Abstract
Accumulating evidence indicates that m6A methyltransferase 3 (METTL3) plays a pivotal role in different malignancies including melanoma. However, the function and underlying mechanisms by which METTL3 contributes to the tumorigenesis of melanoma remain undocumented. The association of METTL3 and long noncoding RNA (lncRNA) small nucleolar RNA host gene 3 (SNHG3) with clinicopathological characteristics and prognosis in patients with melanoma was analyzed by real-time quantitative polymerase chain reaction (RT-qPCR), Western blotting, and The Cancer Genome Atlas data sets. The role of METTL3 in melanoma cells was assessed by in vitro and in vivo experiments. The m6A dot blot, methylated RNA immunoprecipitation (MeRIP), and RT-qPCR were used to verify METTL3-mediated m6A modification of lncRNA SNHG3. The effect of METTL3 on lncRNA SNHG3 was determined by luciferase gene reporter assay, RT-qPCR, and Western blotting. We found that METTL3 was upregulated in melanoma tissue samples and associated with poor survival in patients with melanoma. Knockdown of METTL3 suppressed the growth and invasion of melanoma cells in vitro and in vivo, whereas restored expression of METTL3 promoted these effects. Mechanistic investigations showed that knockdown of METTL3 reduced SNHG3 m6A levels and its messenger ribonucleic acid (mRNA) expression levels. SNHG3 could act as a sponge of microRNA (miR)-330-5p to upregulate the expression of CCHC-type zinc finger nucleic acid binding protein (CNBP). SNHG3 overexpression reversed METTL3-knockdown-caused antitumor effects, miR-330-5p upregulation and CNBP downregulation. SNHG3 had a positive correlation with METTL3 expression but a negative correlation with miR-330-5p expression in melanoma tissue samples. In conclusion, our findings demonstrated that METTL3-mediated m6A modification of lncRNA SNHG3 promoted the growth and invasion of melanoma cells by regulating the miR-330-5p/CNBP axis.
Keywords
Introduction
Malignant melanoma is one of the most aggressive malignancies, and its incidence and mortality rates have increased in recent decades 1 . Once melanoma metastasizes to distant organs, the patients will have a very poor prognosis 2 . Although immunotherapy has been introduced into the treatment of the advanced cases, the 5-year survival rate remains low 3 . Therefore, understanding the molecular biology of melanoma progression is necessary for the early detection and treatment of melanoma.
N6-methyladenosine (m6A) is one of the most common modifications of mRNAs. The methylation status of mRNAs can be dynamically modified by m6A methyltransferase and demethylases 4 . Increasing evidence indicates that deregulated m6A modification is involved in melanoma progression5–7. METTL14-mediated m6A modification of RUNX family transcription factor 2 (RUNX2) accelerates the invasion of choroidal melanoma 8 . The m6A demethylase FTO promotes melanoma tumorigenicity and anti-PD-1 resistance 9 , and m6A reader YTHDF3 enhances the growth of ocular melanoma by promoting catenin beta 1 (CTNNB1) translation 10 .
METTL3, as an m6A methyltransferase 3, is involved in carcinogenesis including melanoma 11 . METTL3 stimulates the proliferation of primary acral melanoma by m6A methylation of thioredoxin domain containing protein 5 (TXNDC5) 12 and contributes to PLX4032 resistance in melanoma by promoting epidermal growth factor receptor (EGFR) translation 13 . In addition, METTL3-mediated m6A modification of uridine-cytidine kinase 2 (UCK2) drives melanoma metastasis through Wnt/β-catenin signaling 14 , whereas depletion of METTL3 facilitates the response to anti-PD-1 treatment in melanoma 15 . These findings indicate that METTL3 is a potential therapeutic target for melanoma.
It has been shown that noncoding RNAs act critical roles in biological processes, including cancer16,17. METTL3-mediated m6A modification of noncoding RNAs in melanoma has not been reported. In this study, we found that upregulation of METTL3 was associated with poor survival in patients with melanoma, whereas METTL3-mediated m6A modification of long noncoding RNA (lncRNA) SNHG3 promoted the growth and invasion of melanoma cells by regulating the microRNA (miR)-330-5p/CCHC-type zinc finger nucleic acid binding protein (CNBP) axis. Our findings provide evidence for METTL3 as a promising therapeutic target for melanoma.
Materials and Methods
Clinical Samples
A total of 474 cases of patients with melanoma as well as their clinicopathological and prognostic data were extracted from The Cancer Genome Atlas (TCGA) database. Ten pairs of melanoma-adjacent tissue samples were stored in liquid nitrogen at −80°C and used for real-time quantitative polymerase chain reaction (RT-qPCR) and Western blotting analysis. Our study was approved by the Ethics Committee of Shanghai East Hospital (K-KYSB-2020-0).
RNA Extraction and RT-qPCR
Total RNA was collected by using an RNA extraction kit (QIAGEN) and cDNA was synthesized by using a reverse transcription kit (Promega, Madison, USA) according to the manufacturer’s instructions. A PCR assay was performed using the SYBR Green Master Mix. After the reactions were finished, the relative expression levels of METTL3, SNHG3, miR-330-5p, and CNBP were quantified using the 2−ΔΔCt. The primer sequences used were summarized in Supplemental Table S1.
Western Blot Analysis
Melanoma tissue samples and A375 and SK-MEL-28 cells were lysed by RIPA buffer (P0013B, Beyotime, Shanghai, China). The supernatants were resolved in SDS-PAGE and transferred onto polyvinylidene fluoride (PVDF) membranes (Millipore, IPVH00010), incubated with anti-METTL3 (DF12020, Affinity, Changzhou, China), anti-CNBP (DF10045, Affinity, Changzhou, China), and anti-GAPDH (AB-P-R 001, Hangzhou, China) overnight at 4°C. Protein bands were scanned by enhanced chemiluminescence (ECL).
Plasmid, Short Hairpin RNA (shRNA), and Cell Transfection
METTL3 plasmid vectors, shRNA targeting METTL3 (sh-METTL3, 5′-GGAGGAGTGCATGAAAGCCAGTGAT-3′), and SNHG3 plasmid vectors were purchased from GenePharma (Shanghai, China). The negative vector (Vector) and si-NC were considered as the control groups. The A375 and SK-MEL-28 cells were planted in six-well plates 24 h prior to si-METTL3, METTL3 plasmids, and SNHG3 plasmid transfection with 50% to 60% confluence according to the manufacture instructions.
MTT, Colony Formation, and Transwell Assays
RNA Immunoprecipitation (RIP) and m6A Methylated RIP (MeRIP)
RNA-Binding Protein Immunoprecipitation Kit (17-700, Millipore), anti-m6A (68055-1-Ig, Proteintech, Wuhan, China), and anti-METTL3 (DF12020) were used for RIP and MeRIP-PCR assays according to the manufacture instructions.
Dual-Luciferase Reporter Assay
HEK293T cells were seeded into 24-well plates and PRL-TK-Luc report systems carrying wild-type (WT) or mutated (Mut) 3’UTR of SNHG3 or CNBP were co-transfected with miR-330-5p mimic or miR-NC into HEK293T cells. After the transfection for 48 h, luciferase activities were measured using a dual-luciferase reporter system.
The Xenograft Tumor Model
Nude male mice (6 weeks old) were purchased from Shanghai Laboratory Animal Central (Shanghai, China). A375 cells (1 × 107) transfected with sh-METTL3 or shNC lentiviruses were subcutaneously injected into the right flank of the mice. After 33 days, the mice were sacrificed and the xenografted tumors were collected for immunohistochemistry analysis. Animal experiments were approved by the Ethics Committee of Shanghai East Hospital (K-KYSB-2020-0).
Statistical Analysis
Statistical analyses were processed with GraphPad Prism 5 (La Jolla, CA, USA). Values are expressed as mean ± standard deviation. Chi-square, Student t test, and analysis of variance were used for comparisons between groups. Kaplan–Meier analysis was used to analyze the association of METTL3, SNHG3, or miR-330-5p with the prognosis of patients with melanoma. A Cox proportional hazard model was used to assess the risk of METTL3 in patients with melanoma. Pearson’s correlation analysis was used to analyze the correlation of SNHG3 with miR-330-5p or METTL3 expression in melanoma. P values <0.05 were considered statistically significant.
Other experimental assays can be provided in the supplementary data.
Results
Upregulation of METTL3 Was Associated With Poor Survival in Patients With Melanoma
METTL3 mRNA expression was measured in 474 melanoma tissue samples from TCGA data set, which indicated that METTL3 expression increased in melanoma tissue samples compared with adjacent normal tissues (Fig. 1A). Elevated expression of METTL3 was validated in 10 pairs of melanoma-adjacent tissue samples by RT-qPCR (Fig. 1B) and Western blotting analyses (Fig. 1C, D). Furthermore, we found that elevated expression of METTL3 showed no correlation with clinical characteristics in patients with melanoma (Supplemental Table S2). Kaplan–Meier analysis revealed that patients with high expression of METTL3 had poor overall survival compared with those with low expression of METTL3, but experienced no significant difference in tumor recurrence (Fig. 1E). Univariate and multivariate Cox regression analyses revealed that high expression of METTL3 was an independent prognostic factor for poor survival in patients with melanoma (Supplemental Table S3).

Elevated expression of METTL3 is associated with poor survival in patients with melanoma. (A) TCGA analysis of the expression of METTL3 in 474 non-paired melanoma samples. (B) RT-qPCR analysis of the expression levels of METTL3 in 10 pair-matched melanoma tissue samples. (C, D) Western blot analysis of the expression of METTL3 in 10 pair-matched melanoma samples. (E) Kaplan–Meier analysis of the association of METTL3 expression with overall survival and tumor recurrence in patients with melanoma. GAPDH: glyceraldehyde-3-phosphate dehydrogenase; METTL3: methyltransferase 3; RT-qPCR: real-time quantitative polymerase chain reaction; TCGA: The Cancer Genome Atlas.
METTL3 Knockdown Inhibited Melanoma Cell Proliferation and Invasion
The transfection efficiencies of sh-METTL3 in melanoma cells were determined by RT-qPCR and Western blotting analyses, which showed that METTL3 expression levels were clearly decreased compared with the short hairpin RNA and negative control (sh-NC) group after transfection with sh-METTL3 into A375 and SK-MEL-28 cells for 48 h (Fig. 2A). The MTT and colony formation assays indicated that METTL3 knockdown suppressed cell proliferation (Fig. 2B) and colony formation capacity (Fig. 2C) in A375 and SK-MEL-28 cells. The Transwell assay also showed that the knockdown of METTL3 weakened the invasive ability of cells in A375 and SK-MEL-28 cell lines (Fig. 2D).

Knockdown of METTL3 hinders melanoma cell proliferation and invasion. (A) RT-qPCR and Western blotting analysis of the expression levels of METTL3 after transfection with sh-METTL3 in A375 and SK-MEL-28 cell lines. (B) MTT analysis of cell proliferation viability after transfection with sh-METTL3 into A375 and SK-MEL-28 cells. (C) Colony formation analysis of cell colony numbers after transfection with sh-METTL3 into A375 and SK-MEL-28 cells. (D) Transwell analysis of cell invasion capacity after transfection with sh-METTL3 into A375 and SK-MEL-28 cells. Data shown are the mean ± SEM of three experiments. GAPDH: glyceraldehyde-3-phosphate dehydrogenase; METTL3: methyltransferase 3; MTT: (3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide); RT-qPCR: real-time quantitative polymerase chain reaction; SEM: standard error of the mean. *P < 0.05; **P < 0.01; ***P < 0.001.
METTL3 Overexpression Accelerated the Proliferation and Invasion of Melanoma Cells
The high expression of METTL3 induced by transfection with METTL3 plasmids was validated by RT-qPCR and Western blotting analyses in A375 and SK-MEL-28 cell lines (Fig. 3A). The MTT and colony formation assays showed that ectopic expression of METTL3 promoted cell proliferation activity (Fig. 3B) and cell colony formation (Fig. 3C) in A375 and SK-MEL-28 cell lines. The Transwell assay revealed that overexpression of METTL3 could enhance the invasion of A375 and SK-MEL-28 cells (Fig. 3D).
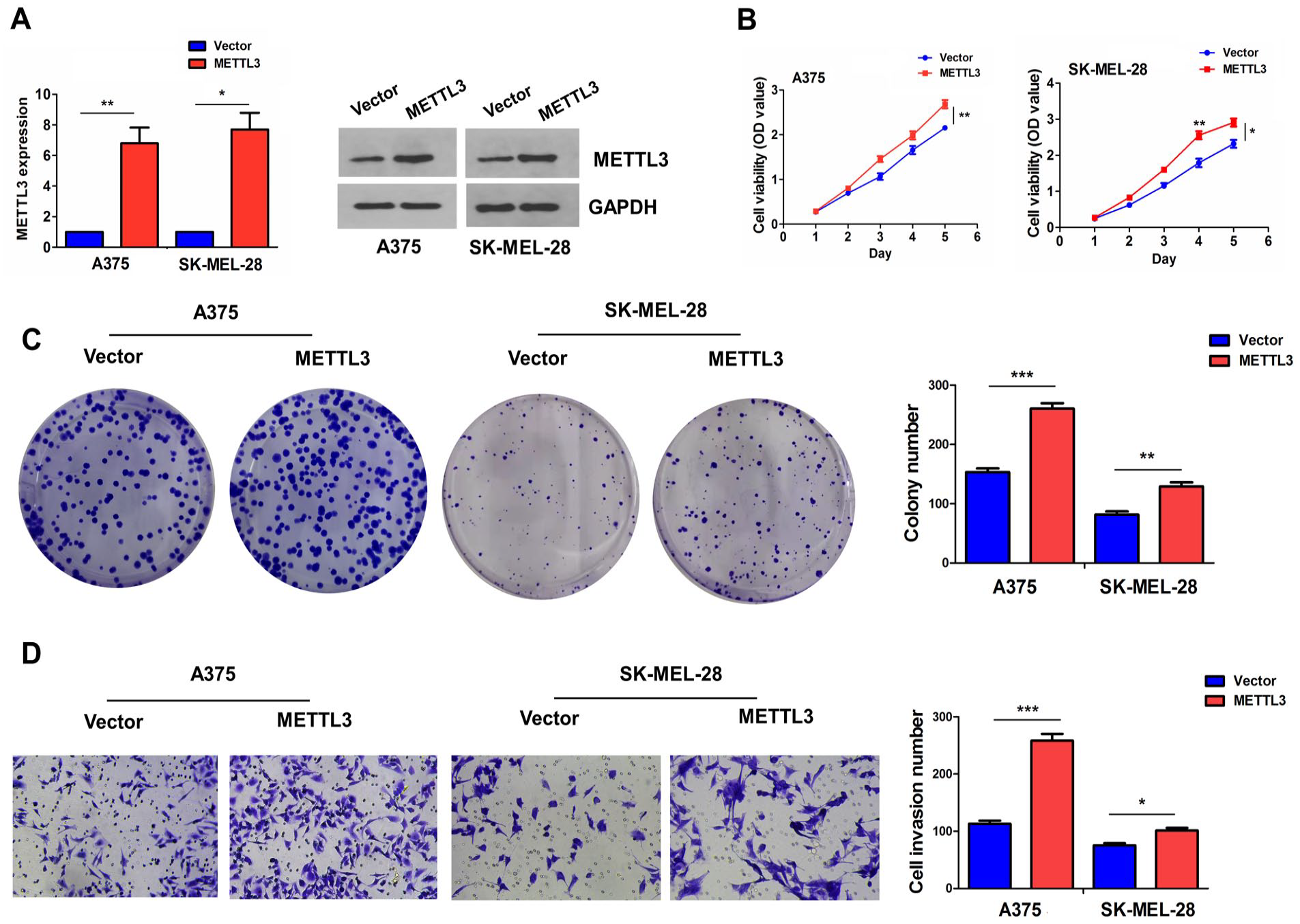
METTL3 enhances melanoma cell growth and invasion. (A) RT-qPCR and Western blotting analysis of METTL3 expression levels after transfection with METTL3 plasmids into A375 and SK-MEL-28 cell lines. (B) MTT analysis of cell viability after transfection with METTL3 plasmids into A375 and SK-MEL-28 cells. (C) Colony formation analysis of cell colony number after transfection with METTL3 plasmids into A375 and SK-MEL-28 cells. (D) Transwell analysis of cell invasion capacity after transfection with METTL3 plasmids into A375 and SK-MEL-28 cells. Data shown are the mean ± SEM of three experiments. GAPDH: glyceraldehyde-3-phosphate dehydrogenase; METTL3: methyltransferase 3; MTT: (3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide); RT-qPCR: real-time quantitative polymerase chain reaction; SEM: standard error of the mean. *P < 0.05; **P < 0.01; ***P < 0.001.
LncRNA SNHG3 Was Identified as an m6A Target of METTL3 in Melanoma Cells
The m6A targets of METTL3 were identified using the Starbase3.0 tool (https://starbase.sysu.edu.cn), which revealed that METTL3 harbored the most promising binding sites on lncRNA SNHG3. We hypothesized that METTL3 might regulate SNHG3 expression through m6A modification in melanoma cells. Analysis of m6A dot blot and MeRIP showed that METTL3 knockdown reduced total m6A levels (Fig. 4A) as well as SNHG3 m6A levels (Fig. 4B) in A375 and SK-MEL-28 cells. Meanwhile, RT-qPCR analysis indicated that METTL3 knockdown decreased SNHG3 expression, whereas overexpression of METTL3 had the opposite effects in A375 and SK-MEL-28 cell lines (Fig. 4C). RIP analysis was used to confirm binding between METTL3 and SNHG3 in A375 and SK-MEL-28 cells, and showed that endogenous levels of SNHG3 were enriched in the METTL3 pellet compared with those of the IgG group (Fig. 4D).

LncRNA SNHG3 is identified as an m6A target of METTL3 in melanoma cells. (A) m6A dot blot analysis of the effects of METTL3 knockdown on total m6A levels in A375 and SK-MEL-28 cell lines. (B) MeRIP-PCR analysis of the effects of METTL3 knockdown on the m6A levels of SNHG3 in A375 and SK-MEL-28 cells. (C) RT-qPCR analysis of the effects of METTL3 knockdown or overexpression on the mRNA expression of SNHG3 in A375 and SK-MEL-28 cell lines. (D) RIP for METTL3 was performed to examine the endogenous expression of SNHG3 in A375 and SK-MEL-28 cells. Data shown are the means ± SEM of three experiments. MeRIP: methylated RNA immunoprecipitation; METTL3: methyltransferase 3; RIP: RNA immunoprecipitation; RT-qPCR: real-time quantitative polymerase chain reaction; SEM: standard error of the mean; SNHG3: small nucleolar RNA host gene 3. **P < 0.01; ***P < 0.001; ****P < 0.0001.
The Expression of miR-330-5p Was Correlated With the lncRNA SNHG3 and a Favorable Prognosis in Patients With Melanoma
Our cohort showed that SNHG3 mRNA levels were markedly elevated in 10 pairs of melanoma-adjacent tissue samples (Fig. 5A), and this result was further verified by TCGA cohort in 387 cases of melanoma samples. METTL3 harbored a positive correlation with the expression of lncRNA SNHG3 in melanoma (n = 352, Fig. 5B). However, SNHG3 did not show any association with clinicopathological characteristics (Supplemental Table S4) and clinical prognosis, including overall survival and tumor recurrence in patients with melanoma (Supplemental Fig. S1). Furthermore, depending on the cellular context, lncRNA SNHG3 had the binding with miR-330-5p, miR-514a, and miR-326, as identified by the StarBaseV3.0 tool, of which only miR-330-5p showed a negative correlation with the expression of SNHG3 in melanoma samples (Figs. 5D and Supplemental Fig. S2A). Our cohort indicated that miR-330-5p mRNA levels were decreased markedly in 10 pairs of melanoma-adjacent tissue samples (Fig. 5C). The expression of miR-330-5p did not show an association with clinicopathological characteristics in patients with melanoma (Supplemental Table S5). However, Kaplan–Meier analysis revealed that patients with miR-330-5p high expression achieved more favorable survival compared with those with low expression of miR-330-5p (Fig. 5E), but showed no difference in tumor recurrence (Supplemental Fig. S2B).
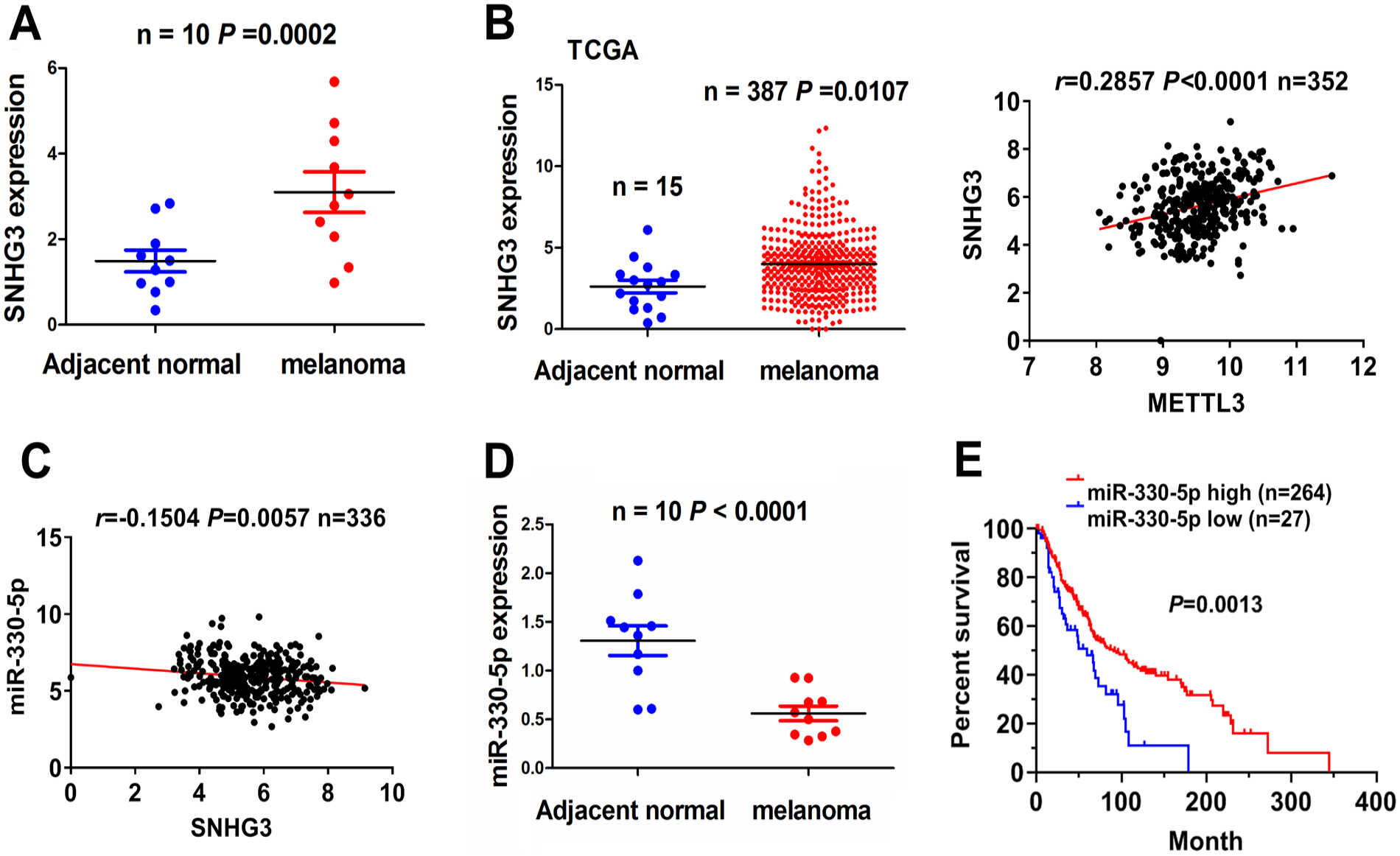
miR-330-5p expression is associated with favorable survival in patients with melanoma. (A) RT-qPCR analysis of the expression of SNHG3 in 10 pairs of matched melanoma-adjacent healthy tissue samples. (B) TCGA analysis of the expression of SNHG3 in 387 melanoma samples and Pearson’s correlation analysis of the correlation between METTL3 and SNHG3 in melanoma samples. (C) Pearson’s correlation analysis of the correlation between miR-330-5p and SNHG3 in melanoma samples. (D) RT-qPCR analysis of miR-330-5p expression in 10 pairs of matched melanoma-adjacent tissue matched samples. (E) Kaplan–Meier analysis of the association of miR-330-5p with overall survival in patients with melanoma. METTL3: methyltransferase 3; RT-qPCR: real-time quantitative polymerase chain reaction; SNHG3: small nucleolar RNA host gene 3; TCGA: The Cancer Genome Atlas.
LncRNA SNHG3 Acted as a Sponge of miR-330-5p in Melanoma Cells
The binding sites between SNHG3 and miR-330-5p are shown in Fig. 6A. We found that miR-330-5p mimics could reduce the luciferase activities of the wildtype SNHG3 3’UTR compared with the mutated SNHG3 3’UTR in HEK293T cells (Fig. 6B). Knockdown of SNHG3 or METTL3 increased miR-330-5p expression, whereas overexpression of METTL3 repressed its expression in A375 and SK-MEL-28 cells (Fig. 6C, D). However, exposure to the miR330-5p inhibitor had no effect on SNHG3 expression in A375 or SK-MEL-28 cell lines (Supplemental Fig. S3). Furthermore, ectopic expression of SNHG3 drove cell proliferation and invasion, but miR-330-5p mimics deteriorated SNHG3-induced tumor-promoting effects in A375 and SK-MEL-28 cells (Fig. 6E–G).

LncRNA SNHG3 acts as a sponge of miR-330-5p in melanoma cells. (A) Schematic representation of the binding sites between miR-330-5p and wildtype (WT) or mutated (Mut) SNHG3 3’UTR. (B) Luciferase gene reporter analysis of the effects of miR-330-5p mimics on the luciferase activities of WT or Mut SNHG3 3’UTR in HEK293T cells. (C) RT-qPCR analysis of the effects of SNHG3 knockdown on miR-330-5p expression in A375 and SK-MEL-28 cells. (D) RT-qPCR analysis of the effects of METTL3 knockdown or overexpression on the expression levels of miR-330-5p in A375 and SK-MEL-28 cells. (E) MTT and (F, G) Transwell analysis of cell viability and invasion after co-transfection with miR-330-5p mimics and SNHG3 plasmids in A375 and SK-MEL-28 cells. Data shown are the mean ± SEM of three experiments. METTL3: methyltransferase 3; MTT: (3-(4,5-dimethyl-2-thiazolyl)-2,5-diphenyl-2-H-tetrazolium bromide); RT-qPCR: real-time quantitative polymerase chain reaction; SEM: standard error of the mean; SNHG3: small nucleolar RNA host gene 3. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001.
A miR-330-5p Inhibitor Abolished SNHG3 Knockdown–Induced CNBP Expression in Melanoma Cells
Using the StarBaseV3.0 database tool, CNBP was identified as a direct target of miR-330-5p. CNBP mRNA and protein levels were significantly elevated in 10 pairs of melanoma-adjacent tissue samples (Fig. 7A, B). The binding sites between CNBP and miR-330-5p are shown in Fig. 7C. We found that miR-330-5p mimics could reduce the luciferase activities of the wildtype CNBP 3’UTR compared with the mutated CNBP 3’UTR in HEK293T cells (Fig. 7D). Furthermore, RT-qPCR and Western blotting analyses indicated that exposure to the miR-330-5p inhibitor increased CNBP expression and attenuated the downregulation induced by SNHG3 knockdown in A375 and SK-MEL-28 cell lines (Fig. 7E, F).

miR-330-5p inhibitor reverses SNHG3 knockdown–induced downregulation of CNBP in melanoma cells. (A) RT-qPCR and (B) Western blotting analysis of CNBP expression in 10 pairs of matched melanoma-adjacent tissue samples. (C) Schematic representation of the binding sites between miR-330-5p and WT or Mut CNBP 3’UTR. (D) Luciferase gene reporter analysis of the effects of miR-330-5p mimics on the luciferase activities of WT or Mut CNBP 3’UTR in HEK293T cells. (E) RT-qPCR and (F) Western blotting analysis of the expression of CNBP after co-transfection with si-SNHG3 and miR-330-5p inhibitor in A375 and SK-MEL-28 cells. Data shown are the mean ± SEM of three experiments. CNBP: CCHC-type zinc finger nucleic acid binding protein; GAPDH: glyceraldehyde-3-phosphate dehydrogenase; Mut: mutated; RT-qPCR: real-time quantitative polymerase chain reaction; SEM: standard error of the mean; SNHG3: small nucleolar RNA host gene 3; WT or Mut: wildtype or mutated. *P < 0.05; **P < 0.01; ***P < 0.001.
Knockdown of METTL3 Repressed Melanoma Tumorigenesis
To validate whether METTL3 influences in vivo melanoma tumorigenesis, sh-METTL3 or shNC stably transfected A375 cells were used to establish subcutaneous xenograft tumor models by injecting the engineered cells into the flank of nude mice. When the mice were sacrificed, we found that the volumes of xenograft tumors formed by sh-METTL3 transfected A375 cells were smaller than those of sh-NC transfected cells (Fig. 8A). The tumor growth curve demonstrated that the xenografted tumors in the sh-METTL3 group showed a relatively slow growth trend in a time-dependent manner (Fig. 8B). Both tumor volume and weight were much smaller in the sh-METTL3 group than in the shNC group (Fig. 8C). Immunohistochemistry analysis indicated that the positive expression of VEGF, a growth-related marker, was markedly reduced in the sh-METTL3 group compared with the NC group (Fig. 8D).

Knockdown of METTL3 represses in vivo melanoma tumorigenesis. (A) Schematic representation of xenograft tumors between sh-METTL3 and shNC transfected groups. (B) A tumor growth curve analysis of the tumor growth trend in sh-METTL3 and shNC transfected groups. (C) Comparison of the tumor volume and tumor weight between sh-METTL3 and shNC transfected groups. (D) Immunohistochemistry analysis of the positive expression of VEGF in sh-METTL3 and shNC transfected groups. Data shown are the mean. METTL3: methyltransferase 3; VEGF: vascular endothelial growth factor.
Discussion
An increasing number of studies have shown that m6A methylation plays a crucial role in cancer, including melanoma4,11. METTL3 is upregulated in gastric cancer 18 , bladder cancer 19 , and in intrahepatic cholangiocarcinoma 20 , and predicts a poor prognosis in these cancer patients18–20. However, METTL3 is downregulated in breast cancer and associated with favorable survival in patients with triple-negative breast cancer (TNBC) 21 . METTL3 is upregulated in patients with acral melanoma and associated with a higher tumor stage 12 . Accordingly, we found that METTL3 was upregulated in melanoma samples and was associated with poor survival, acting as an independent prognostic factor in patients with melanoma. Our findings suggested that METTL3 could be a potential prognostic biomarker for melanoma.
METTL3 has a dual role in different malignancies. In most cases, it can act as an oncogenic factor to accelerate tumor proliferation and metastasis18–20, but it shows a suppressive role in TNBC 21 . Current studies indicate that METTL3 plays a carcinogenic role in melanoma cells12,14. In our study, we found that METTL3 knockdown inhibited melanoma cell proliferation and invasion, but overexpression of METTL3 promoted its growth in vitro and in vivo, suggesting that METTL3 could be an oncogenic factor in melanoma cells.
Mechanistically, METTL3 can regulate cancer progression by m6A-dependent modification of mRNAs and noncoding RNAs22,23. For example, METTL3-mediated m6A modification of homo sapiens hepatoma-derived growth factor (HDGF) mRNA drives gastric cancer metastasis 24 and METTL3 facilitates the stability of metastasis-associated lung adenocarcinoma transcript 1 (MALAT1) in an m6A-dependent manner and stimulates glioma and prostate cancer progression25,26. In addition, METTL3 displays the oncogenic properties by m6A modification of lncRNA THAP7-AS1 27 , SNHG7 28 , and forkhead box D2 adjacent opposite strand RNA 1 (FOXD2-AS1) 29 . Our results showed that inhibition of METTL3 reduced the m6A methylation levels of SNHG3, which in turn could act as a sponge for miR-330-5p to regulate melanoma cell proliferation and invasion. LncRNA SNHG3 favors the metastasis of colorectal cancer by regulating the miR-539/RUNX2 axis 30 and miR-330 can suppress the growth of melanoma cells 31 . Altogether, these findings suggest that METTL3-mediated m6A modification of the SNHG3/miR-330-5p axis can promote melanoma cell growth.
CNBP has been reported to promote cancer cell proliferation and invasion by upregulation of matrix metalloproteinase (MMP)-2/-14 32 . The miR-330-3p/CNBP axis can enhance proliferation and invasion of colorectal cancer 33 . Herein, we found that CNBP was upregulated in melanoma samples and exposure to the miR-330 inhibitor increased CNBP expression and counteracted the downregulation of CNBP induced by SNHG3 knockdown in melanoma cells. These results indicated that METTL3 mediated the SNHG3/miR-330-5p/CNBP axis to promote growth and invasion of melanoma cells.
In summary, our study demonstrated that upregulation of METTL3 was associated with poor survival in patients with melanoma, and METTL3-mediated m6A modification of lncRNA SNHG3 promoted the growth and invasion of melanoma cells by regulating the miR-330-5p/CNBP axis. Our findings provide a promising target for the treatment of patients with melanoma.
Supplemental Material
sj-docx-1-cll-10.1177_09636897231188300 – Supplemental material for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis
Supplemental material, sj-docx-1-cll-10.1177_09636897231188300 for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis by Shaojun Chu, Yulong Li, Baojin Wu, Guo Rong, Qiang Hou, Qin Zhou, Dexiang Du and Yufei Li in Cell Transplantation
Supplemental Material
sj-tif-2-cll-10.1177_09636897231188300 – Supplemental material for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis
Supplemental material, sj-tif-2-cll-10.1177_09636897231188300 for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis by Shaojun Chu, Yulong Li, Baojin Wu, Guo Rong, Qiang Hou, Qin Zhou, Dexiang Du and Yufei Li in Cell Transplantation
Supplemental Material
sj-tif-3-cll-10.1177_09636897231188300 – Supplemental material for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis
Supplemental material, sj-tif-3-cll-10.1177_09636897231188300 for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis by Shaojun Chu, Yulong Li, Baojin Wu, Guo Rong, Qiang Hou, Qin Zhou, Dexiang Du and Yufei Li in Cell Transplantation
Supplemental Material
sj-tif-4-cll-10.1177_09636897231188300 – Supplemental material for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis
Supplemental material, sj-tif-4-cll-10.1177_09636897231188300 for METTL3 Promotes the Growth and Invasion of Melanoma Cells by Regulating the lncRNA SNHG3/miR-330-5p Axis by Shaojun Chu, Yulong Li, Baojin Wu, Guo Rong, Qiang Hou, Qin Zhou, Dexiang Du and Yufei Li in Cell Transplantation
Footnotes
Ethical Approval
This study was approved by the Ethics Committee of Shanghai East Hospital.
Statement of Human and Animal Rights
All procedures in this study were conducted in accordance with the Ethics Committee of Shanghai East Hospital.
Statement of Informed Consent
Written informed consent was obtained from the patients for their anonymized information to be published in this article.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the east hospital affiliated to Tongji University–introduced talent research startup fund (DFRC2019008) and the featured Clinical Discipline Project of Shanghai Pudong (PWYts2021-07).
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
