Abstract
Objectives
To evaluate the efficacy of non-invasive prenatal testing in detecting sex chromosome aneuploidies.
Methods
A total of 67,099 pregnant women were recruited in this retrospective observational study at Peking Union Medical College Hospital; 300 cases at high risk of sex chromosome aneuploidies were screened and 36 cases refused invasive prenatal diagnosis after detailed prenatal genetic counselling. The clinical data and prenatal diagnosis results were collected.
Results
101 cases were in accordance with non-invasive prenatal testing results after invasive prenatal diagnosis: 95 cases had follow-up confirmation with abnormal sex aneuploidies (14 for monosomy X; 36 for 47, XXY; 22 for trisomy X and 23 for 47, XYY). The total positive predictive value was 38.26% (101/264). The accuracy of different Z scores was calculated, respectively, with positive predictive value ranging from 33.96% to 41.38% and a decreased sensitivity with rise of the Z score.
Conclusions
Non-invasive prenatal testing is an effective screening method for sex chromosome trisomy, rather than monosomy X, with relatively high accuracy. Z score = 3 is an optimum risk threshold in sex chromosome aneuploidies.
Keywords
Introduction
Lo et al. discovered the presence of circulating fetal cell-free DNA (cfDNA) in the plasma of pregnant women in 1997, 1 giving rise to the development of non-invasive prenatal testing (NIPT). NIPT has consistently higher screening performance in the detection of fetal normal chromosomal aneuploidies than any of the traditional screening approaches.2,3 Meta-analysis showed the sensitivity of NIPT for trisomy 21, trisomy 18 and trisomy 13 to be 98.80% (95% CI = 97.81–99.34), 98.83% (95% CI = 95.45–99.71) and 100% (95% CI = 0–100), respectively. 3 NIPT has been widely applied in clinical settings throughout China.
Since 2022, the American College of Medical Genetics and Genomics strongly recommends that NIPT should be offered in screening for fetal sex chromosome aneuploidies (SCAs) for singleton pregnancies, 2 and the International Society for Prenatal Diagnosis also proposes a positive attitude after taking societal, economic, cultural and ethical factors into consideration. 4 However, opinions on whether to use NIPT screening for SCAs are still not completely unified on account of the questionable accuracy and feasibility. A 2016 report from China's General Office of the National Health and Family Planning Commission stated that for chromosomal abnormalities detected by NIPT, other than common chromosomal aneuploidies, pregnant women should be informed and given further consultation and diagnosis. 5 The new expert consensus in China maintains a neutral attitude and only recommends routine reporting of Turner syndrome (45, X) among SCAs. 6 This study retrospectively analyzed data from a single prenatal diagnostic center to provide a reference and basis for NIPT screening of SCAs.
Materials and methods
Study population and samples
The retrospective study enrolled pregnant women aged 18–50 with a gestational age of 13–27 weeks for NIPT at Peking Union Medical College Hospital, Beijing, China, from January 2018 to April 2025. A total of 67,099 singleton pregnancies were included. Pre-test clinical counselling was provided to all participants. SCAs were detected in 300 cases, and of these 36 refused invasive prenatal diagnosis after detailed prenatal genetic counselling.
The study has been approved by the Institutional Ethics Committee of Peking Union Medical College Hospital (reference number K23C1179) and all participants signed written informed consent prior to the test. The research was conducted in accordance with the relevant guidelines and clinical norms and followed the STROBE guidelines.
NIPT
A whole blood sample of 10 mL was collected into ethylene diamine tetra acetic acid anticoagulant vacutainers. After two centrifugal separations of 10 min at 1600 g and 10 min at 16000 g within 48 h, maternal plasma was separated and reserved. With the use of a DNA extraction and purification kit (R0011, Berry Genomics Corporation, Hangzhou, China), the cell-free DNA (cfDNA) was extracted, and the concentration of total cfDNA was measured by Qubit 3.0 fluorometer. If the concentration of total cfDNA qualified (reference range 0.05–0.70 ng/mL), the samples were subjected to library construction using the Library Preparation and Purification Kit (R0022, Berry Genomics Corporation). Massively parallel sequencing (MPS) was performed on the Illumina NextSeq CN 500 sequencing platform (high-throughput sequencing kit), and the sequencing data was mapped to the human genome reference sequence (hg19, NCBI build36). The mapping results were analyzed by the Bambini Test system. Z score was calculated upon repetitive sequences removal, effective read calculation, and GC correction. The Z score represents the standardized deviation of the read count proportion of a given chromosome in a test sample from a reference euploid population, which was calculated as the difference between the observed and expected chromosomal representation divided by the standard deviation of the reference dataset. In the context of NIPT, an increased or decreased Z score reflects an over- or under-representation of sequencing reads from a specific chromosome, corresponding to potential chromosomal aneuploidy. The fetal aneuploidy status for all 24 chromosomes was determined based on Z scores (normal range, −3 < Z < 3). The absence of results in a sample was attributed to insufficient fetal fraction (<4%) of cfDNA or failure to pass quality control measures.
In cases before 2021, the Z scores of chromosome X and Y should have a linear relationship with each other for the male samples, and the Z scores of chromosome X should range from −2.91 to +2.91 while the Z scores of chromosome Y should be less than 3 for the female samples, since the copy numbers of sex chromosomes were proportional to the percentage of fetal DNA in the maternal plasma. According to the calculation method proposed by Liang in 2013, 7 those samples that were outliers to these two categories were potential SCA samples. After 2021, the analytical algorithm of Z scores of sex chromosomes was updated and the Z scores of both chromosome X and Y should range from −3 to +3. Those samples that had abnormal Z scores of sex chromosomes would be considered as SCA samples.
Invasive prenatal diagnosis
Following the results of NIPT, amniocentesis was performed under ultrasound guidance at 17–22 gestational weeks. Chromosome karyotyping analysis (G banding, 300–400), fluorescence in-situ hybridization analysis (F01001–01, Beijing GP Medical Technologies, Ltd, Beijing, China) and chromosomal microarray analysis using a CytoScan 750 K array (Affymetrix, Santa Clara, CA, USA) were performed according to standard protocols.
Conformation of maternal karyotypes by genomic DNA sequencing
The maternal leukocyte layer was used to extract maternal DNA and to construct the library. The Illumina NextSeq CN 500 sequencing platform was used for MPS, and the genomic reference sequence (hg19, NCBI build36) was used to map the sequencing results. Using the fused lasso algorithm, the gain or loss of chromosome regions was identified. 8
Statistical analysis
Version 22.0 of the Statistical Product and Service Solutions (SPSS) software (IBM, Armonk, NY, USA) was used for statistical analysis. Measurement data were presented in the form of medians ± interquartile ranges (IQRs) and count data adoption rate (%). The sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV) of NIPT for detecting fetal SCAs were calculated.
Results
A total of 300 (0.45%) cases were detected by NIPT as being at high risk of SCAs among 67,099 singleton pregnancies, including 138 results for monosomy X, 68 for XXY, 56 for XXX and 31 for XYY. In seven cases NIPT results indicated maternal aneuploidy for chromosome X, with six reporting X (increased)-M and one reporting X (decreased)-M (M = maternal). The median ± IQR maternal age of the 300 patients was 33 ± 6 years and the median gestational age at testing was 15 ± 4 weeks. After detailed prenatal genetic counselling, 264 (88.0%) patients agreed to invasive prenatal diagnosis. The entire process of the study is shown in Figure 1.
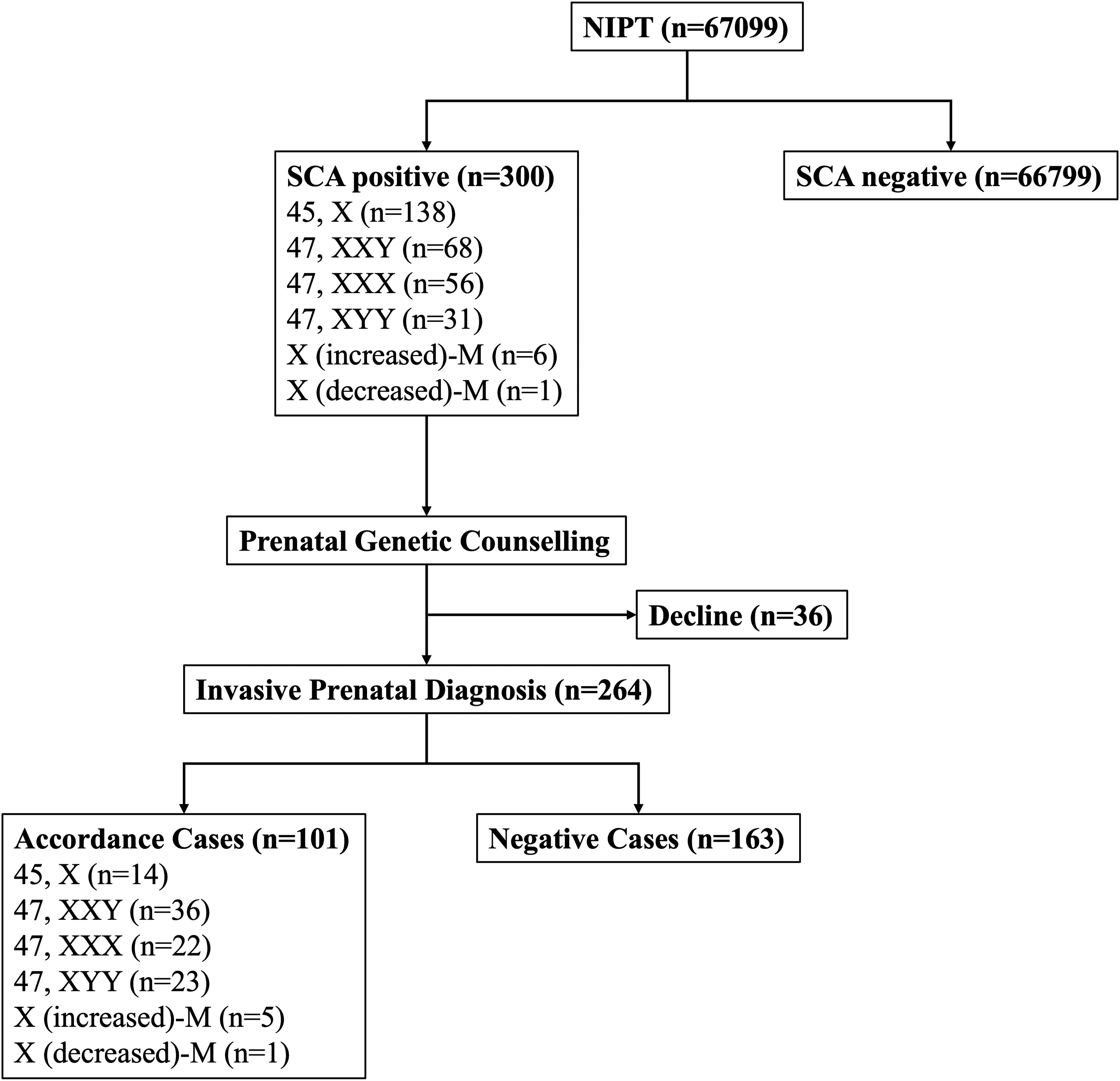
Study flowchart summarizing NIPT for detecting fetal SCA. NIPT: non-invasive prenatal test; SCA: sex chromosome aneuploidy; M: maternal.
Accuracy of NIPT in the screening of SCAs
In 101 cases, the NIPT results were consistent with invasive prenatal diagnostic findings (see Table 1). There was follow-up confirmation in 95 cases of abnormal sex aneuploidy (14 for monosomy X; 36 for 47, XXY; 22 for trisomy X; 23 for 47, XYY). There were seven cases indicating maternal chromosome X aneuploidy, with six cases confirmed. The only case of X(decreased)-M was confirmed as maternal mosaic X chromosome karyotype: 30% 45, X and 70% 46, XX. The total incidence rate was 0.15 (101/67099) and the PPV was 38.26% (101/264).
NIPT and invasive prenatal diagnosis results for 300 cases.
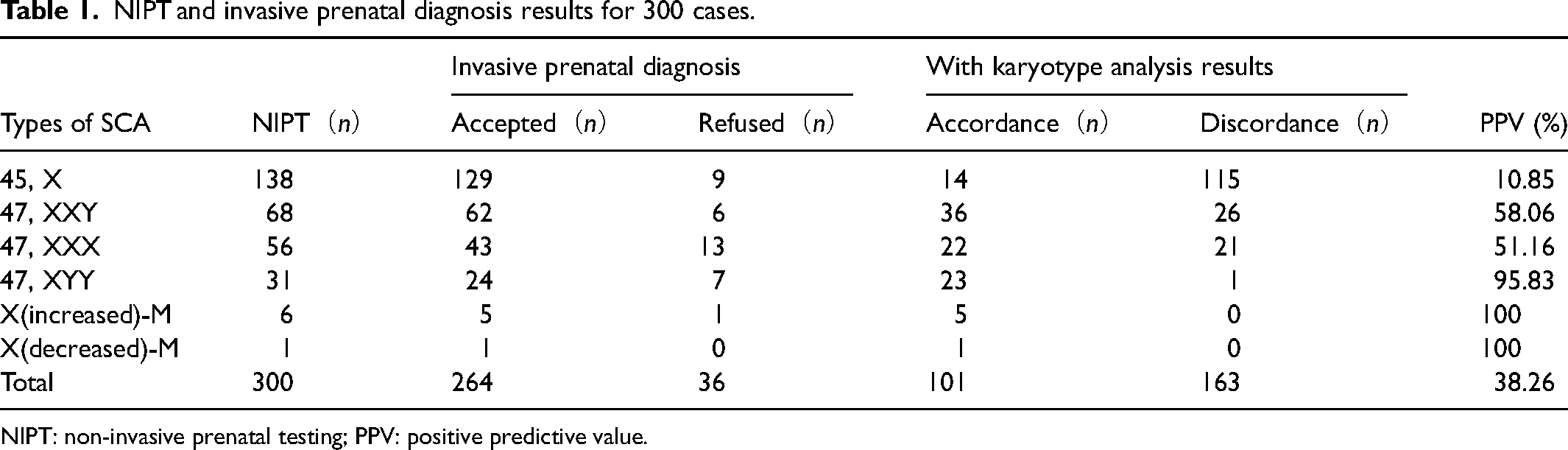
NIPT: non-invasive prenatal testing; PPV: positive predictive value.
Comparison of NIPT in the screening of SCAs with different Z scores
After 2021, 139 (0.37%) of 37,364 cases were reported as being at high risk of SCAs with Z scores beyond the interval range of 3. The screening performance of NIPT was evaluated using increasing Z score thresholds (see Table 2). The PPV remained relatively stable across different thresholds, ranging from 33.96% to 41.38%, and the NPV remained consistently high (≥99.93%). As the Z score threshold increased from ≥3 to ≥7, sensitivity decreased progressively from 100% to 40.91%, while specificity increased from 99.77% to 99.91%. Correspondingly, the false-positive rate declined from 0.23% to 0.09%, whereas the false-negative rate increased from 0% to 59.09%. These results indicate that lower Z score thresholds maximize sensitivity and minimize the risk of false-negative SCAs, whereas higher thresholds reduce false-positive rates but are associated with a substantial increase in missed cases.
Application of NIPT in the detection of fetal SCAs with different Z scores.
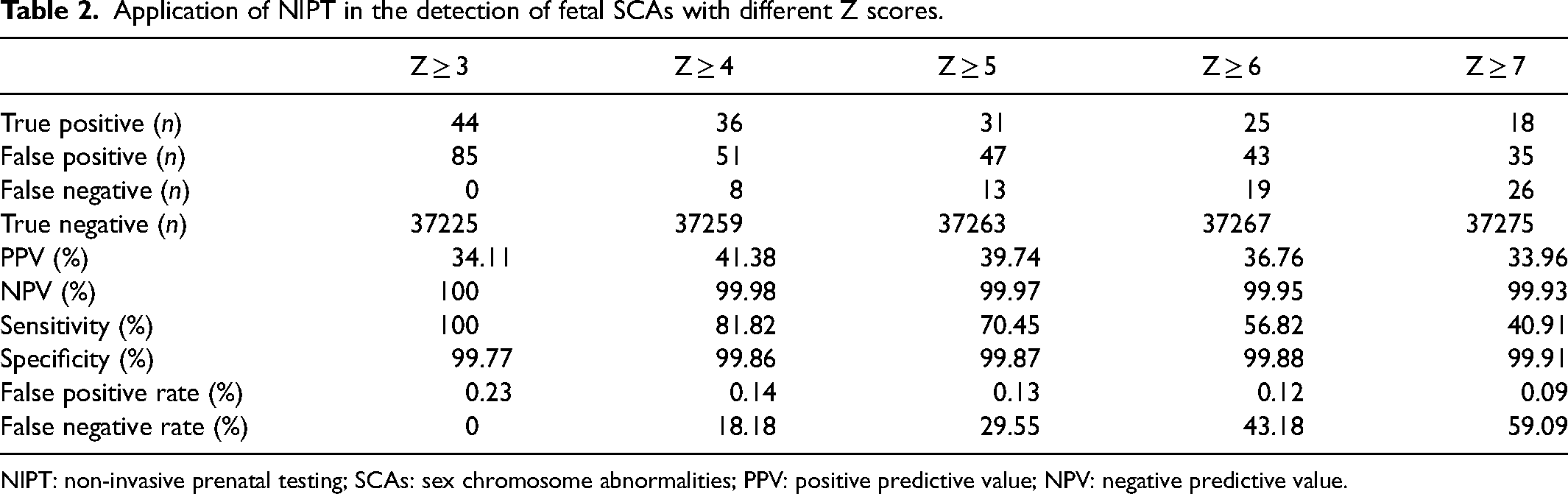
NIPT: non-invasive prenatal testing; SCAs: sex chromosome abnormalities; PPV: positive predictive value; NPV: negative predictive value.
Discussion
This single-center study presents data from a Chinese cohort of pregnancies with high risk of SCAs detected by NIPT. Our study demonstrated that: (1) NIPT is an effective screening method for SCAs, with higher detection efficiency in sex chromosome trisomy than monosomy X; (2) Z score = 3 is an optimum risk threshold in SCAs.
The screening performance
NIPT for SCAs is more complicated than for autosomal aneuploidy and can be confounded by a variety of maternal and fetal biological phenomena. From 2022, there has generally been a positive attitude in statements about NIPT screening for SCAs.2,4 The sensitivity and specificity of NIPT for SCAs have been reported as 99.63% and 99.80%, respectively, and PPVs, which varied slightly, were about 30%–74% and false positive rates 0.0%–0.1%.3,9–13
In our cohort of 66,799 singleton pregnancies, all low-risk NIPT results were followed up until delivery, yielding no confirmed false-negative cases. Based on this large dataset, the sensitivity of NIPT for SCAs was 100%, with specificity ranging from 99.83% to 99.99% (see Table 3), which is largely consistent with the previously published reports.
Comparative performance metrics of NIPT for SCAs: single-center cohort versus systematic review.
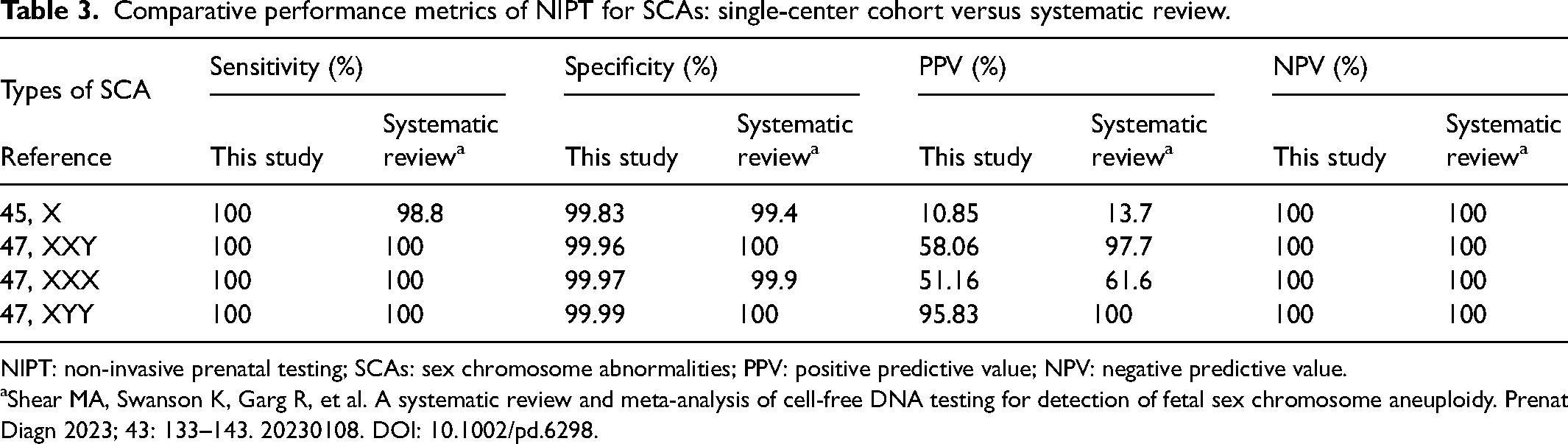
NIPT: non-invasive prenatal testing; SCAs: sex chromosome abnormalities; PPV: positive predictive value; NPV: negative predictive value.
Shear MA, Swanson K, Garg R, et al. A systematic review and meta-analysis of cell-free DNA testing for detection of fetal sex chromosome aneuploidy. Prenat Diagn 2023; 43: 133–143. 20230108. DOI: 10.1002/pd.6298.
The overall positive predictive value (PPV) in our cohort was 38.26%, with PPVs for 45, X, 47, XXY, 47, XXX, and 47, XYY being 10.85%, 58.06%, 51.16%, and 95.83%, respectively. These values were compared with a previous systematic review, 14 which reported PPVs of 13.7%, 97.7%, 61.6%, and 100% for the same karyotypes (Table 3). Overall, the trend in PPVs is similar, although our single-center PPVs are lower for several SCAs. This may be due to multiple factors, including referral patterns to a tertiary hospital where a mixture of high-risk pregnancies and borderline cases increases the number of false positives, the relatively small absolute number of positive cases in a single center leading to wider statistical fluctuations, and minor differences in analytical algorithms used across studies.
Among the 300 cases screen-positive for SCAs, 36 women declined invasive prenatal diagnostic testing and therefore did not have cytogenetically confirmed fetal karyotypes. Within the regional maternal and child healthcare system, all pregnancies are routinely followed through secondary and primary health institutions, and pregnancy outcomes are systematically reported to our center. According to postnatal follow-up records, no clinically apparent abnormalities were identified in the newborns delivered by these women based on pediatric examinations. In the present study, screening performance metrics were calculated exclusively using cases with confirmed invasive prenatal diagnostic results, in order to ensure consistency and methodological rigor.
Importantly, the negative predictive value (NPV) in our cohort was 100%, consistent with the systematic review. 14 This indicates that NIPT is highly reliable for ruling out SCAs in a large population, and the risk of missed cases (false negatives) is extremely low, which is crucial for prenatal screening and clinical counselling.
Comparison across different Z scores
Since each platform has its own unique analysis algorithms and different labs apply different cut-offs, Z scores, or normalization controls, the same maternal blood sample examined at different labs may produce inconsistent SCA screening findings. 15 At present, Z score = 3 is generally used as the risk threshold in common chromosomal aneuploidies,5,16–18 although some assays may use different values. Research has shown that posterior risk is effectively independent with a lower a priori risk only being reached at Z score > 5,19,20 but large-scale clinical studies to validate this hypothesis are lacking. However, some groups have indeed conducted research on the setting of Z score and drawn corresponding conclusions. Yun Tian et al. found that the occurrence of confined placental mosaicism may slightly elevate a Z score in NIPT, higher than 3 but lower than 5 for normal chromosomal aneuploidies. 21 Xinxin Tang et al. discovered that for rare autosomal trisomies, when the Z score is ≥15 the occurrence of adverse pregnancy outcome will see a rise. 22
In the screening of SCAs, there is currently no unified Z score threshold for NIPT. To better evaluate the impact of different thresholds on screening performance and clinical utility, we analyzed NIPT results using a series of increasingly stringent Z score cut-offs (Z ≥ 3, ≥ 4, ≥ 5, ≥ 6, and ≥ 7). In our cohort, increasing the Z score threshold was associated with a progressive reduction in the false positive rate, accompanied by a substantial rise in the false negative rate. While the PPV showed only modest variation across thresholds (ranging from 33.96% to 41.38%), the sensitivity declined markedly from 100% at Z ≥ 3 to 40.91% at Z ≥ 7, indicating a significant loss of detectable cases as stricter cut-offs were applied. In contrast, the NPV remained consistently high (>99.9%) across all thresholds. False positive rate decreased slightly with higher Z score thresholds; however, this improvement was achieved at the expense of a pronounced increase in missed cases. Taken together, our data indicate that a Z score threshold of 3 provides the most appropriate balance between sensitivity and specificity for SCA screening, supporting its use as a clinically compatible risk cut-off for NIPT-based population screening and genetic counselling.
Reasons for lower PPV in 45, X
Although current guidelines recommend NIPT for detecting SCAs in singleton pregnancies, its screening performance varies substantially across different SCAs. In our cohort, NIPT performed well for sex chromosome trisomy but showed a markedly lower PPV for fetal monosomy X, consistent with previous reports.3,14 Several large-scale studies in China have reported similar findings, with PPVs for monosomy X ranging from approximately 12% to 29%.23–27 Despite the introduction of genome-wide and comprehensive NIPT approaches into clinical practice,28,29 several studies have reported no significant improvement in the detection accuracy for monosomy X, with the reported PPV ranging from approximately 17.5% to 25.8%.30–32 Our slightly lower PPV may reflect the high rate of confirmed prenatal diagnosis and rigorous post-test follow-up in our study, which provides a more realistic representation of NIPT's detection efficacy for 45, X in real-world settings.
Several biological and technical factors contribute to the lower PPV for monosomy X. These include a higher rate of placental mosaicism, 33 maternal mosaicism or age-related X chromosome loss,9,20,34,35 and technical challenges associated with sequencing and amplifying the X chromosome.27,36 In our cohort, 22 of 115 discordant cases with NIPT-predicted monosomy X were found to be maternal monosomy X upon verification of maternal peripheral blood chromosomes. Additional intrinsic biological factors, including low fetal fraction of cfDNA, confined placental mosaicism, and other maternal or fetal variants, further limit the detection accuracy. 27 Because only a small fraction of cfDNA in maternal plasma is fetal in origin, fetal variants can only be reliably detected when the genetic signal surpasses assay variability.
Recent advances, including second-generation NIPT approaches such as coordinative allele-aware target enrichment sequencing (COATE-seq), 37 have shown promise in improving detection of 45, X. In a prospective study 38 of 1191 pregnancies at elevated genetic risk, comprehensive NIPT using this approach increased the PPV for 45, X to 88.2%. While encouraging, these findings still require validation in large-sample, population-based studies.
Implementation experience
Our NIPT screening program at Peking Union Medical College Hospital has offered valuable insights beyond standard performance metrics. Several challenges were encountered, including the low PPV for monosomy X, which presents the most severe clinical phenotype among SCAs, making genetic counselling particularly complex, and the small proportion of patients who declined invasive prenatal diagnosis. Key lessons learnt highlight the importance of thorough pre- and post-test counselling, long-term follow-up, and ongoing optimization of analytical thresholds and algorithms, including Z score thresholds. The program has evolved from initially screening only common chromosome aneuploidies to including SCAs and copy number variants, and the recent implementation of COATE-seq has improved overall PPV. From a cost-effectiveness perspective, a prior study 39 in high-risk populations has demonstrated that NIPT is more efficient, safer, and economically favorable compared with invasive diagnostic strategies. Finally, the program's strengths include a consistently high NPV (100%) and a robust referral and follow-up system across tertiary and community hospitals, ensuring nearly complete coverage and reliability in real-world clinical practice.
Strengths and limitations
Compared with other similar studies, our study has the strengths of a larger sample size, and being the first to assess the Z score setting in NIPT for SCAs. There are also several study limitations. First, we followed all 66,799 women with singleton pregnancies who obtained low-risk results in NIPT until parturition from the community hospital to our tertiary hospital, but we didn’t conduct placental karyotyping to determine placental mosaicism. Second, Z score = 3 for SCAs in NIPT was only applied after 2021, so only 37,364 cases were analyzed for the relationship between Z score and detection efficiency. Finally, due to being a single-center study, we could not compare NIPT SCA screening efficiency using different technology platforms. Further studies are needed to determine if there is an optimal NIPT methodology for the detection of SCA.
Conclusions
NIPT is an effective screening method for sex chromosome trisomy, rather than monosomy X, with relatively high accuracy. Z score = 3 was shown to be an optimum risk threshold in SCAs. Detailed and comprehensive genetic counselling before and after testing is of great importance. All cases with high-risk results should undergo invasive prenatal diagnosis and karyotype examination of maternal peripheral blood for confirmation, decision, intervention, and treatment. There should also be a reminder of residual risks for cases with low-risk results.
Footnotes
Acknowledgments
We would like to express our gratitude to all the patients included in the study, as well as the clinical and research personnel who supported the recruitment, care, and follow-up of these participants.
Author contributions
YQY: data collection, analysis and interpretation, and manuscript writing. QWQ, XYZ, YLJ, NH, MML, and XTY: data collection. JTL: study conception and design, data collection, and revision. QWQ: study conception and design, data collection, analysis and interpretation, manuscript writing, editing, and revision. All the authors have read and approved the manuscript.
Ethical considerations
This study was approved by the Institutional Ethics Committee of Peking Union Medical College Hospital (Ethics Code: K23C1179). This research was conducted ethically in accordance with the World Medical Association Declaration of Helsinki.
Consent to participate
Written informed consent was obtained from all participants.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by National High Level Hospital Clinical Research Funding (2022-PUMCH-B-076) and the National Key Clinical Specialty Construction Project (U114000).
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data availability
The datasets generated and/or analyzed during the current study are not publicly available to comply with hospital and IRB policies. However, they are available from the corresponding author upon reasonable request.
