Abstract
Objective
To investigate the factors associated with cell-free DNA test failure, and the optimal subsequent management of these pregnancies.
Methods
This was a retrospective study of 27,363 singleton pregnancies undergoing cell-free DNA testing. Women with cell-free DNA test failure were divided into a high-risk group and a low-risk group according to their indications. The subsequent management and pregnancy outcomes of these women were followed up.
Results
The rate of cell-free DNA test failure at the first sampling was 1.49%, and 78.4% of failures were due to a low fetal fraction. Of the 66 women who refused any subsequent management, an adverse pregnancy outcome was seen in 5 cases, all belonging to the high-risk group. Of the 13 low-risk women who chose second-trimester maternal serum screening, all obtained a low-risk maternal serum screening result and an unaffected pregnancy outcome. A redraw was chosen by 171 women, which yielded a result in 75.4% and their pregnancy outcomes were unaffected; 42 women had an uninformative result again and received an amniocentesis. As 158 women had an amniocentesis after the first sampling, this procedure was offered in 200 cases altogether. Abnormal genetic testing results were shown in six (3%, 6/200) cases, all in the high-risk group.
Conclusions
High-risk pregnant women with cell-free DNA test failure are at increased risk of adverse pregnancy outcomes. A second sampling for cell-free DNA test or maternal serum screening might be suggested to low-risk women. Invasive prenatal diagnosis should be offered to the high-risk patients, especially those with a second cell-free DNA test failure.
Keywords
Introduction
Next-generation sequencing-based cell-free DNA (cfDNA) testing has been a part of clinical practice since 2011. Although the cfDNA test has been proved to be the most effective screening method for common fetal chromosomal aneuploidies, there is a certain rate of test failure, which means the results are not reported, indeterminate, or uninterpretable after first sampling. In a recent meta-analysis, Gil et al. suggested three main reasons for the failure to obtain a result from a cfDNA test: (a) problems with blood collection, labeling and transportation, including inadequate blood volume, hemolysis, incorrect labeling of tubes, and delay in arrival at the laboratory; (b) low fetal fraction (FF), usually below 4%; and (c) assay failure for a variety of reasons, including failed DNA extraction, amplification, and sequencing. 1
Two previous multicenter studies have discussed the factors associated with cfDNA test failure. In their historical cohort study, Chan et al. analyzed 12,033 cases of cfDNA test performed using two test platforms (Harmony® and GeneSyte™), and the rate of test failure was 1.1%. They concluded that women who failed to obtain a result from a cfDNA test were at increased risk of adverse pregnancy outcomes, in particular chromosomal aneuploidy, gestational diabetes, and pre-eclampsia. 2 In the prospective study of Galeva et al., 23,495 singleton pregnancies undergoing cfDNA testing at 10 + 0 to 14 + 1 weeks’ gestation were analyzed and the rate of no result after first sampling was 3.4%. They concluded that maternal age, weight, racial origin and parity, and gestational age, method of conception, and serum levels of free β-human chorionic gonadotropin (β-hCG) and pregnancy associated plasma protein A (PAPP-A) are independent predictors of cfDNA test failure. 3
Though the two studies implied an association between cfDNA test failure and adverse pregnancy outcomes, they did not discuss the subsequent management. Both patients and physicians are faced with a dilemma after receiving a cfDNA test failure result. Should they choose to have a redraw, an alternative aneuploidy screening method, or an invasive diagnostic test? Currently there are no guidelines on the subsequent management of pregnancy after an uninformative cfDNA test.
In this study, we retrospectively reviewed 27,363 consecutive cases of cfDNA test performed at a single tertiary center in China during 1 January 2016 to 31 December 2019, and identified 408 cases in which an initial sampling gave a failed result. In this paper, a cfDNA test failure is defined as an inconclusive result at first sampling due to FF below 4% or other laboratory assay failures. The aim of this study was to investigate the rate of cfDNA test failure, the factors associated with test failure, and the variables that affect the chance of an informative redrawn cfDNA test. Understanding these questions should be helpful for choosing optimal subsequent management of these pregnancies.
Methods
Study design, participants, and setting
This was a retrospective study. The participants were among the singleton pregnant women who undertook cfDNA testing in Peking Union Medical College Hospital, a tertiary center for prenatal diagnosis and genetic counseling, during 1 January 2016 to 31 December 2019. Consecutive cases who failed to achieve a result after first sampling were included in the study. All women signed a written informed consent before blood collection.
Procedure
Before the cfDNA test, all the women underwent a sonographic assessment to make sure that the thickness of nuchal translucency of the fetus was normal and no other abnormalities were detected. Pre-test counseling regarding the scope, risk, and limitations of cfDNA testing was provided by obstetricians. Those cases meeting one of the following criteria were excluded: fetal structural abnormalities detected by ultrasound; blood transfusion, stem cell therapy, or immunotherapy within a year; known malignant tumors; either one of parents with chromosomal abnormalities; a history of birth defects. The gestational weeks of all the women undergoing cfDNA testing were between 12 + 0 and 20 + 0 weeks. The high-risk group consisted of women of advanced maternal age (AMA) (aged ≥35) or with a high-risk maternal serum screening (MSS) result (cut-off 1/270 for trisomy 21 and 1/380 for trisomy 18). The low-risk group consisted of women with an intermediate-risk MSS result (cut-off between 1/270 and 1/1000 for trisomy 21 and between 1/380 and 1/1000 for trisomy 18), and those who had no risk factors, i.e. aged younger than 35 and asked for cfDNA test directly.
cfDNA test
From each pregnant woman, 10 ml peripheral blood was collected by EDTA anticoagulant tube and the plasma was separated following a double centrifugation protocol within 48 h of collection. cfDNA was extracted using a DNA extraction and purification kit (Berry Genomics Corporation) and then measured using a Qubit 3.0 fluorimeter (Thermo Fisher Scientific). The reference range of qualified cfDNA was 0.05–0.70 ng/µl. cfDNA library preparation, sequencing, and data analysis were performed using the Bambni™ assay (Berry Genomics Corporation). Briefly, cfDNA was blunt-ended and A-tailed, followed by ligation of Illumina Truseq adaptors and purification. Library concentration was determined by fluorescence quantitative PCR (Kapa Biosystems). NextSeq CN500 sequencing platform (Berry Genomics Corporation) was used for massive parallel sequencing (MPS). The sequencing data were mapped with human genome reference sequence (hg19, NCBI build36) and processed with the Z-score-based Bambni data analysis system (Berry Genomics Corporation).4,5 The sequences mapped to each chromosome were counted and the GC content was calculated. A reference set of female diploid samples was generated. The normalized chromosome representation (NCR) = count of the sequences uniquely mapped to the chromosome of interest/total count of the sequences uniquely mapped to all the autosomal chromosomes. The NCR values were plotted against the GC content and the slope was calculated by simple linear regression. The NCR was further GC-corrected as follows: NCRgc = NCR – (GC – GCaverage_ref)/Sloperef, where GC was the chromosome GC content of the test sample, and GCaverage_ref and Sloperef were the average values of the reference samples, respectively. For chromosome 14 and chromosome Y, Z-score for chromosome i in test case = ((NCRi in test case) − (mean NCRi in reference controls))/(standard deviation of NCRi in reference controls). 6 For any other chromosomes, Z-score for chromosome i in test case = ((NCRgci in test case) − (mean NCRgci in reference controls))/(standard deviation of NCRgci in reference controls). 6 Chromosomes with a Z-score between –3 and 3 were defined as disomic (namely, copy number = 2), whereas chromosomes with a Z-score of ≥3 were considered trisomic (namely, copy number = 3).
Criteria for cfDNA test failure
Criteria for cfDNA test failure were as follows: (1) low FF, which was defined as <4%; (2) high concentration of total cfDNA (>0.70 ng/µl), which indicated the contamination of genomic DNA; and (3) data fluctuation, including GC fluctuation and Z-score fluctuation, which were caused by sample degradation, library preparation, and other technical reasons. GC fluctuation was defined as GC% ≥42.5% or GC% ≤36.5%. Z-score fluctuation means that the sample harbors significant multiple chromosomal aneuploidies, most likely due to test failure but not bona fide chromosomal aneuploidies. Z-score fluctuation was defined as: (I) Z-score of any two of chromosome 13, 18 and 21 > 3 or (II) Z-score of any three or more chromosomes >3. In those cases with high concentration of total cfDNA or data fluctuation, a repeat assay was routinely performed with the original sample in our laboratory. A test failure was considered for those who failed again in the repeated test with the original sample.
Estimate of fetal DNA fraction
For male pregnancy samples, the FF was measured by quantifying the presence of chromosome Y using the methodology described by Hudecova et al. 7 The chromosome Y sequences in maternal plasma are contributed by the male fetus (chrY% fetus ) and sequences originating from the maternal background DNA (chrY% background ) that were incorrectly assigned to chromosome Y. ChrY% background is measured from female pregnancy samples and chrY% male is obtained from an adult male individual. 7
For female pregnancy samples, we used SeqFF containing two regression models (elastic net (Enet) and weighted rank selection criterion (WRSC)) to predict the FF. 8 Briefly, the genome was divided into 50-kb bins, the counts of single-end sequence reads within each bin from autosomal chromosomes (except 13, 18 and 21) were normalized and used as predictor variables, and the model coefficients were determined by making use of Enet and WRSC standard multivariate regression models trained on a large number of male pregnancy samples. 8
Clinical follow-up for cfDNA test failure cases
Further genetic counseling was offered to the women with cfDNA test failure. For the low-risk patients, a fresh blood sample for cfDNA testing or second-trimester MSS (alpha fetoprotein, free β human chorionic gonadotropin, unconjugated estriol) was recommended. For high-risk patients, invasive prenatal diagnosis (IPD) such as amniocentesis was suggested first. Those who refused IPD would sign a refusal form, and then a fresh blood sample for cfDNA test would be recommended. Those who refused any subsequent management would sign a refusal form.
Karyotyping was performed after amniocentesis, and chromosomal microarray analysis (CMA) was suggested to those with a fetal structural abnormality detected by ultrasound. Information including genetic testing results, ultrasound examination results, and pregnancy outcomes were collected after amniocentesis. The pregnancy outcomes were followed up three months after delivery.
Karyotyping analysis
Karyotyping analysis using GTG-banding was done according to standard procedures. A total of 20 metaphase cells were analyzed for each sample. Karyotypes were described according to the International System for Human Cytogenetic Nomenclature.
Chromosomal microarray analysis
Genomic DNA of the fetus was extracted from amniotic fluid using the QIAamp DNA Blood Mini Kit (Qiagen, Valencia, CA, USA). The DNA was digested, ligated with adaptors, amplified, purified, and labeled with biotin. Then, the labeled DNA was hybridized to the Affymetrix® CytoScanTM 750K Array (Affymetrix, Santa Clara, CA, USA). The arrays were washed with Affymetrix GeneChip® Fluidics Station 450 and scanned with an Affymetrix GeneChip® Scanner 3000 according to the manufacturer’s protocol. CEL files obtained by scanning the arrays were analyzed with Chromosome Analysis Suite v33.1 software. The GRCh37 (hg19) genome was used for annotation. Copy number variations that were larger than 200 kb or affected more than 50 markers were considered.
Statistics
The statistical differences in clinical characteristics (weight, height, gestational age of fetus, body mass index (BMI)) between those who obtained a cfDNA test result and those with a test failure were determined using student’s t test. The statistical difference in the distribution of high-risk patients between each series of women with different reasons for a cfDNA test was determined using chi-squared test. All data were analyzed by SPSS 19.0 software (SPSS Inc, USA). A p value of < 0.05 was considered statistically significant.
Results
Study population
A total of 27,363 women with singleton pregnancies received a cfDNA test during 1 January 2016 to 31 December 2019 in Peking Union Medical College Hospital. Among these pregnancies, 1070 were conceived by in-vitro fertilization (IVF), accounting for 3.91% of the population. Compared with those women who obtained a cfDNA test result, the maternal weight and BMI of the cfDNA test failures were significantly higher (p < 0.001 and p < 0.001 respectively), and the gestational age was significantly lower (p < 0.001). There were no differences in the average maternal age and maternal height between these two populations (p = 0.941 and p = 0.598 respectively). Maternal and pregnancy characteristics of the 27,363 single pregnancies are summarized in Table 1.
Maternal and pregnancy characteristics of the 27,363 single pregnancies.
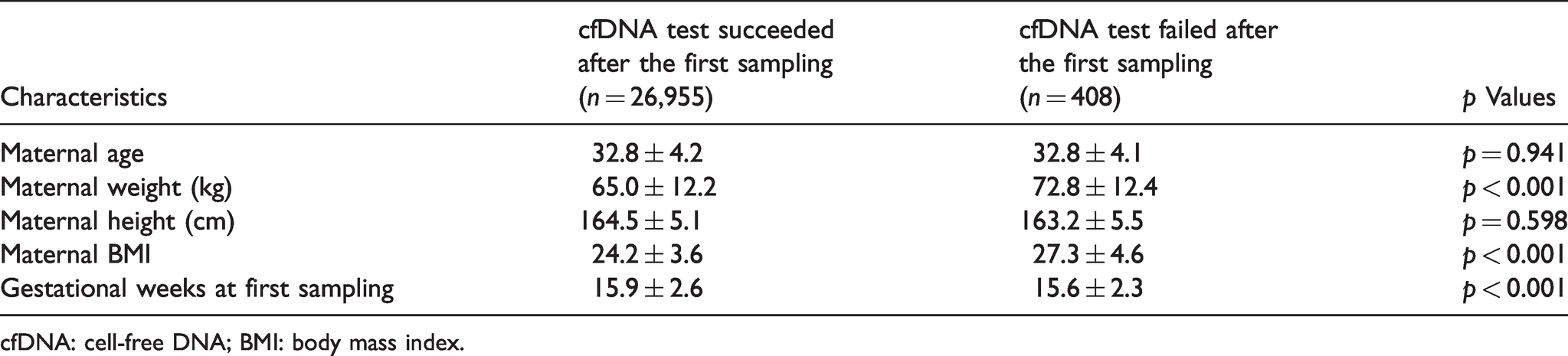
cfDNA: cell-free DNA; BMI: body mass index.
The rate of cfDNA test failure and characteristics of the cases
Altogether, 408 cases failed to obtain a result after first sampling, giving a test failure rate of 1.49% (408/27,363). Among the 1070 cases of IVF pregnancy, 17 failed to obtain a result (1.59%, 17/1070). There was no statistical difference in the rate of cfDNA test failure between these two populations (p = 0.796). Because the sample size among IVF pregnancies was relatively small, we do not discuss the subsequent management of the IVF group in this study.
Among the 408 cases of cfDNA test failure at the first sampling, 320 (78.4%, 320/408) were because of low FF, 18 (4.4%, 18/408) because of high concentration of total cfDNA, and 70 (17.2%, 70/408) because of data fluctuation.
Of the 320 cases with a low FF, 134 (41.9%, 134/320) were in the high-risk pregnancy group, including 130 cases of AMA and 4 cases of high-risk MSS result. The remaining 186 (58.1%, 186/320) cases in the low-risk group included 38 cases of intermediate-risk MSS result and 148 cases with no risk factors.
Of the 18 cases with high concentration of total cfDNA, 6 (33.3%, 6/18) were high risk, and they were all AMA women. The other 12 (66.7%, 12/18) were low risk, all with no risk factors.
Of the 70 cases of data fluctuation, 28 (40%, 28/70) were high risk, including 27 cases of AMA and 1 with a high-risk MSS result. The other 42 (60%, 42/70) cases were low risk, including 4 with intermediate-risk MSS result and 38 with no risk factors.
In total, of the 408 cases with a cfDNA test failure result, 168 (41.2%, 168/408) belonged to the high-risk group, including 163 cases of AMA and 5 with a high-risk MSS result. The other 240 (58.8%, 204/408) cases belonged to the low-risk group, including 42 with intermediate-risk MSS result and 198 with no risk factors.
The distribution of the cases with cfDNA test failure after first sampling is shown in Table 2. Chi-squared test revealed that the distribution of high-risk and low-risk patients among the three groups with different reasons for cfDNA test failure has no statistical difference (p = 0.76).
The distribution of cases with a cfDNA test failure result.
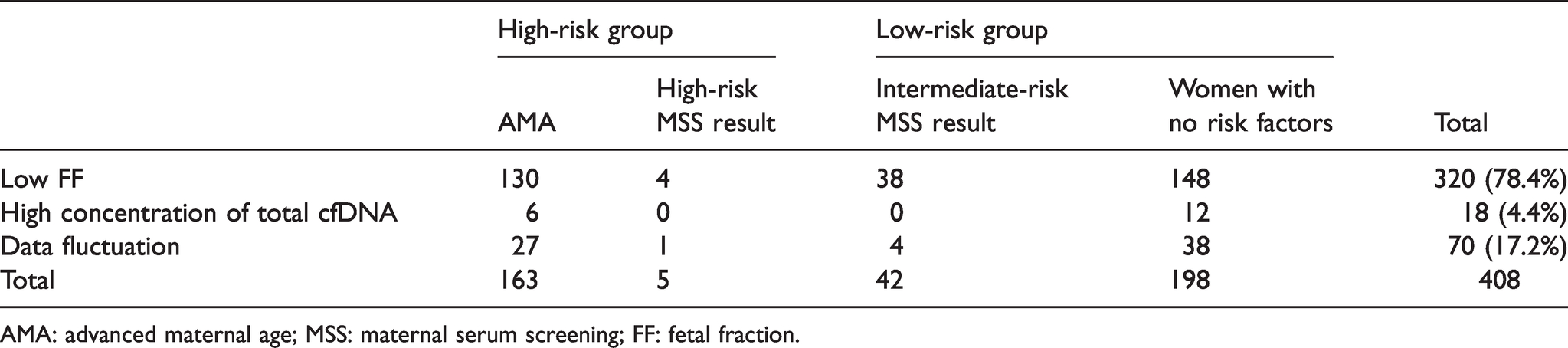
AMA: advanced maternal age; MSS: maternal serum screening; FF: fetal fraction.
Subsequent management of cfDNA test failures and pregnancy outcomes
Further genetic counseling was offered to those women with cfDNA test failure, and IPD was suggested to high-risk patients. A flowchart of subsequent management and pregnancy outcome of cfDNA test failures at the first blood draw is shown in Figure 1.
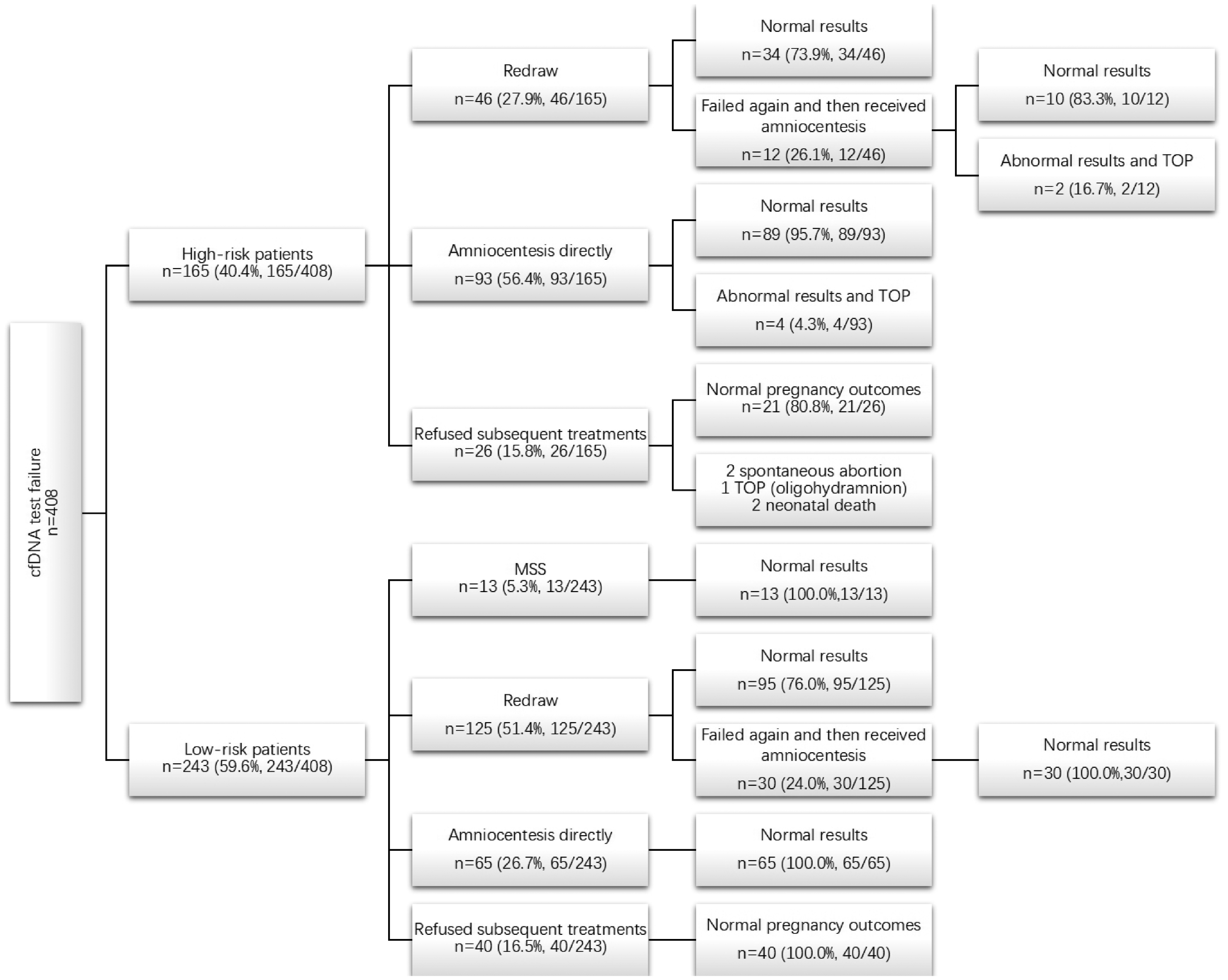
Flowchart of cases with a first result of cfDNA test failure.
After the post-test genetic counseling, 66 women refused any subsequent management. Of these, 26 belonged to the high-risk group and 40 to the low-risk group. Adverse pregnancy outcomes were seen in five cases, including two of spontaneous abortion, one of termination of pregnancy because of oligohydramnion, and two of neonatal death. These five women belonged to the high-risk group and refused any further genetic evaluation. The pregnancy outcomes of the other 61 cases were unaffected.
Of the remaining 342 cases who chose subsequent management, 13 low-risk women chose the second-trimester MSS. All 13 had low-risk MSS results and unaffected pregnancy outcomes.
Redraw for a second blood test was chosen by 171 (of 342) women, including 142 cases of low FF, 24 of data fluctuation, and 5 of high concentration of total cfDNA. Among these, 129 obtained an informative cfDNA test result (75.4%, 129/171), including 34 (26.4%, 34/129) cases of high risk and 95 (73.6%, 95/129) of low risk. The other 42 cases (24.6%, 42/171) had an uninformative result again, which accounted for 0.15% (42/27,363) of the total number of cfDNA test failures. These 42 women then chose amniocentesis, including 12 (28.6%, 12/44) cases of high risk and 30 (71.4%, 30/42) low risk.
The remaining 158 (of 342) women had an amniocentesis directly after the first sampling, including 93 high-risk and 65 low-risk cases. Adding the 42 cases with a second blood test failure, amniocentesis was offered to 200 women altogether, including 105 (52.5%, 105/200) high-risk and 95 (47.5%, 95/200) low-risk cases. Karyotyping was performed, and CMA was suggested to those with a fetal structural abnormality detected by ultrasound. Ultimately, abnormal genetic test results were found in six cases, all high-risk, and accounting for 5.7% (6/105) of the high-risk group. The abnormal results of these six cases are shown in Table 3.
Abnormal prenatal genetic diagnostic test results.
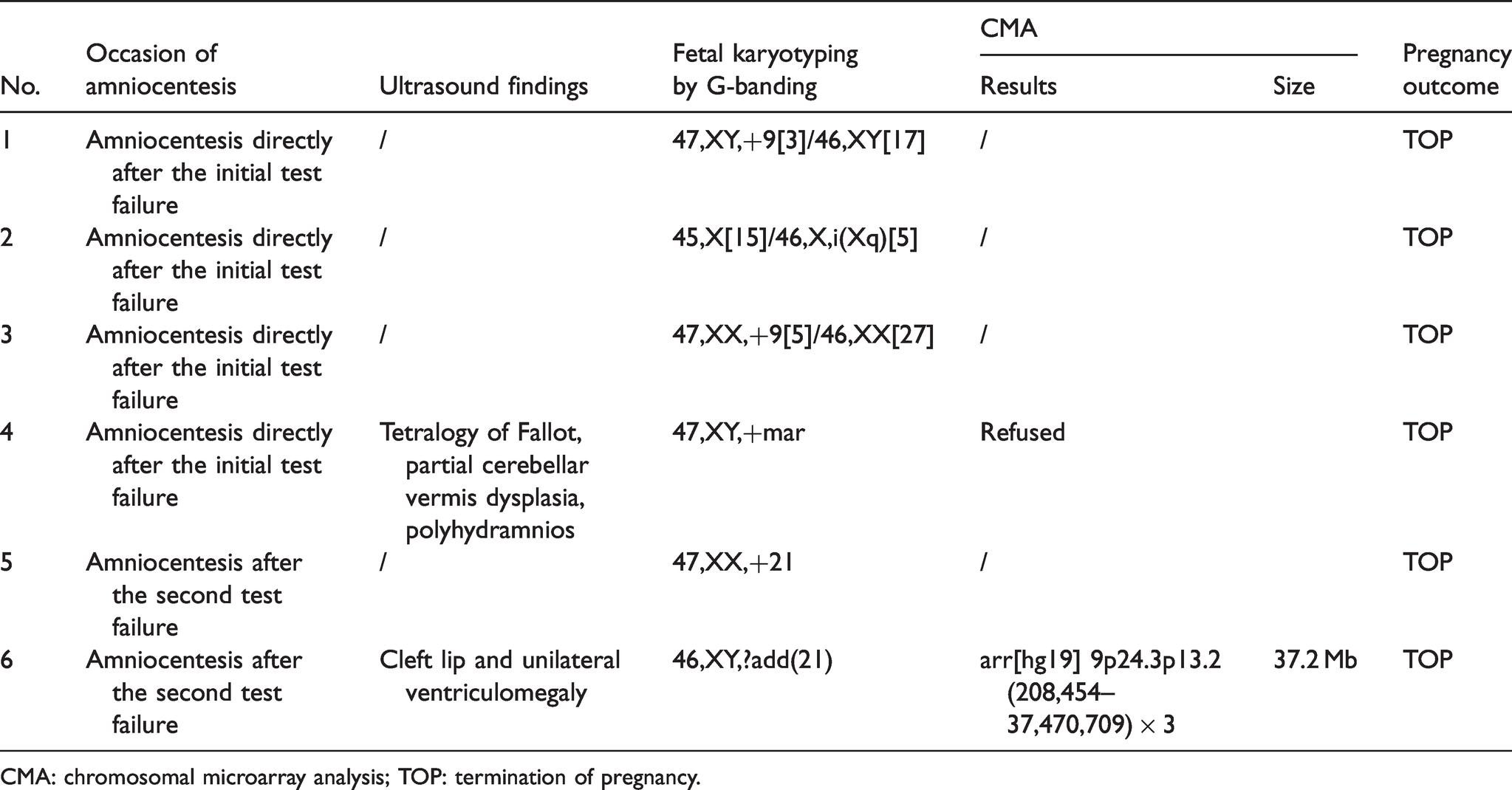
CMA: chromosomal microarray analysis; TOP: termination of pregnancy.
Discussion
To the best of our knowledge, this is the first retrospective study focused on consecutive cases with a cfDNA test failure result in a single tertiary center. This is also the first study to discuss the subsequent management of such cases.
Rate of cfDNA test failure
The rate of cfDNA test failure reported ranges from 1 to 8% depending on the assay technology. MPS-based methods were reported to have the lowest test failure rate (1.58%), followed by chromosome-specific sequencing (3.56%), and the single‐nucleotide-polymorphism-based method had the highest rate (6.39%). 9 In the current study, an MPS-based sequencing platform was used and the rate of cfDNA test failure was 1.49%, which is marginally lower than in the literature reported.9–11 This might be because the samples were drawn at later gestational weeks than in the previous studies, and thus with a higher FF.
Reasons for cfDNA test failure
Many factors, biological and technical, may be associated with cfDNA test failure. In this study, a failure is defined as an inconclusive result due to low FF or other laboratory assay failure, i.e. high concentration of total cfDNA or data fluctuation. In those cases with other assay failure, a repeat assay was routinely performed with the original sample, for confirmation.
The most common reason for test failure is low FF.3,9 There is no universal FF threshold that is applicable to all platforms and there have not been any differences demonstrated in cfDNA test performance by sequencing platform or method of FF calculation. Most cfDNA test providers have a common FF cut-off at 3–4% for a valid result. 12 In the position statement of the American College of Medical Genetics and Genomics (ACMG) in 2016, it was suggested that the lower limit of cfDNA for a reliable result was approximately 4%. 13 Some methodologies may be reliable with FF as low as 2%. 14 In this study, we chose to use the suggested cut-off of 4%.
Among 408 cases of cfDNA test failure after the first sampling, 78.4% were due to a low FF. This result corresponds with previous studies. The main contributors to low FF are small placental mass, high maternal weight, and impaired placentation. There are large interindividual variations in FF, with many biological factors influencing the maternal and placental cfDNA contributions. Any conditions that increase the maternal contribution and/or reduce the placental contribution may lower the FF.8,15–17 The proportion of women with FF < 4% increases with maternal weight. 15 In their recent review, Juul et al. found that the failure rate of the cfDNA test increased with increased maternal obesity. The test failure rate ranged from 0 to 4.2% for women with BMI 18.5–24.9, and from 5.4 to 70.1% for women with BMI > 40. 18 In our study cohort, the weight and BMI of women with a cfDNA test failure result were significantly higher than of those women with an informative result (p < 0.001 in both cases), and the gestational week was significantly earlier (p < 0.001), consistent with the previous literature.
We divided women with a cfDNA test failure result into high-risk and low-risk groups, according to their indications (age and MSS result). Chi-squared testing revealed that there was no statistical difference (p = 0.76) in the distribution of high-risk and low-risk patients among the three groups with different reasons for cfDNA test failure after the first sampling. This indicated that the failure was mainly technical, and not due to factors associated with the woman and her pregnancy.
Subsequent management and pregnancy outcomes
There is a lack of consensus among professional societies around the clinical management of patients with a cfDNA test failure result. In our study cohort, further genetic counseling was offered in all cases, and IPD was suggested to the high-risk patients. Women chose the subsequent management according to their own preferences.
Performance of a second blood test
Limited published data exist to inform a decision to obtain a repeat specimen versus pursue alternative testing.19,20 Guidelines from the position statement of the International Society of Prenatal Diagnosis recommend reappraisal based on clinical factors such as gestational age and ultrasound findings. 21 According to the statement of the ACMG, a repeat blood draw is not appropriate in the case of a cfDNA test failure due to low FF if maternal blood for the cfDNA test was drawn at an appropriate gestational age. 13 This rested on uncertainty over whether repeat collection would overcome a low FF. In their recent study, White et al. concluded that the probability of obtaining a result with repeat cfDNA test decreases with higher maternal weight and increases with the interval between draws. 22 In their cohort study, about 53% of patients overall received a result upon repeat test. In another two previous studies, approximately 50–60% of women with a failed cfDNA test result had success with a second blood draw.23,24 In our cohort, 171 women chose a second blood draw, and this yielded a result in 75.4%. The mean gestational age at the initial sampling and the second sampling was 15.6 ± 2.3 weeks and 18.7 ± 2.5 weeks, respectively; both higher than in the previous reports. This might explain the greater success rate of a second blood test in our study.
Of the 171 women who chose a second blood test, 42 (42/171, 24.6%) again failed to obtain an informative result and then underwent amniocentesis. In this series of patients, 12 were from the high-risk group and 30 from the low-risk group. Two cases from the high-risk group had abnormal prenatal genetic diagnosis results. The karyotype analysis showed that one case was trisomy 21 and the patient chose to terminate the pregnancy. Ultrasound of the other case found a cleft lip and unilateral ventriculomegaly of the fetus. CMA testing of the amniotic fluid showed a 37.2 Mb gain at the region of 9p24.3p13.2. The patient chose to terminate the pregnancy. Both women were AMA, accounting for 16.67% (2/12) of these high-risk patients.
In another study on prenatal reflex DNA screening, the cfDNA test failure rate for the first aliquot of plasma during initial blood draw was 8.7% (152/1756). 25 Interestingly, before a second blood draw, cfDNA testing was performed using the second aliquot of plasma and the success rate was 80.7% (121/150). This suggests that the failure could be technical and repeating the library preparation would reduce the failure rate; thus, using the second aliquot of plasma from the initial blood draw could be the next choice after cfDNA test failure.
Performance of alternative aneuploidy screening method
From the low-risk group, 13 women chose second-trimester MSS; they all obtained a low-risk MSS result and unaffected pregnancy outcome. MSS has been widely used for nearly 30 years. The screening methods and techniques have been proven to be reliable and the protocols are constantly improving. Moreover, a significant advantage over cfDNA testing is that MSS has almost no risk of failure. Although the sample size was small, the results suggest that low-risk patients who fail to obtain an informative cfDNA test result at first sampling could choose MSS as an alternative aneuploidy screening method.
Performance of IPD
In total, amniocentesis was offered to 200 patients, 52.5% of whom were high risk. Six of these high-risk patients (five cases of AMA and one case of high-risk MSS) ultimately received abnormal genetic test results. Further ultrasonic examination was performed in two cases and structural fetal anomalies were found (Table 3, cases 4 and 6).
Of the six cases, four came from the group who chose amniocentesis directly after an initial cfDNA test failure, giving a rate of abnormal genetic diagnosis of 2.53% (4/158), and accounting for 4.30% (4/93) of that series of high-risk patients. The other 2 cases were of the 42 who chose amniocentesis after a second cfDNA test failure, giving a rate of abnormal genetic diagnosis of 4.76% (2/42), and accounting for 16.67% (2/12) in that series of high-risk patients. The rate of abnormal genetic diagnosis was much higher among those patients with a second cfDNA test failure, although there was no statistical difference compared with the women who chose amniocentesis directly after the initial cfDNA test failure. It might be because of the relatively small sample size of these two series.
Three of the six cases with abnormal prenatal genetic diagnosis were associated with abnormality of chromosome 9 (two cases of mosaic trisomy 9 and one case of pathogenic copy number variant from chromosome 9), accounting for 50% of the genetic abnormalities in this study. This might reflect an association with smaller placentas which is likely to be responsible for the abnormal cfDNA levels and subsequent failure to obtain a result from cfDNA testing. 26 The other three cases were one of trisomy 21, one of mosaic of 45,X/46,X,i(Xq), and one of marker chromosome but further CMA was refused (Table 3, case 4).
Pregnancy outcome of those patients who refused subsequent management
Sixty-six women refused subsequent management after a first cfDNA test failure. Adverse pregnancy outcomes were seen in 5 of the 26 high-risk cases in this series (19.2%). All five patients refused further genetic evaluation. The pregnancy outcomes of the other 61 cases were unaffected. Thus, for patients with high-risk pregnancy, particularly those with a cfDNA test failure result, a confirmative prenatal test is recommended.
Conclusions
cfDNA test failure is intimately related to both biology and the technology used. Clinically, failure to achieve a result from a cfDNA test is associated with a higher risk of adverse pregnancy outcome and increased risk for aneuploidy; these cases cannot be considered high risk per se. In our study cohort, all the patients with an affected pregnancy outcome were high-risk patients. Such cases require careful counseling and further genetic diagnosis. The pregnancy outcomes of the low-risk patients were unaffected, which implies that a second sampling for cfDNA test or an alternative aneuploidy screening method might be considered.
In their recent review, Hui and Bianchi suggested that clinicians do not need to know the details behind the biological and technological reasons for cfDNA test failure, but they do need to have a “plan B” prepared for affected patients in their clinical practice. 27 Although waiting before redrawing a fresh sample will increase the probability of obtaining a result, the clinical utility of doing so must be considered. Redraws only resolve some of the cases and significantly increase turnaround time. IPD should also be considered in light of gestational age, patient preference, background aneuploidy, and results of detailed fetal ultrasound examination. We suggest that IPD be offered to the high-risk patients, especially those with a second cfDNA test failure.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
