Abstract
Long noncoding RNAs (lncRNAs) are crucial in porcine preimplantation embryonic development, yet their regulatory role during zygote genome activation (ZGA) is poorly understood. We analyzed transcriptome data from porcine fetal fibroblasts (PEF), induced pluripotent stem cells (iPS), and preimplantation embryos, identifying ZGA-specific lncRNAs like XLOC-040580, and further predicted its potentially interacting genes TPRA1 and BCL2L1 via co-expression network. XLOC-040580 was knocked down by siRNA microinjection and the expression of ZGA-related genes was detected by qRT-PCR. After microinjecting siRNA targeting TPRA1 and BCL2L1 at the one-cell stage, we counted the blastocyst development rate. The blastocyst development rate was consistent with the results from si-XLOC-040580 after si-TPRA1. Through dual-luciferase reporter assays, we found that XLOC-040580 was a downstream target of TPRA1. To further elucidate the mechanism of XLOC-040580, Single-cell mRNA sequencing after XLOC-040580 knockdown revealed its regulatory network involved in embryonic developmental defects. Transcriptome analysis revealed that XLOC-040580 was specifically expressed during zygote activation. Knockdown of XLOC-040580 decreased the blastocyst development rate and reduced both the total blastocyst cell number and TE cell number. TPRA1 and BCL2L1 were specifically co-expressed with XLOC-040580 during ZGA stage, and TPRA1 could interact with the promoter region of XLOC-040580 and regulate its expression. Knockdown of TPRA1 or XLOC-040580 blocked porcine embryonic development by affecting the expression of ZGA-related genes. We found and validated that lncRNA XLOC-040580 played a key role in the ZGA process, which was regulated by TPRA1. These results implied that the functional axis of TPRA1-XLOC-040580-downstream genes involved in ZGA-related functions also coordinated early embryonic development in porcine.

Keywords
Background
During early embryonic development, zygote genome activation (ZGA), cell polarization, asymmetric division, and the dynamic balance of pluripotency and differentiation-related genes are essential for embryonic development, any developmental defects will have a profound impact on individual embryos1,2. In the process of embryonic development, the first key process is ZGA, during which embryos undergo DNA replication and modification, transcription factor and enhancer activation, protein modification and response to external signals. The normal progress of ZGA is crucial to embryo cleavage. Any level of error will lead to the termination of embryonic development3,4. Previous studies have shown that mouse ZGA occurs during late one-cell to late two-cell embryonic stages 5 , and ZGAs in both pigs and humans occur from four-cell to eight-cell embryonic stage 6 .
Pigs share many similarities with humans in anatomical structure, physiological metabolism and pathogenesis of diseases, and are considered as ideal for development and disease models. Therefore, in-depth study of early porcine embryos helps to deepen the understanding of human embryonic development and related diseases7–10. LncRNAs have tissue-specific expression and spatiotemporal-specific expression, there are a large number of lncRNAs in early porcine embryos, and they are specifically expressed at different stages 11 . Several novel porcine lncRNAs have been identified in previous studies, and their unique expression patterns and structures make them play important roles in fetal development 12 , for instance, abnormal expression of Xist may be associated with midterm abortion in porcine nuclear transfer embryos 13 . Other lncRNAs were also identified in specific tissues, such as lung 14 , endometrium 15 and frontal cortex 16 .
LncRNAs are identified to play important roles during ZGA. Knockout of lncRNA-IL17D in mouse one-cell stage embryos leads to embryonic death, which can be rescued by IL17D protein at the four-cell stage in vitro 17 . LincGET related to embryonic cleavage can form RNA-protein complexes with hnRNP U, FUBP1 and ILF2 to regulate the transcription of ZGA 18 . LincGET can also form complexes with CARM1 to regulate the expression of Sox2, Nanog and Sox21 necessary for early embryonic development in mice 19 . In addition, lncRNA-37 was found to target TTLL12 and NDST1 to mediate ZGA in goat embryos 20 . Previous studies have shown that lncRNAs are involved in the regulation of pig ZGA and play an important role in early embryonic 21 . Here, we identified that XLOC-040580 was specifically expressed during porcine zygote activation, and confirmed that XLOC-040580 affects ZGA and further embryonic development. In addition, we also verified the effect of TPRA1 and BCL2L1 as co-expressed genes of XLOC-040580 on embryonic development. Our results may provide a theoretical basis for the regulation mechanism of porcine early embryonic development, and also provide a technical support for the improvement of the quality of porcine embryos produced in vitro.
Materials and methods
Porcine oocyte collection, parthenogenetic activation (PA), and embryos culture
Porcine ovaries were collected at the slaughterhouse and transferred to the laboratory within 3 h in sterile physiological saline at 37℃ with 100 IU/l penicillin and 100 g/ml streptomycin sulfate. The ovaries were rinsed with normal saline (38℃) three times. The cumulus-oocyte complexes (COCs) were extracted from normal follicles with a diameter of 2–8 mm on the surface of the ovaries by a syringe and placed in a 9-cm culture dish to select COCs with more than three layers of intact and dense cumulus cells. The selected oocytes were washed three times in IVM solution and cultured in a CO2 incubator, at 38.5℃ for 42–44 h. Mature oocytes were transferred to non-electrolyte solution containing 0.25 mM mannitol, 0.1 mM CaCl2, 0.1 mM MgSO4, 0.01% PVA (w/v), 0.5 mM HEPES (pH 7.0–7.2) and washed three times after removal of cumulus cells by 0.1% hyaluronidase. Then, the somatic hybridization instrument (Shimadzu SSH-2, Kyoto, Japan) (pH 7.0–7.2) was used to activate with two DC pulses of 1.56 kV/cm. The activated embryos were cultured in PZM-3 medium 22 at 38.5℃. All experiments involving animal embryos were approved and conducted according to the guidelines of the Institutional Animal Care and Use Committee of China Agricultural University (approval no. AW02113202-3-1).
In vitro GFP mRNA synthesis
Before microinjection, the expression vectors need to be linearized, then extracted with phenol/chloroform and precipitated with ethanol. The linearized DNA was transcribed in vitro according to the MESSAGE T7 mMACHINE® Kit (Ambion). The synthesized product was recovered with lithium chloride and dissolved in nuclease-free water.
siRNA microinjection
siRNAs were designed and synthesized by Suzhou Jima Pharmaceutical Co Ltd. Negative control siRNA without a specific target and siRNA of lncRNA XLOC-040580 are the following (Table S1). Use Eppendorf for siRNA microinjection. There is 3 μl RNA solution (50 nM siRNA, 30ng/ul GFP mRNA) in the injection needle 23 .
Plasmid electro transfection
siRNAs were transformed into shRNAs, and primers for PCR amplification were designed using porcine four-cell to eight-cell embryo cDNA as the template. The fragment was digested with Bgl II and Hind III and inserted into pSuper (Addgene) plasmid by T4 ligase Plasmids were extracted using EndoFree Plasmid Maxi Kit (QIAGEN, 12391). Then 4 μg of shRNAs plasmid were transfected to 5×104 pgEpiSCs or iPS using Lonza Amaxa Nucleofector-2b program A023 24 . After electric transfection, cells were taken back into a 37℃ incubator for further culture.
XLOC-040580 overexpression
For cloning of XLOC-040580, we designed primers for PCR (Table S1) amplification using porcine four-cell to eight-cell embryo cDNA as a template. The fragment was digested with Hind III and Pac I, and inserted into the pCAG plasmid by T4 ligase. And pCAG-XLOC-040580 was in vitro transcribed with 4PB plasmid into linearized DNA according to the message T7 Kit (Ambion). Overexpression of XLOC-040580 was finally achieved by microinjection into embryos.
Immunocytochemistry
For immunocytochemical analysis, embryos were fixed with 4% paraformaldehyde (PFA) in DPBS for 20 min at room temperature. Fixed embryos were washed three times with DPBS, incubated in 0.2% Triton X-100 buffer for 15 min, and washed three times with DPBS. After blocking in 2% BSA blocking buffer for 1 h, embryos were incubated at 4°C overnight in 1% BSA buffer containing primary antibodies. The following primary antibodies were used: anti-OCT4 (Santa Cruz), and anti-CDX2 (Cell Signaling Technology). The embryos were washed in DPBS and incubated for 1 h with secondary antibody. For nuclear staining, the cells were incubated for 2 min with DAPI (Life Technologies). The images were captured using Nikon microscope.
RNA purification and RT-PCR
Total cellular RNA of 100 embryos was extracted using RNeasy Mini Kit (QIAGEN). Reverse transcription was performed using an oligo-dT primer and M-MLV Reverse Transcriptase (Promega). Quantitative RT-PCR (q RT-PCR) analyses were performed using the LightCycler 480 SYBR Green I Master Kit (Roche) and detected with LightCycler 480II (Roche). The data were analyzed using the comparative CT (2-ΔΔCT) method. The ΔCT was calculated using Actin as an internal control. All experiments were performed with more than three biological replicates.
Identification of lncRNAs by RNA-seq
LncRNAs were identified by a series of analysis procedures using porcine pre-implanted embryonic RNA-seq data. First, we performed cufflink to construct and exclude protein-coding transcripts, and subsequently excluded the single exon, short exonic length (<200 nt) and short open reading frame (<300 nt) transcripts. Second, to distinguish the coding and non-coding sequences we performed both the Coding Potential Calculator (CPC) and Coding-Potential Assessment Tool (CPAT) to calculate the coding potential, then we also used PhyloCSF to analyze the conserved regions of the multi-species genome sequence alignment of all candidate transcripts. Finally, the remained transcripts were searched against the Pfam database of hidden Markov models (HMMs) to filter the sequences that contained protein domains. Thus, we obtained the lncRNA candidates that affect ZGA in porcine pre-implanted embryos.
Single-cell RNA-seq and reads mapping
The si-RNA(including si-XLOC-040580, si-TPRA1, si-BCL2L1 and negative control) were microinjected into embryos at one-cell stage, then the eight-cell stage embryos were selected by mouth-suction tube under the stereomicroscope, digested with 0.5% streptomycin, and washed three times in 1% PBS, then stored in cold scRNA-seq cell lysate. We used a modified STRT-seq protocol to construct the libraries 25 . Single-cell cDNA was obtained by terminal repair, reverse transcription, and PCR amplification. The unit bar code obtained by Single-cell RNA-seq matches the end sequence, and read 1 should be attached when reading the UMI sequence in reads 2. The transcript abundance of each gene was estimated using HTSeq (v0.6.1), and the reads with duplicate UMIs of each gene were excluded 26 . Finally, the expression levels of genes were normalized to the number of TPMs (transcripts per million reads).
Alignment of RNA-seq data and differential expression analysis
Raw reads from scRNA-seq were split by 8 bp cell barcodes located on read 2 allowing two mismatches. In addition, the 8 bp unique molecular identifiers (UMIs) located on read 2 were switched to the identifier line of paired read 1, then read 1 was processed to remove the template switch oligo (TSO) primer, low-quality bases, and polyA sequence. The trimmed reads were aligned against the Ensembl Sus scrofa reference genome (Sscrofa11.1) using STAR software with default parameters. The gene expression levels were estimated by counting the UMIs assigned to the genes using feature Counts, yielding an expression matrix consisting of UMI counts for each cell and gene. Differential expression genes were identified by DESeq2 package, and functional enrichment for Gene Ontology (GO) and KEGG were performed with GOstats package 27 . LncRNA regulatory network was visualized by Cytoscape.
LncRNA interacting gene prediction and dual-luciferase reporter assay
LncRNA interacting genes were identified by co-expression modules in a previous study 11 , then we use LncTar 28 to verify the potential of lncRNA-mRNA interaction, which included promoter, CDS and 5’-UTR region of each gene, and potential genes were filtered with the minimum normalized binding free energy (ndG). The promoter of XLOC-040580 (chr12:5,597,712–5,603,712) is fused to a firefly luciferase reporter and is co-transfected into 293T cells with an internal control vector (PRL-TK) and overexpression plasmid (pMX-TPRA1) to express renilla luciferase (Table S1), both the firefly and renilla luciferases are measured using a dual-luciferase reporter assay system by Dual-Glo Luciferase Assay System (E2920, Promega). The relative luciferase activity is presented as a ratio of firefly luciferase intensity to renilla luciferase intensity.
Statistical analysis
Each experiment was repeated three times, and the results are presented as the mean standard error. Data were analyzed by one-way analysis of variance (ANOVA) by LSD tests and χ2 using SPSS software. P < 0.05 was considered to be statistically significant.
Results
LncRNA XLOC-040580 is specifically expressed during porcine ZGA
We analyzed the transcriptome data of porcine oocytes, one-cell, two-cell, four-cell, eight-cell stage embryos, morula, blastocyst intracellular cell mass (ICM), trophoblast cells, iPS, and PEF cells 11 , and organized the results into 24 modules according to the expression levels of lncRNAs in embryos and cells. Nine of those modules were associated with early embryonic development in porcine, including module 1, 3, 5, 6, 8, 11, 17, 22, 24 (Fig. 1a). The lncRNAs and protein-coding genes specifically expressed during ZGA, which occurs during the stage from four-cell to eight-cell embryos in porcine, were mainly enriched in module 17. The heatmap of lncRNAs expression profile at different stages in module 17 indicated that lncRNA XLOC-040580 (chr12:5,603,712–5,608,560) is the most significantly differential expressed lncRNA from four-cell to eight-cell stage (Fig. 1b). We next confirmed that XLOC-040580 had the highest expression in eight-cell stage by RNA-seq data (Fig. 1c). In addition, expression levels of XLOC-040580 in ZGA were further verified by qRT-PCR using porcine iPS, PEF, oocyte, and PA embryos at different stages (Fig. 1d). These results suggested that XLOC-040580 may play an important modulator during porcine zygotic activation progress.
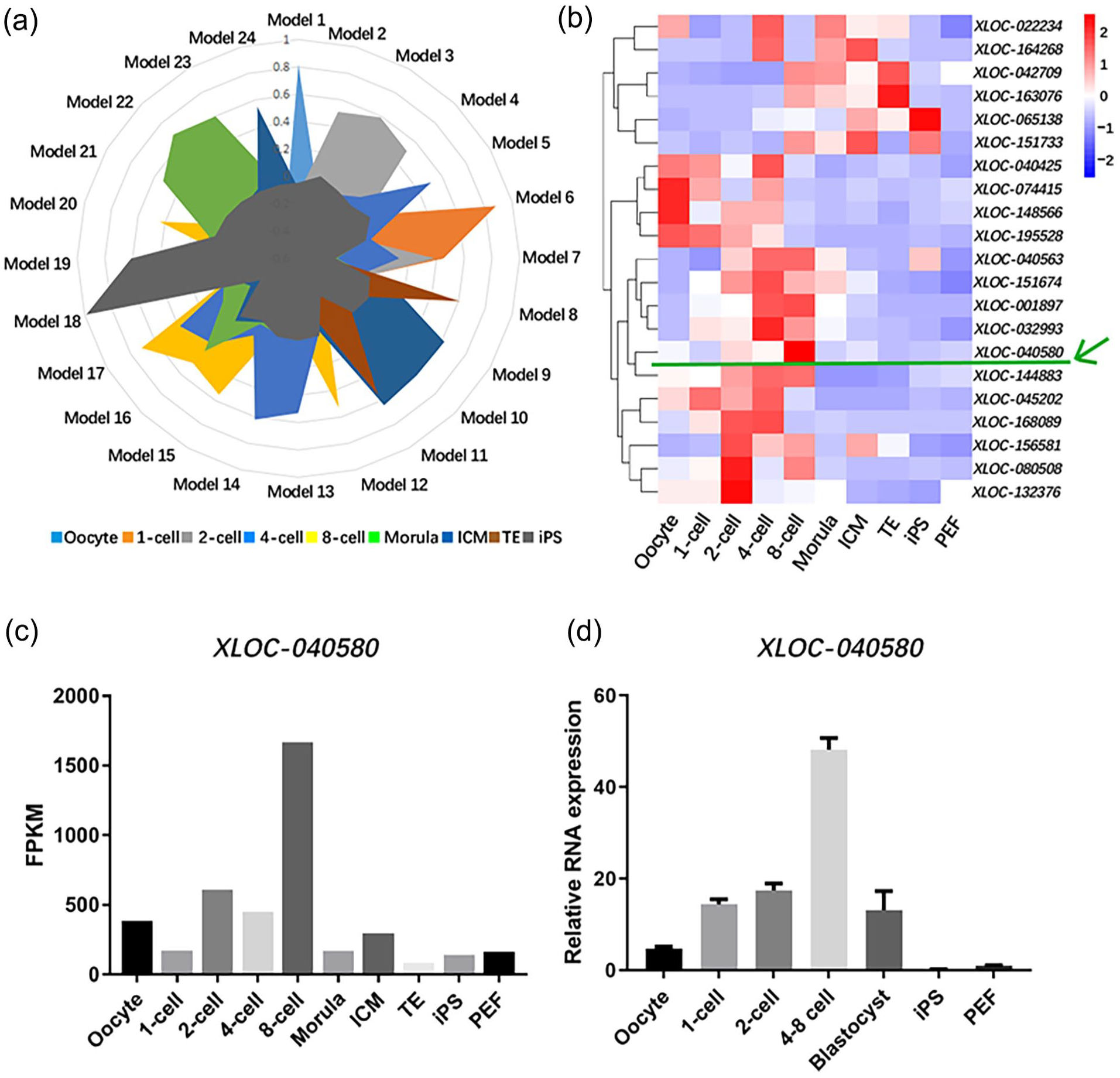
Identification of the specifically expressed lncRNAs during porcine ZGA. (a) Function module diagram of lncRNAs. Different colors represent functional blocks in different periods, the closer to the periphery from the center, the higher lncRNAs expression. (b) Heat map of lncRNA differential expressions in module 17. The green arrow indicates the candidate RNA XLOC-040580. (c) Expression of XLOC-040580 in embryos at different stages or in different cell types. (d) qRT-PCR analysis of XLOC-040580 differential expression in embryos at different stages or in different cell types.
XLOC-040580 knockdown influences embryonic development
We first designed siRNA and evaluated the knockdown efficiency of XLOC-040580 on iPS cells, the results showed that all three targets of si-XLOC-040580 significantly decreased their expression (Fig. 2a). Then, porcine one-cell embryos were injected with the three lncRNA-siRNA mixes at a concentration of 25 μM to investigate the effectiveness of knockdown expression of XLOC-040580 on embryonic ZGA. The control group injected with the control-siRNA that had no knockdown effect on any gene. To verify whether the injection of siRNAs was successful, we mixed GFP mRNA into siRNA solution, and only GFP-positive embryos were selected for subsequent experiments (Fig. 2b). qRT-PCR was used to detect the efficiency of siRNAs at the four-cell to eight-cell stage, and the results showed that the knockdown efficiency of XLOC-040580 was about 70% (Fig. 2c). To investigate whether XLOC-040580 played a regulatory role during embryonic development, the blastocyst formation ratio was calculated between the knockdown and control group. We found that there was no significant difference in four-cell to eight-cell embryo formation rate between the si-XLOC-040580 group and si-control group (P = 0.0636), while the blastocyst formation rate of the si-XLOC-040580 group was significantly lower than that of the si-control group (P = 0.0008) (Table 1, Fig. 2d).

Knockdown of XLOC-040580 resulted in reduced blastocyst development ratio. (a) qRT-PCR analysis of the knockdown efficiency of XLOC-040580 by siRNAs. *P < 0.5. (b) Microinjection system: one-cell stage embryos were injected with lncRNA-siRNAs and GFP mRNA, and the GFP expression after 24h indicates successful injection. Scale bar = 200 μm. (c) qRT-PCR analysis of the knockdown XLOC-040580, the efficiency of siRNAs was detected at the stage of four to eight cells. *P < 0.5. (d) Representative photos of wild type (WT) black group, si-control and si-XLOC-040580 embryos developed to blastocyst stage. Scale bar = 200 μm.
Development of porcine embryos after injection with XLOC-040580 siRNA in the one-cell stage embryos.

Values in the same column with different letters (a, b) differ significantly (P < 0.05).
XLOC-040580 knockdown reduces the number of TE cells
Subsequently, to verify the influence of XLOC-040580 on porcine ZGA, we first knock down XLOC-040580 by one-cell embryonic injection and interestingly we found that the number of trophectoderm (CDX2 positive) cells in E6.5 blastocysts, as well as the total number of blastocysts in the si-XLOC-040580 group, was significantly reduced in the XLOC-040580 knockdown groups (P < 0.0001 and P = 0.0245 respectively) (Fig. 3a, b). While there was no significant difference in embryonic development between si-XLOC-040580 and control groups after XLOC-040580 overexpression (Fig. S1), this may be due to the high expression level of zygotic genome during ZGA29–31 or the diverse regulation mechanisms of lncRNAs 32 . The results indicated that XLOC-040580 affects normal embryonic development by interfering with TE cells. In addition, we found that the expression levels of ZGA-related genes such as eIF1a, SMG, ACADL and HSP70.2 were downregulated after knocking down XLOC-040580 (Fig. 3c), which suggests that XLOC-040580 affects embryonic development by regulating ZGA-related genes.

Knockdown of XLOC-040580 affects ZGA. (a) Immunofluorescence histochemical staining of CDX2 (indicated the TE cells) and DAPI (indicates the cell nucleus) in E6.5 blastocysts. Scale bar, 20 μm. (b) Effect of XLOC-040580 knockdown on the cell number of TE and ICM in the treated blastocyst at 6.5d. *P < 0.5, ****P < 0.001. (c) qRT-PCR analysis of the expression levels of ZGA related genes after XLOC-040580 knockdown at the eight cell stage, *P < 0.5, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Knockdown of TPRA1 or BCL2L decreases early embryonic development rate
To clarify the regulation mechanism of XLOC-040580 on porcine ZGA, lncRNA interacting genes were identified by co-expression modules. Co-expression results implied that several candidate genes might interact with XLOC-040580 during the ZGA, such as TPRA1(Transmembrane protein adipocyte associated 1), BCL2L1(B cell leukemia/lymphoma 2 like 1), MED27(Mediator complex subunit 27), BABAM1(BRISC and BRCA1 A complex member 1), VPS72(Vacuolar protein sorting 72 homolog) and UBE2A (Ubiquitin conjugating enzyme E2 A). It was previously reported that TPRA1 was closely related to cell growth and embryonic differentiation pathways, and BCL2L1 was involved in apoptosis regulation33,34. Therefore, we selected XLOC-040580 co-expressed ZGA-related genes TPRA1 and BCL2L1 for follow-up experiments (Fig. 4a). LncTar 28 was performed on those candidate genes to verify the potential of lncRNA-mRNA interaction, which included promoter, CDS and 5’-UTR region of each gene, and potential targets were filtered with the minimum normalized binding free energy (ndG) (Table S2). Subsequently, we validated the interactions between lncRNA XLOC-040580 and co-expressed genes by bioinformatic analysis and dual-luciferase reporting system, the results displayed that there is a binding region in the upstream of XLOC-040580 and implied that interaction may arise between TPRA1 and XLOC-040580 (Fig. 4b). While there is no direct binding signal between BCL2L1 and XLOC-040580 in region of upstream and gene body by LncTar, which suggested that BCL2L1 may be an indirect regulatory gene. To further verify the effects on early embryonic development of potential interactional counterpart of XLOC-040580, siRNAs of TPRA1 and BCL2L1 were designed. The knockdown efficiencies were validated by transfection using porcine pgEpiSCs 24 , and then the optimal siRNAs were selected for subsequent experiments (Figs. 4c and S2A). We continued to knock down TPRA1 and BCL2L1 at the one-cell stage of porcine embryos using the optimal siRNAs and the results showed that the four-cell to eight-cell embryonic development rate and blastocyst formation rate in si-TPRA1 group and si-BCL2L1 group were significantly lower than WT black group and si-control group (Tables 2 and S3, Figs. 4d and S2B), which suggested that the silencing of ZGA-related genes could inhibit early embryonic development similar to the function of XLOC-040580. Notably, the in silico analysis and dual-luciferase reporter assay exhibiting the interaction between TPRA1 and XLOC-040580, which implied that XLOC-040580 was a key target of TPRA1 that regulated the co-expression of XLOC-040580 and finally involved in the early embryonic development.

Predicting the ZGA-related gene of XLOC-040580 and verifying their effects on embryonic development. (a) The differential expression of ZGA-related gene of XLOC-040580 in porcine early embryos was shown by thermal map. (b) Analysis of the target relationship between TPRA1 and XLOC-040580 by dual luciferase assay, which are presented as the relative ratio of firefly luciferase activity to renilla luciferase activity. ** P < 0.01. (c) qRT-PCR analysis of the knockdown efficiency of TPRA1 siRNAs. ****P < 0.001. (d) Representative photographs of the blastocysts in WT black group, si-control and si-TPRA1 groups. Scale bar = 200 μm.
Development of porcine embryos after injection with TPRA1 siRNA and BCL2L1 siRNA in the one-cell stage embryos.

Values in the same column with different letters (a, b) differ significantly (P < 0.05).
Knockdown of TPRA1 or BCL2L1 disrupts early embryonic development related function
To further explore the potential causation of embryonic development failure after knocking down the TPRA1 and BCL2L1, we used single cell RNA-seq to analyze differentially expressed profiles between the control porcine embryos and those with knock down TPRA1 and BCL2L1. Further analysis showed that 100 genes were downregulated and 233 genes were upregulated after knocking down TPRA1. After knocking down BCL2L1, 153 genes were downregulated and 179 genes were upregulated (|log2(fold-change)| ≥ 1; adjusted P-value ≤0.05) (Figs. 5a and S2C). We found that TPRA1 related gene AGO2 was involved in oocyte development by GO enrichment analysis, BCL2L1 related gene SNRNP and PRDX5 were involved in oocyte meiosis and maturation (|log2(fold-change)| ≥ 1) (Figs. 5b and S2D). In order to understand the key pathways that TPRA1 and BCL2L1 can affect the development of porcine embryos, we performed KEGG enrichment analysis, and both ZGA-related genes were involved in MAPK signaling pathway (Figs. 5c and S2E). In addition, TPRA1 is also involved in progesterone-mediated oocyte maturation, oocyte meiosis and other signaling pathways related to early embryonic development (adjusted P-value <0.05) (Fig. 5c). These results coincide with the initial conjecture that lncRNA is essential in the zygote activation stage.
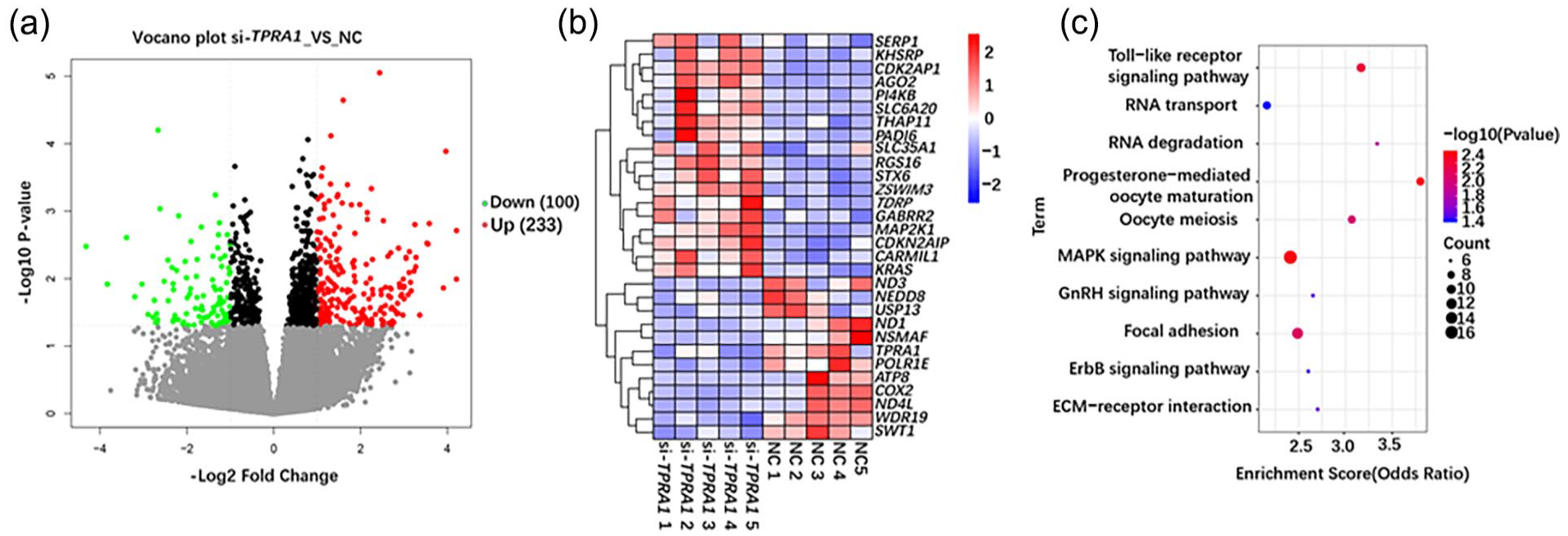
Knockdown of TPRA1 disrupts early embryonic development. (a) Volcano map showed the differential expression of related genes in porcine embryos after TPRA1 knockdown. (| log2(fold-change)| ≥ 1; adjusted P-value ≤0.05) (b) Heat maps showing differential gene expression in TPRA1 knockdown embryos. The scale bar of heatmap represents Z-score normalized expression of genes. (| log2(fold-change)| ≥ 1) (c) KEGG pathway is enriched in porcine embryos knocking down TPRA1. (adjusted P-value <0.05).
TPRA1-XLOC-040580 downstream gene regulatory axis in embryonic development
Since knocking down TPRA1 and BCL2L1 affects embryonic development, and TPRA1 simultaneously regulates the lncRNA XLOC-040580, we performed single cell RNA-seq for eight-cell stage of knockdown of XLOC-040580 to further investigate how the TPRA1-XLOC-040580-downstream gene regulatory axis influences embryonic development. Transcriptome analysis indicated that high similarity between samples from the si-XLOC-040580, NC, and WT groups (Fig. 6a). Compared with the WT group, the si-XLOC-040580 group exhibited 421 differentially expressed genes, with 345 upregulated and 76 downregulated (Fig. 6b–c). GO and KEGG enrichment analysis indicated that, after XLOC-040580 knockdown, down regulated genes were significantly enriched in the cell cycle pathway (P = 0.048), and GO functions related to transcription regulation, such as RNA binding (P = 0.001), was also significantly enriched (Fig. 6d). Up regulated genes were significantly enriched in metabolic pathways (P = 1.29e–05) according to KEGG, and GO analysis showed enrichment in ATP synthesis coupled with electron transport (P = 0.037) (Fig. 6e). Finally, a regulatory axis was constructed with TPRA1-XLOC-040580-downstream genes (Fig. 6f), which indicated that XLOC-040580 is involved in regulating several core functional networks, including endonuclease activity, cell cycle, cellular ketone metabolic process, ribonuclease MRP complex, ribonucleoprotein complex binding, and the acetyl-CoA biosynthetic process from pyruvate. These results indicated that XLOC-040580 plays important roles during zygotic activation and early embryonic development by regulating cell cycle, transcriptional expression, and metabolic alteration.
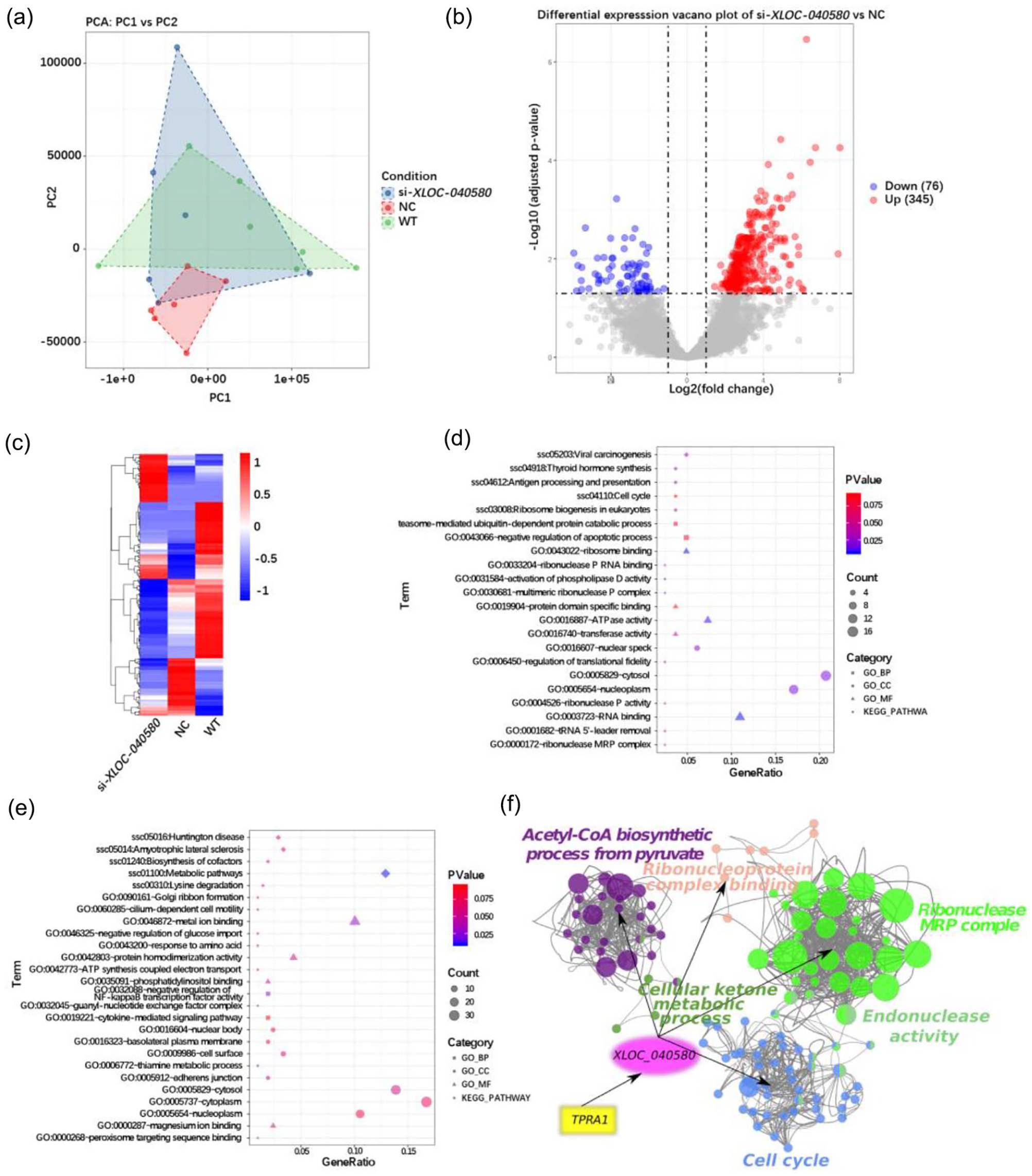
Investigate XLOC-040580 Downstream Gene Regulatory Network in Embryonic Development. (a) PCA plot of si-XLOC-040580, NC and WT group. (b) Differentially expressed genes display in volcano plot, x-axis indicate the Log2 fold change, Y-axis indicate the -Log10 adjusted P-value, red point is up regulated genes and blue point is down regulated genes, NS indicate the not significant genes which in gray. (c) Heatmap of total differentially expressed genes between si-XLOC-040580 and WT. The scale bar of heatmap represents Z-score normalized expression of genes. (d) The GO and KEGG enrichment of down regulated genes between si-XLOC-040580 and WT. (e) The GO and KEGG enrichment of up regulated genes between si-XLOC-040580 and WT. Count indicate the gene count in each term, category of GO function and KEGG displayed in different shape. (f) Functional network of TPRA1-XLOC-040580-Downstream genes. Color indicate the cluster of function and the size indicate the gene count of each function term.
Discussion
Zygote genome activation is essential for early embryonic development with a series of physiological and biochemical reactions, including transcription factor activation, DNA methylation, protein modification, RNA binding, metabolic changes, and maternal factor degradation35,36. In this study, we identified a novel lncRNA XLOC-040580 specifically expressed at ZGA stage based on the transcriptome analysis of early porcine embryos at different stages. We found that the rate of blastocyst formation decreased and the number of TE cells in blastocysts was significantly reduced after knocking down XLOC-040580, which suggested that lncRNA XLOC-040580 is indispensable in ZGA process. In addition, we found that the two co-expressed genes of XLOC-040580 were also involved in the regulation, especially, XLOC-040580 may serve as an intermediary between TPRA1 and the subsequent functional network during early embryonic development in porcine. These results provide a valuable reference for further study of porcine ZGA-related lncRNAs.
At the early stage of embryonic development before ZGA, the transcription of embryonic genome is completely regulated by maternal mRNAs and proteins and the maternal materials are gradually degraded during ZGA37,38. There is no significant difference in the gene expression between in vitro fertilization (IVF) and PA embryos during ZGA and their early development 39 . More studies indicated that KDM7A 40 , PDHA1 41 , KDM5B, KDM5C 42 , and other ZGA-related transcription factors also share similar expression patterns in IVF and PA embryos. In this study, the expression pattern of XLOC-040580 in natural preimplantation embryos is similar to that in PA embryos, which was confirmed by qRT-PCR (Fig. 1c, d), so we used PA embryos in the subsequent experiments.
In porcine, RNA-sequencing suggests that lncRNAs play important roles in tissue and embryo development43,44, while few lncRNA functions have been identified in specific developmental events. In this study, we found the novel lncRNA XLOC-040580, which specifically expressed in porcine four-cell-stage to eight-cell-stage embryos. Importantly, we verified that XLOC-040580 modulates porcine zygote activation progress by the assays such as siRNA, over-expression, and single cell RNA-seq etc. For example, XLOC-040580 knockdown reduced the expression levels of ZGA-related genes including eIF1a 45 , Smg, 46 ACADL 47 and Hsp70 48 , which were considered to be the marker genes for ZGA (Fig. 3c). Interestingly, XLOC-040580 knockdown decreased the rate of blastocyst formation and the number of TE cells suggesting that ICM and TE programs first started in ZGA consisting with the recent study about transcription factors in ZGA 49 .
In order to better explore the regulatory network of lncRNA XLOC-040580, we first focused on 2 co-expressed genes of XLOC-040580, namely TPRA1 and BCL2L1. We found that the expression abundance of TPRA1 and BCL2L1 increased from four-cell stage to eight-cell stage, and peaked at eight-cell stage, which were consistent with ZGA expression pattern. lncRNA XLOC-040580 is directly targeted by TPRA1, which identified by dual-luciferase reporter assay (Fig. 4b). It has been reported that TPRA1 mediates intracellular cAMP level, mainly affects cell growth and differentiation by regulating Hedgehog signaling pathway, and has a positive regulatory effect on embryonic formation and tissue homeostasis 33 . In addition, BCL2 protein family is a key regulator of apoptosis and participates in a variety of signaling pathways related to cell development50,51. To clarify the mechanism of TPRA1 and BCL2L1 regulations in early embryonic development, we performed single-cell transcriptome sequencing for the embryos with knockdown of the two genes during ZGA. The analysis results indicated that TPRA1 was related to oocyte gene inhibitor AGO252,53, male reproductive factor TDRP 54 and WDR19 55 . BCL2L1 is related to the two-cell cleavage and pretreatment mRNA factor SNRNP 56 , the PRDXs that affect mitochondrial activity throughout embryonic development 57 , and the apoptosis and senescence-related gene GADD45B 58 . We further drew the regulation network diagram of XLOC-040580 and TPRA1. Among them, TPRA1 is associated with many genes in the G protein-coupled receptor (GPR) family, and GPR107 is the key gene for embryonic lethality 59 (Fig. 5d). KEGG enrichment analysis indicated that TPRA1 and BCL2L1 jointly participate in MAPK signaling pathway, which plays a key role in spindle connection, microtubule organization and asymmetric division during oocyte maturation 60 , and is closely related to many intracellular events such as cell proliferation, cell differentiation and apoptosis. In addition, TPRA1 is also involved in signaling pathways such as oocyte maturation, gonadotropin-releasing hormone signal and oocyte meiosis (Fig. 5c). In summary, the results showed that lncRNA XLOC-040580 played a key role in pig ZGA process, which was regulated by TPRA1.
The mechanisms that XLOC-040580 regulates embryonic development based on the function of TPRA1 might be an initiation. To address the role of XLOC-040580, we performed single-cell RNA-seq on embryonic samples injected with si-XLOC-040580 to identify its downstream target genes. By comparing the si-XLOC-040580 group with the WT group, we identified 421 differentially expressed genes. To further investigate whether these genes could be potential targets of XLOC-040580, we used LncTar predictions, including GDNF, ADGRG7, VXN, TMC3, and SNT, which highlighted genes involved in cell fate determination and differentiation. However, none of those potential targets showed positive results in the dual-luciferase reporter assay. LncRNAs can function as cis-acting elements or trans-acting factors, and their regulatory actions are not always direct targeting relationship 61 . Enrichment analysis of differentially expressed genes allowed us to explore the functional regulatory networks in which XLOC-040580 may participate, highlighting several key clusters and further revealing lncRNAs’ regulatory mechanisms in embryonic development. One of the mechanism appears to be the regulation of epigenetics 62 , this cluster focused on the Acetyl-CoA biosynthetic process. Studies show that trophoblast stem cells (TSCs) maintain basal glycolysis, becoming sensitive to glycolytic deficiency, while Acetyl-CoA synthesis influences histone acetylation, thereby reducing TSC differentiation potential 63 . Another key mechanism lncRNA-mediated regulation of gene transcription, processing, and degradation, which is essential for the dynamic spatiotemporal gene expression during embryonic development 64 . The clusters centered on ribonucleoprotein complex binding, ribonuclease MRP complex, and endonuclease activity, which highlight the role of lncRNAs in interacting with proteins to form ribonucleoprotein complexes, altering cellular homeostasis through gene regulation 65 . In addition, lncRNAs are involved in regulation by controlling the expression of cell cycle regulators or acting as mediators of cell cycle-regulated gene expression66,67. In this study, genes related to the cell cycle pathway were significantly enriched among the downregulated genes in the si-XLOC-040580 group, potentially leading to impeded embryonic development. Therefore, a functional insights into lncRNA XLOC-040580 its downstream genes provide a valuable understanding of how lncRNA regulate porcine ZGA and embryonic development.
Conclusions
A novel lncRNA XLOC-040580 targeted by TPRA1 is identified to regulate ZGA process during early porcine embryo development, Our findings also emphasize the complexity of lncRNA-mediated regulatory networks and their roles in porcine ZGA and early developmental processes.
Supplemental Material
sj-docx-1-cll-10.1177_09636897251332527 – Supplemental material for LncRNA XLOC-040580 targeted by TPRA1 coordinate zygotic genome activation during porcine embryonic development
Supplemental material, sj-docx-1-cll-10.1177_09636897251332527 for LncRNA XLOC-040580 targeted by TPRA1 coordinate zygotic genome activation during porcine embryonic development by Mengxin Liu, Enhong Li, Haiyuan Mu, Zimo Zhao, Xinze Chen, Jie Gao, Dengfeng Gao, Zhiyu Liu, Jianyong Han, Liang Zhong and Suying Cao in Cell Transplantation
Footnotes
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
