Abstract
The current investigation explores the possibility of using skin microbiomes to solve criminal cases. The study investigates the skin microbial samples as a marker from four professions, that is, garbage collectors (GC), morgue staff (MW), construction workers (CW), and housekeeping staff (HK). The findings show the cluster-wise separation of Bacillus spp. in four different professions. The heat map, scatter plot and cluster dendrogram show the discrete pattern of separation of 16S rDNA on the basis of occupation. The results clearly demonstrate that the CW and GC samples may share some common features that are not shared by the MW and HK samples. Most of the CW samples are related to each other in most of the branches and related to most of the GC samples. This suggests that the bacterial communities on the palms of CW and GC samples are more similar to each other than they are to the other groups. The cluster dendrogram shows some of the HK and MW samples together in the branch. This similarity can help in establishing the link between the corroboratory evidence and the suspect and/or crime scene. However, more samples should be studied for involving it in a full-fledged criminal investigation. Studies on transient and resident bacterial communities on surfaces are crucial. This study shows the promising method of skin microbiome for forensic applications. With continued research and development, this technology could become a valuable and cost-effective tool for criminal investigations.
Introduction
The skin acts as a vital barrier between the human body and the external environment, with its diverse microbiota contributing to skin health. The topographical and temporal diversity of the human skin microbiome was investigated using genomic techniques in a study carried out by Grice, et al. 1 To make microbial forensics viable for investigations, rigorous validation and ethical consideration 2 and the establishment of reliable methods and protocols are required. Forensic practitioners require proficiency and standardized training. The human skin is home to a wide variety of bacteria that are easily transferred to surfaces through touch and are resistant to environmental stresses. Skin microbiomes differ from person to person but remain stable over time, with potential forensic applications as unique “fingerprints.” Personal contact with environmental sources can cause bacterial communities on the skin to persist on surfaces for days to weeks. Microbial DNA fingerprinting from touched surfaces could help forensic investigations by providing information on individual hosts, including their geographic origins. This method may be useful in cases where there are no matching DNA profiles of individuals. Molecular characterization of bacteria from crime scene exhibits, understanding bacteria’s role in fluid degradation, and investigating spatial and temporal variations in human bacterial microflora are all part of the exploration of microbial forensics. 3 The molecular characterization of mitochondrial DNA of different species can be used to rule out the origin of the degraded blood samples. 4 Microbial profiling currently lacks the accuracy required for forensic trace evidence, necessitating additional research to identify potential limitations and effective mitigation strategies. 5 Amorim’s 2010 proposal for microbial forensics shows promise in solving sexual offenses by linking perpetrators and victims years after the incident using bio-indicators such as HIV (Human Immunodeficiency Virus) and HCV. 6 As Budowle and Skopp illustrate, clear protocols, technique validation and dependable databases are critical before broad usage in forensic investigations. 6 Few of the researchers delve into unprecedented detail to examine the skin microbiota in health and illness. Comparative investigations of microbiome sequencing data can produce ideas regarding disease-causing microbes, which can guide targeted culture, whole-genome sequencing, and functional research for dysbiosis and pathogen treatment development. 7
With the help of extensive sequence databases, the 16S rDNA gene serves as a molecular marker for bacterial identification. Understanding of human microbiota, emphasizing that, contrary to popular belief, the number of microbial cells in the body is comparable to somatic cells. The microbiome, which represents the DNA content of these microbes, outnumbers the diversity of the human genome. The emphasis is on microbial forensics, specifically skin microbiome analysis and on individual variations and potential forensic applications. Molecular sequencing advances, particularly next-generation sequencing, improve forensic capabilities. The skin microbiome, which is diverse and influenced by a variety of factors, provides a distinct microbial signature that could aid in human identification, complementing traditional forensic methods such as fingerprint and DNA analysis. 8
Forensic applications by demonstrating individual microbiome persistence into the early postmortem period and the traceability of microbial communities on surfaces back to individuals. 9 Using targeted sequencing of microbial markers results indicate that microbial strain composition is more individualizing than phylogenetic relationships, emphasizing the potential of microbial composition for precise identification, possibly reflecting host-environment interactions that maintain a distinct microbial profile. 10 Bacterial abundance in source communities predicted transmission and residence times, demonstrating that transient encounters with environmental sources can drive skin microbiome variability if sufficient biomass is present. 11
Overall, the study provides a thorough overview of the difficult and time-consuming task of defining bacterial species. It emphasizes the significance of employing multiple methods of analysis and the need to take into account factors such as recombination and ecological niche. 12 The skin’s micro-organisms, known as the cutaneous microbiota, are thought to play important roles in the prevention and treatment of skin diseases. This culture-independent genetic profiling sheds light on potentially novel human cutaneous microbiota components. 13 Current and past studies in microbial applications of forensic sciences promise betterment and more accurate investigation techniques like DNA or any biological or chemical trace evidence analysis. The crucial reason behind it is a universal distribution of microbes and their persistence for a longer time and the change occurs when drastic environmental changes occur. 14 They are even unique to a particular person and even to a body fluid and body part. 15 This aspect can be used for identifying burglaries. The need for research and development and method validations, standardization and ethical considerations are required to present microbes as evidence in the courts.16, 17 The study suggests that human-associated microbes show variations in personal discrimination. The detailed study of microbial and non-microbial study together will prove even better to link a suspect to the evidence and crime scene. This field of investigation does not only depend on microbial techniques, but it needs the dependencies on science like genomics, phylogenetics, bioinformatics, etc. while presenting the evidence for criminal and microbial outbreaks. 18 Post anthrax letter case in 2001 in US news agency, immediately after the Twin tower blast, the microbial accenting in an investigation has increased. 19 To start the full-fledged microbial forensic lab, instruments and techniques like gene sequencing, hybridization, microarrays, spectrophotometry, and PCR and expertise are required. 20 The application of traces of touch microbiome will require the study in the direction of research in the cult of how the exchange of hand microbes is happening between the touched surface, the persistence of these microbes in various surface types and contamination of other bacteria. Rigorous practice is required to improve the gap between laboratory-controlled experiments and regular forensic analysis. This requires an understanding of the strength, sensitivity, and shortcomings of microbial forensic techniques. At the University of California, Microbial DNA profiling was done to identify the individual on the basis of the microbial signature associated with the objects that they touch. They found that plastic and ceramic objects were linked more accurately to the individuals with the help of microbial signatures. 21 In another type of experiment of mock burglary crime scene set up in households in Chicago and Florida, they could find significant changes in the microbial signatures across time on the household items to the people. 22 The airborne dust in the dormitory was used to predict the gender on the basis of microbial composition. This technique holds a promising approach in identifying the place where the indoor crime has occurred which was occupied by male or female. 23 Meta-analysis of bacterial 16S rRNA proved to be a much more accurate technique to identify the patterns in indoor environments. However, the validation will require rigorous study in various conditions for results comparisons and to enhance the reliability of these results by setting up methodologies. 24 we spend the majority of our time in indoor environments. Consequently, environmental exposure to microorganisms has important implications for human health, and a better understanding of the ecological drivers and processes that impact indoor microbial assemblages will be key for expanding our knowledge of the built environment. In the present investigation, we combined recent studies examining the microbiota of the built environment in order to identify unifying community patterns and the relative importance of indoor environmental factors. Ultimately, the present meta-analysis focused on studies of bacteria and archaea due to the limited number of high-throughput fungal studies from the indoor environment. We combined 16S ribosomal RNA (rRNA Also, attempts are going on to track the micro-organisms to their source in the context of epidemiology and forensic criminal investigations to implement them in regularized investigation techniques. 25 The research is also going on to complement microbial DNA with human DNA to support the forensic investigations associated with individualization. 26 One of the studies suggests that the changes in the hand microbial community are dependent on the sex, which hand you use often and hand wash frequencies. The deep study of this will also help in identifying the individuals and their daily routines. 27 Microbial community structures on items and palm prints are successfully matched with corresponding palm skin in research. 28 The environmental changes, urbanization, and altered dietary patterns are important for maintaining a balance between environmental and human microbiomes. 29 According to the findings, human-associated microbial communities have enough variation to distinguish individuals over time. As microbiome data is linked to individuals without additional identifying information, ethical implications arise, calling into question the assumptions of complete privacy. 30 Microbial signatures may include ethnicity, which could be useful as evidence in forensic investigations.
Methodology
Isolation of Bacteria from Hands
A total of 40 impressions of hands were collected on labeled Nutrient Agar (NA) plates from the people of four different professions (10 each, i.e., garbage collectors [GC], morgue staff [MW], construction workers [CW], and housekeeping staff [HK] staff) and plates were incubated for 24 hours at 37ºC for the growth of bacteria. Isolates showing similar colony characteristics were selected for further studies.
Isolation of Bacteria from Mobile
Sterile cotton swabs were rubbed on the phones separately in the areas where most of the finger contact occurs and were streaked onto NA plates separately. Plates were then incubated for 24 hours at 37ºC. Isolates showing colony characteristics similar to those from hand impressions of respective individuals were selected for further studies.
Identification of Bacteria
The selected isolates were subjected to gram’s staining for preliminary identification. The isolates were then subjected to DNA Isolation and 16S RDNA sequencing for identification up to the species level. 31
DNA Isolation and PCR Amplification
The DNA from the selected bacterial cultures were isolated using the Phenol-Chloroform method. 32 Bacterial 16S rDNA was amplified using primers 27F forward and 1492R reverse sequence 33 and obtained a 1550 bp long sequence. 34 The amplified DNA samples were subjected to Sanger sequencing and then identification of the bacteria was done by NCBI-BLAST. 35
These sequences were then used for clustering according to the species from different professions 36 and establishing the phylogenetic relatedness to visualize the diversity of the species.
Sequence Processing
The genetic relatedness and clustering of the sequences were established with the help of R Studio by using the packages like seqinr, magrittr, dplyr, ggpubr, gplots, tidyverse, cluster, factoextra, seqinr, adegenet, ape, ggtree, DECIPHER, viridis, ggplot2. 37 The sequences were aligned. The distance matrix calculation, alignment and graphical representation were obtained by writing codes for each step. Then used for plotting clusters, a phylogenetic tree in R studio for establishing relationships between the isolated sequences.
Results and Discussions
Out of 40 persons sampling bacteria from hands and mobile, a total of 250 isolates were selected with similar colony characteristics from person’s hand and mobile of all the professions. The majority of them were gram-positive cocci in clusters and few were gram-positive rods in chains and gram-negative coccobacilli in singles.
A total of 23 isolates which were sequenced were then further analysed by the scatter plot for four professions, that is, GC, CW, HK, and MW. The GC cluster shows overlap with all other clusters of the professions. The variability in all four professions could be due to the professions that they work in. For example, the reason GC overlaps with other professions is due to the reason that GC collects garbage from all other professions and hence the transfer of the organism occurs. After GC, the graph shows the distribution of morgue workers as they handle the bodies from vast areas and there is no clear demarcation from where the bodies come in the morgue. Then it is clearly observed that CW and housekeeping are very well confined to their works and hence show the clustering of the professions (Figure 1).
MDS Plot of the 16S rDNA Sequences.
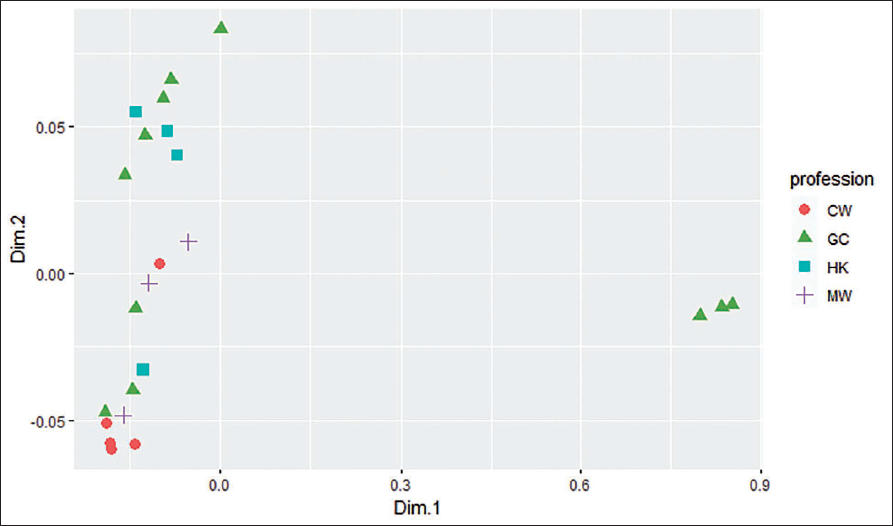
The reason behind the clustering of the Bacillus spp. bacteria present on the hand can be mainly due to materials that they are exposed to, for example, GC and CW are exposed to soil, outdoor dust environment, etc. The hygiene maintained by individuals, lifestyle, location, etc. might have contributed to the formation of the clusters on the basis of profession. For example, finding bacteria typical of morgue workers on a tool could suggest use in that setting. A study conducted on hair and pubic hair microbiota of different hair types and geolocations in California and Maryland found that the microbiota falls into two different clusters according to the places that they belong to. This was represented by non-metric multi-dimensional scaling plots. 38
Heatmap
The specific types of bacteria involved in these similarities can further refine the identification of potential suspects. The heat map generated shows the similarities, slight similarities and dissimilarities on the basis of color demarcation. Dark black colors represent greater dissimilarity, while light white or white colors represent greater similarity and gray colors represent slight similarity among the sequences. The key findings state that the heatmap also shows some interesting relationships between the different types of Bacillus spp. among CW. For example, the CW samples appear to be more similar to the GC samples than they are to the MW or HK samples. This suggests that the CW and GC samples may share some common features that are not shared by the MW and HK samples. The bacterial colonies which were seen evidently in GC, MW, and CW, were not observed in HK evidently. Still, we found a few difficult to interpret and compare with other professions. The collected bacteria from HK showed similarity with all other professions as they touched various objects including dust, organic waste, etc. GC samples show more similarity to HK and CW samples as the nature of their work includes dealing with organic waste, which leads to the transfer of bacteria from their workplace to their hand (Figure 2).
Heatmap of the 16S rDNA Sequences.
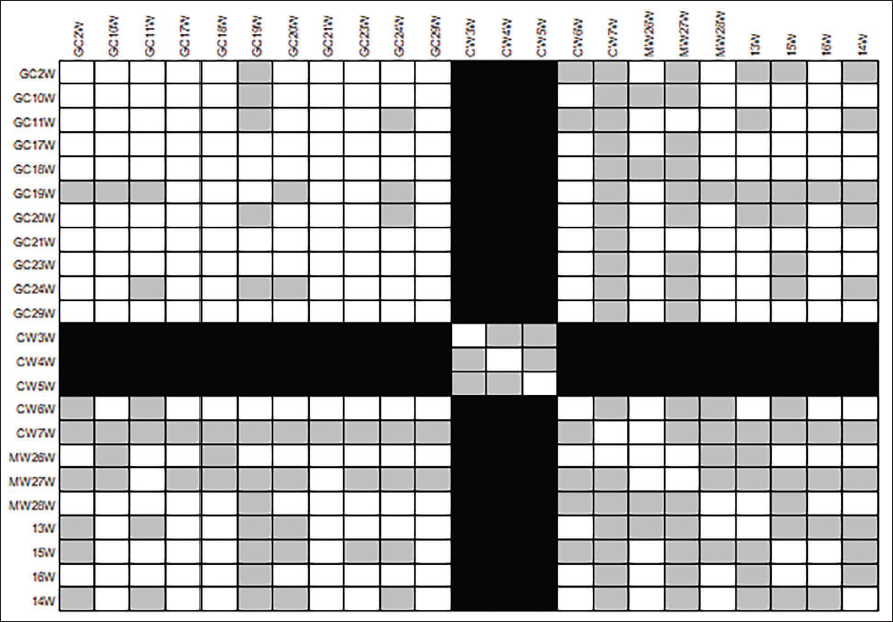

A similar type of analysis was conducted to find out the probability of finding the personal objects with the skin microbiome of the deceased people, where they had rightly predicted the owner of the object. 9 The sequencing of the selected bacteria with similar colony characteristics in terms of color and shape has provided insight into the use of specific bacteria as the markers of the professions. Similar-looking colonies from the suspect’s hands can be collected and compared with the collected microbial swabs from the objects of the crime, where the chances of handling the object are high, like a doorbell, door handle, phones, etc. The careful sample collection and analysis will help us exclude the suspect. For example, finding a bacterial profile similar to HK samples on a weapon could suggest the involvement of someone from that profession. The presence of matching or slightly similar bacterial communities between samples (like objects and hands) can strengthen evidence of contact or proximity.
Cluster Dendrogram
The dendrogram shows three main clusters at a height of about 0.10 on the y-axis. Most of the CW samples are related to each other in most of the branches and to most of the GC samples. This suggests that the bacterial communities on the palms of CW and GC are more similar to each other than they are to the other groups. The cluster dendrogram shows some of the HK and MW samples together in the branch. This suggests that the bacterial communities on the palms of HK and morgue workers are more similar to each other than they are to the other groups (Figure 3).
Locard states that every object leaves a trace on the person who is touching it and the person contributes the traces to the object. 39 The above results support the fact that the person exhibits a microflora similar to their workplace. 40 The bacterial evidence, when aligned with other investigative findings, can strengthen the case against a suspect or provide support for a specific timeline of events. the unique bacterial profiles of different groups offer a new avenue for investigation, potentially identifying previously unexamined suspects or locations.
Cluster Dendrogram of the 16S rDNA Sequences.
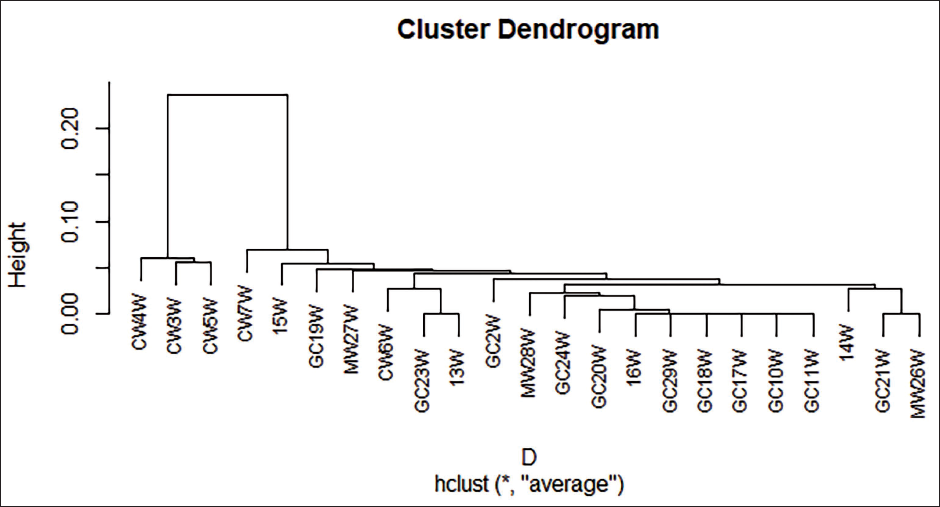
Footnotes
Acknowledgement
We thank Dr. Syed Dastager and his laboratory, without whose engagement this study would not have been successful. We would also like to thank the volunteers who contributed by providing the samples.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship and/or publication of this article.
Ethical Approval
The ethical clearance was approved by the University Ethics Committee under the reference number JU-EC-/022-SC/FS/PhD-JUL2023.
Funding
The authors received no financial support for the research, authorship and/or publication of this article.
Informed Consent
All participants provided written informed consent prior to enrollment in the study.
