detecting repetitive sequences on the longest allele. The use of more than one STR locus may increase the accuracy of the discrimination analysis. From the research scenario, we can find out who owns the DNA in the sample being examined.
Footnotes
Abbreviations
CODIS: Combined DNA index system
DNA: Deoxyribonucleic acid
Phenol CIAA: Phenol chloroform isoamyl-alcohol
PCR: Polymerase chain reaction
STR: Short tandem repeat
Acknowledgements
We thank the laboratory assistants in the molecular biology laboratory and the Department of Nursing and Health Sciences at Universitas Muhammadiyah Semarang, which has provided the materials and equipment for carrying out the research.
Authors’ Contributions
ARE was responsible for the conception and design of the study (lead) and formal analysis (lead). DAK, YA, and RS contributed to the sample collection. DAK performed an investigation in the laboratory experiment. YA and RS helped in the laboratory experiment. RS is responsible for operating the software and the visualization of study results. YA contributed to the analysis of data and interpretation. AIK was responsible for conception (supporting), writing review, and editing. ARE wrote the original draft (lead). DAK wrote the original draft (supporting). All authors read and approved the final manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
Ethical approval to carry out this research was obtained through ethical clearance number 209/VI/2022/KomisiBioetik issued by the Institutional Review Board and Bioethics Committee of the Faculty of Medicine, Universitas Islam Sultan Agung.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Informed Consent
All individuals who participated in this research provided informed consent and permitted the dissemination and publication of the results. Privacy and participant confidentiality were strictly maintained throughout the study.
