Abstract
Objectives
METTL3 as an m6A methyltransferase acts in diverse malignancies including gastric cancer (GC). We aimed to reveal the underlying mechanisms by which METTL3 contributes to gastric carcinogenesis.
Methods
The association of METTL3 and SNHG3 with GC was analyzed by qRT-PCR, Western blot, and TCGA cohort. The functional experiments were implemented to uncover the role of METTL3 in GC. m6A dot blot and MeRIP were used to determine METTL3-mediated m6A modification of lncRNA SNHG3. The effect of METTL3 on SNHG3-mediated miR-186-5p/cyclinD2 axis was evaluated by luciferase gene report, RT-qPCR, and Western blot assays.
Results
We found that METTL3 was remarkably elevated in GC tissues and correlated with poor survival in patients with GC. Silencing of METTL3 impaired GC cell growth and invasion, whereas restored METTL3 expression promoted these effects. Mechanistically, reduced expression of METTL3 decreased SNHG3 m6A level and caused a decrease in SNHG3 expression, which could further act as a sponge of miR-186-5p to upregulate cyclinD2. Overexpression of SNHG3 attenuated METTL3 knockdown-induced anti-proliferating and miR-186-5p upregulation and cyclinD2 downregulation.
Conclusion
We find that METTL3-mediated m6A modification of lncRNA SNHG3 accelerates GC progression by modulating miR-186-5p/cyclinD2 axis.
Introduction
Gastric cancer (GC) harbors the fifth incidence and the third cancer-related mortality. 1 Although enormous efforts have been made to improve poor prognosis in GC, the advanced patients still display discouraging prognosis duo to the widespread spread of tumors. 2 Deregulated epigenetic modification is implicated in the pathogenesis and progression of GC. 3 Therefore, further investigations of the molecular pathogenesis of GC is indispensable for the treatment of GC.
N6-methyladenosine (m6A) is dynamically regulated by m6A methyltransferases such as methyltransferase-like 3/14 (METTL3/14)/WTAP/KIAA1429 and removed by m6A demethylase FTO and ALKBH5. 4 Previous studies showed that m6A methylation plays a critical role in GC. The dysregulation of METTL3/14/WTAP/KIAA1429 or FTO/ALKBH5 participates in GC progression.5–10
METTL3 as an m6A methyltransferase participates in multiple malignancies including gastric carcinogenesis. 11 METTL3 has been indicated to be upregulated in GC, 5 prostate cancer (PCa), 12 non-cell lung cancer (NSCLC), 13 esophageal squamous cell carcinoma (ESCC), 14 bladder cancer (BCa), 15 oral squamous cell carcinoma (OSCC), 16 and colorectal cancer (CRC), 17 associated with poor prognosis5,14,16,17 and facilitated tumorigenesis and metastasis.5,12–17 STM2457, a selective catalytic inhibitor of METTL3 can be used as a potential therapeutic strategy against acute myeloid leukemia. 18 However, METTL3 is downregulated in renal cell carcinoma (RCC), associates with favorable prognosis, and suppresses cell proliferation and invasion. 19
METTL3 has been shown to promote tumorigenesis and invasion of GC.5,20 However, the effects of METTL3 on GC tumorigenesis remain undocumented. We herein found that elevated expression of METTL3 was related to poor overall survival in GC, and METTL3-mediated m6A modification of lncRNA SNHG3 facilitated GC growth by regulating miR-186-5p/cyclinD2 axis.
Materials and methods
Clinical samples
The Cancer Genome Atlas database (TCGA) 21 was used for analysis of the prognosis of METTL3, SNHG3, and miR-186-5p with GC. 10 pairs of GC samples were used for analysis of the expression of METTL3, SNHG3 and miR-186-5p. Our study protocol was approved by the Ethics Committee of Shanghai Tenth People’s Hospital (2021-YS-009).
RNA extraction and real-time quantitative PCR (RT-qPCR)
Total RNA was collected by using Trizol (Ambion, cat No. 15596-026), and cDNA was synthesized by using HiScript® II Q Select RT SuperMix (VAZYME, cat No. R233) according to the manufacturer’s instructions. The qPCR was performed using the SYBR Green Master Mix (VAZYME, cat No. Q111-02). After the reactions were finished, the expression levels of METTL3, SNHG3, miR-186-5p, and cylinD2 relative to the expression of GAPDH or U6 were quantified using the 2−ΔΔCt. The primer sequences used are shown in Supplementary Table S1.
MTT, colony formation, and transwell assays
These assays were performed as according to the previous report. 6
The other experimental methods used in this paper could be provided in the supplementary methods.
Results
Upregulation of METTL3 is an independent prognostic factor of poor survival in patients with GC
The mRNA expression levels of METTL3 were assessed in 380 GC tissue samples from the TCGA cohort,
21
which showed that METTL3 expression was dramatically elevated in 34 pare-matched and 380 non-matched GC tissue samples (Figure 1(a)). The increased expression of METTL3 in GC tissue samples was further confirmed in the transcriptional (Figure 1(b)) and translational levels (Figure 1(c) and Supplementary Figure S1). Then, increased expression of METTL3 displayed no link to the pathological features in GC (Supplementary Table S2). ALL and GSE14210 cohorts unveiled that the GC patients with METTL3-high expression indicated an unfavorable survival as compared with those with METTL3-low expression (Figure 1(d)). Multivariate Cox regression analysis uncovered that METTL3 was a prognostic marker for poor survival in GC (Supplementary Table S3). The elevated expression of METTL3 was associated with poor survival in GC. (a) Analysis of the levels of METTL3 in GC. (b) Examination of the levels of METTL3 in GC. (c) Western blot for detection of the levels of METTL3 in GC. (d) Kaplan–Meier analysis of the association of METTL3 expression with overall survival in patients with GC.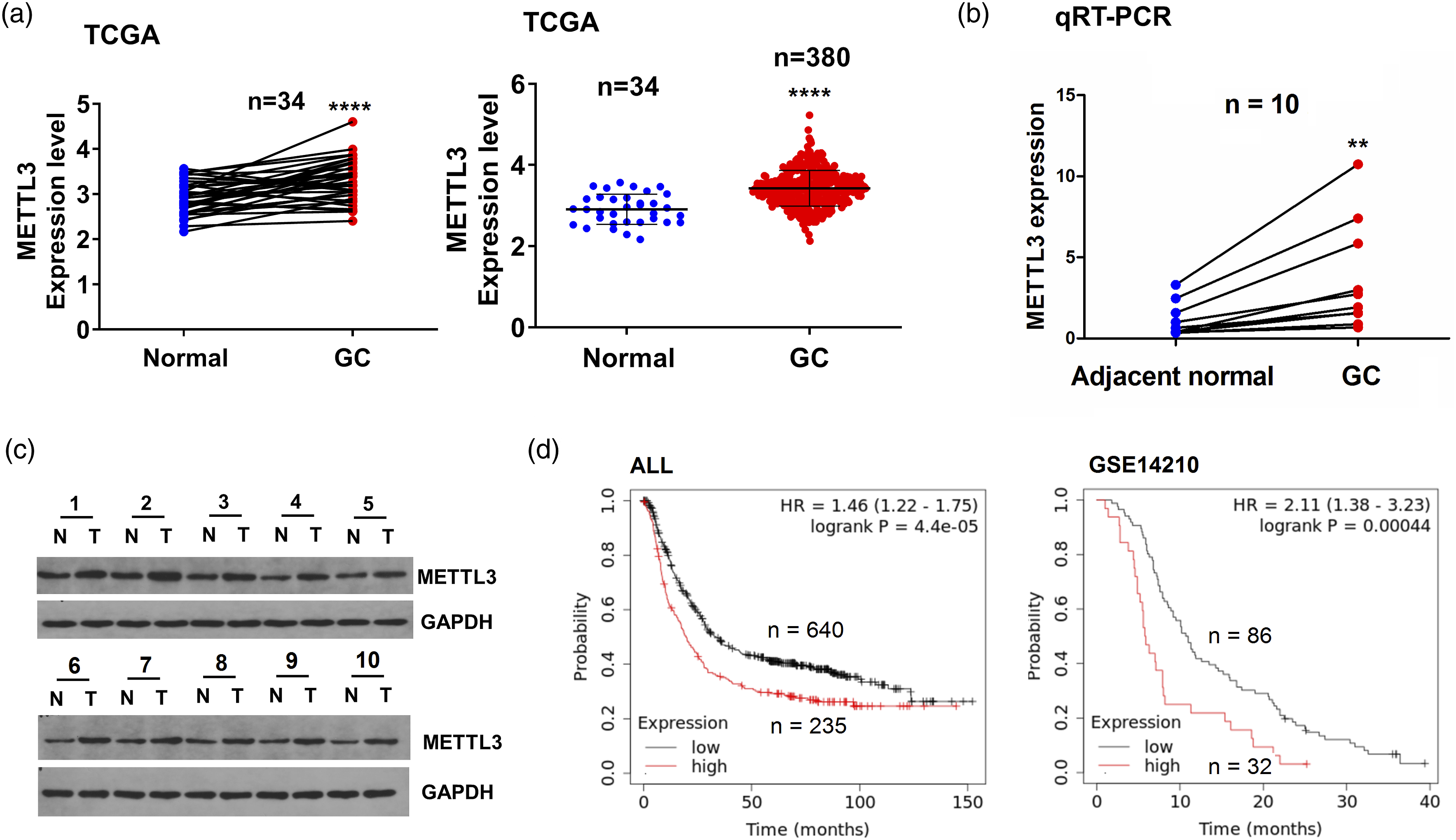
Knockdown of METTL3 represses the growth and invasion of GC cells
We detected the expression levels of METTL3 in GC cell liens and found that METTL3 expression was raised in MKN-45 and BGC-823 cell lines as compared with the normal GES-1 cells (Figure S2). The transfection levels of si-METTL3 in MKN-45 and BGC-823 cells were examined by RT-qPCR (Figure 2(a)) and Western blot analyses (Figure 2(b)), indicating that METTL3 mRNA and protein levels had significantly reduced expression in MKN-45 and BGC-823 cells after transfection with si-METTL3 for 48 h. Proliferation assays indicated that reduced expression of METTL3 repressed the cell proliferation activities (Figure 2(c)) and colony formation (Figure 2(d)) in MKN-45 and BGC-823 cells. Transwell assay was performed to assess the cell invasion abilities, indicating that reduced expression of METTL3 blocked the cell invasive capabilities in MKN-45 and BGC-823 cells (Figure 2(e)). Reduced METTL3 expression impaired GC cell growth and invasion. (a) RT-qPCR and (b) Western blotting analysis of the mRNA and protein levels of METTL3 after the treatment with si-METTL3 in MKN-45 and BGC-823 cells. (c) Examination of cell proliferation after treatment with si-METTL3 in MKN-45 and BGC-823 cells. (d) Examination of cell colony number after treatment with si-METTL3 in MKN-45 and BGC-823 cells. (e) Examination of cell invasion after treatment with si-METTL3 in MKN-45 and BGC-823 cells.
Restored expression of METTL3 facilitates GC cell growth and invasion
The transfection levels of METTL3 overexpression plasmids were determined by RT-qPCR (Figure 3(a)) and Western blot analyses (Figure 3(b)), suggesting that the METTL3 mRNA and protein levels had elevated expression in MKN-45 and BGC-823 cells after transfection with METTL3 plasmids for 48 h. Proliferation assays showed that restored expression of METTL3 enhanced cell proliferation activities (Figure 3(c)) and colony formation abilities (Figure 3(d)) in MKN-45 and BGC-823 cells. Transwell was used to measure cell-invasive capabilities, indicating that overexpression of METTL3 boosted cell-invasive capabilities in MKN-45 and BGC-823 cells (Figure 3(e)). METTL3 overexpression drove GC cell growth. (a) RT-qPCR and (b) Western blotting analysis of the mRNA and protein levels of METTL3 after the transfection with METTL3 plasmids into MKN-45 and BGC-823 cell lines. (c) Examination of the cell proliferation after treatment with METTL3 overexpression in MKN-45 and BGC-823 cells. (d) Examination of cell colony number after treatment with METTL3 overexpression in MKN-45 and BGC-823 cells. (e) Examination of cell invasion after treatment with METTL3 overexpression in MKN-45 and BGC-823 cells.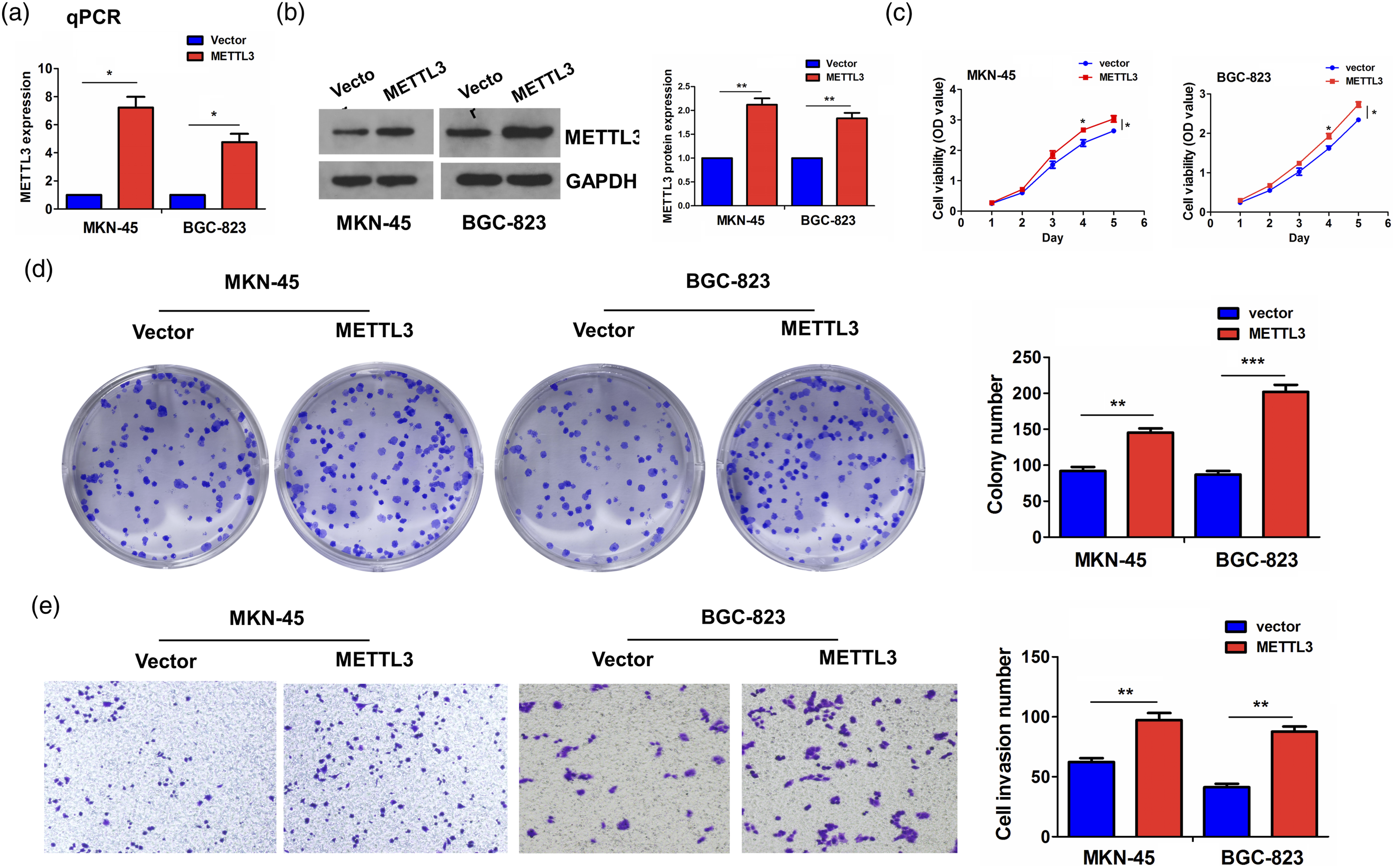
The lncRNA SNHG3 is identified as an m6A target of METTL3 in GC cells
We screened the m6A targets of METTL3 by starbase3.0, which has the binding sites with lncRNA SNHG3. The m6A dot blot indicated that knockdown of METTL3 decreased the whole m6A levels in MKN-45 and BGC-823 cells (Figure 4(a)). MeRIP-PCR showed that knockdown of METTL3 decreased the m6A levels of SNHG3 in MKN-45 and BGC-823 cells (Figure 4(b)). The RT-PCR showed that knockdown of METTL3 reduced the expression levels of SNHG3, but overexpression of METTL3 promoted its expression in MKN-45 and BGC-823 cells (Figure 4(c)). RIP assay was employed to assess METTL3-specific binding with SNHG3 in MKN-45 and BGC-823 cells and found the endogenous levels of SNHG3 pulled-down from METTL3-expressed cells were predominantly enriched in the METTL3 pellet compared with those in the IgG control (Figure 4(d)). In addition, we performed the MTT proliferation assay to illustrate the effects of METTL3 on GC cells that could be regulated by SNHG3 in GC cells. We found that knockdown of METTL3 repressed the cell proliferation, while overexpression of SNHG3 promoted the cell proliferation and reversed METTL3-knockdown-induced anti-proliferation effect in GC cells (Figure 4(e) and (f)). Identification of SNHG3 as an m6A target of METTL3. (a) The total m6A levels after treatment with si-METTL3 in GC cells. (b) MeRIP-PCR analysis of the m6A levels of SNHG3 after treatment with si-METTL3 in MKN-45 and BGC-823 cells. (c) Investigations of the effects of METTL3 knockdown or overexpression on the mRNA levels of SNHG3 in MKN-45 and BGC-823 cells. (d) RIP analysis of the interaction of SNHG3 mRNAs with METTL3 protein in MKN-45 and BGC-823 cells. (e) MTT analysis of the cell viability after co-transfection with si-METTL3 and SNHG3 overexpression plasmids in MKN-45 and BGC-823 cells.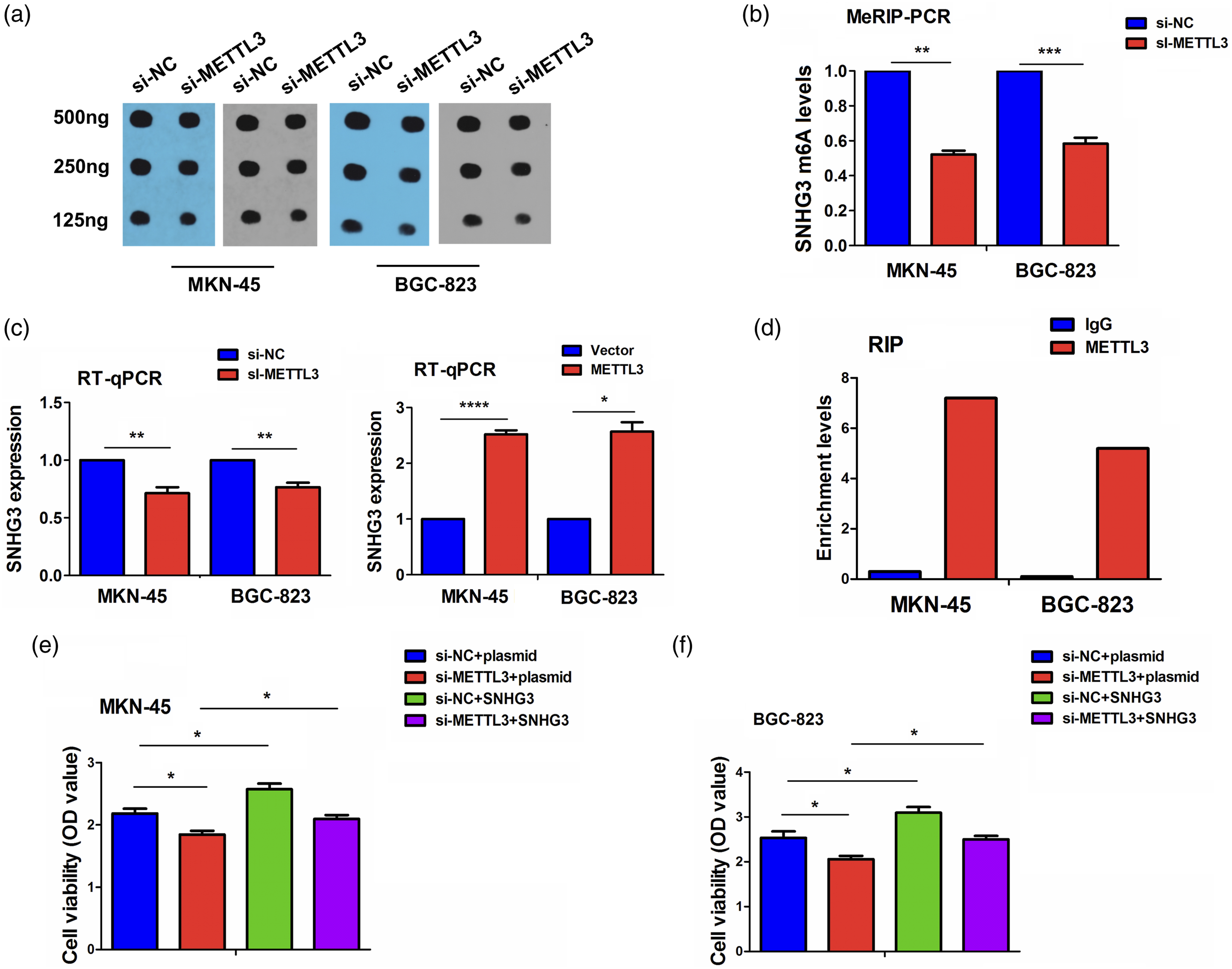
Upregulation of SNHG3 is a prognostic marker of poor survival in patients with GC
The mRNA expression levels of SNHG3 were assessed in 380 GC tissue samples from the TCGA cohort,
21
which showed that SNHG3 expression was dramatically elevated in 34 pare-matched and 380 non-matched GC tissue samples as compared with adjacent normal tissues (Figure 5(a)). The expression of SNHG3 was validated to be increased in 10 pairs of GC tissue samples in our cohort by qRT-PCR (Figure 5(b)) and exhibited a positive correlation with METTL3 in 380 GC tissue samples from TCGA cohort (Figure 5(c)). We also found that METTL3 expression was increased in MKN-45 and BGC-823 cell lines as compared with the normal GES-1 cells (Figure S3). Then, SNHG3 exerted no correlation with the pathological features in GC (Supplementary Table S4). ALL and GSE22377 cohorts unveiled that the GC patients with SNHG3-high expression indicated a poor overall survival as compared with those with SNHG3-low expression (Figure 5(d)). Multivariate Cox regression analysis uncovered that SNHG3 was a prognostic marker in GC (Supplementary Table S5). Increased SNHG3 expression is associated with poor survival in GC. (a) Detection of the levels of SNHG3 in 34 pair-matched and 380 non-paired GC. (b) Detection of the levels of SNHG3 in 10 pairs of GC. (c) The correlation between METTL3 and SNHG3 expression in GC. (d) Kaplan–Meier analysis of the association of SNHG3 with overall survival in patients with GC.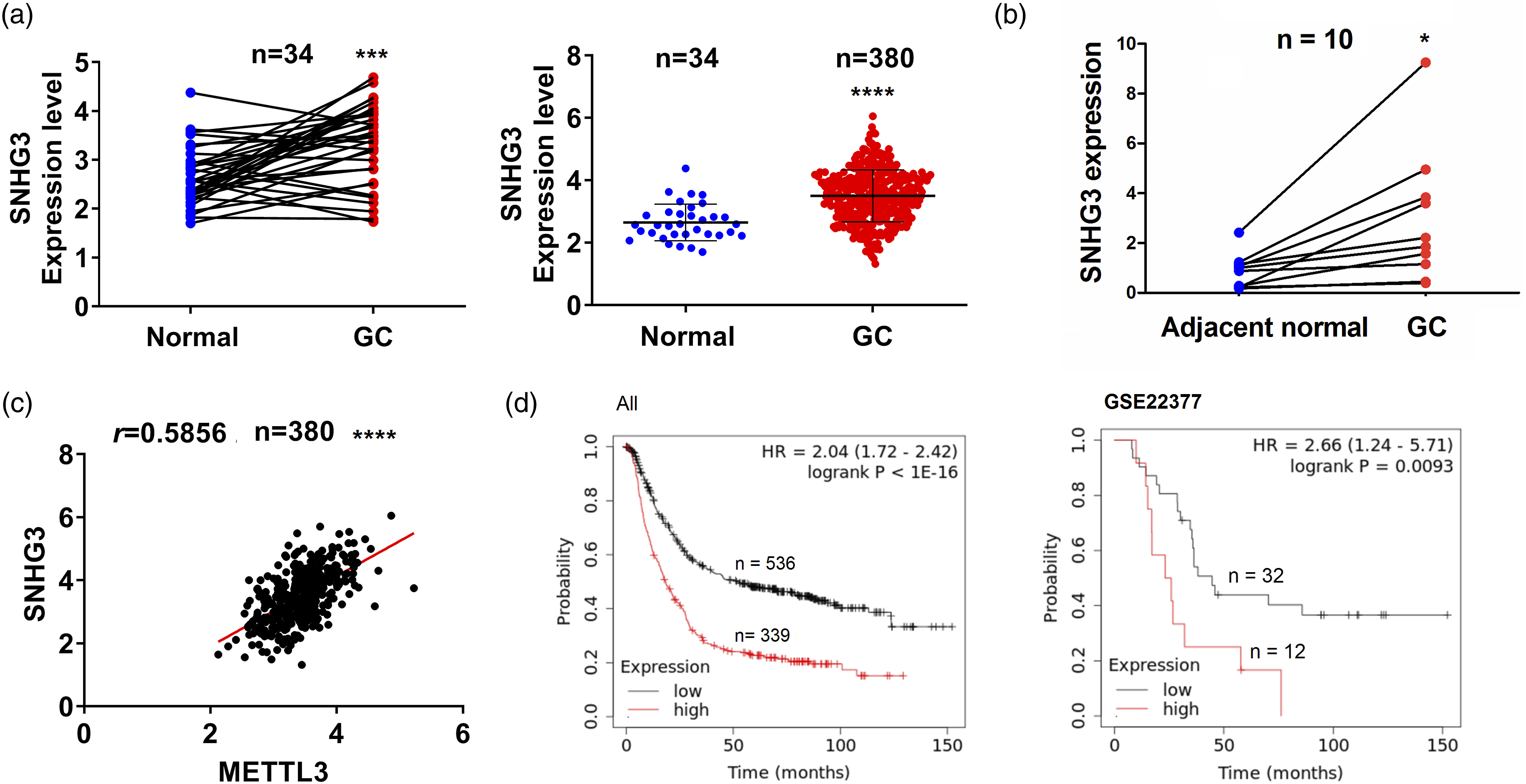
The miR-186-5p is sponged by SNHG3 and indicates favorable survival in GC
According to the condition set (very high stringency > 5), we found that lncRNA SNHG3 only has the potential to bind to miR-186-5p. The binding sites between SNHG3 and miR-186-5p can be shown in Figure 6(a). The expression of miR-186-5p was significantly decreased in 10 pairs of GC tissue samples in our cohort by qRT-PCR (Figure 6(b)) and possessed a negative correlation with SNHG3 expression in 368 GC tissue samples from TCGA cohort (Figure 6(c)). GC patients with miR-186-5p-high expression indicated a favorable overall survival as compared with those with miR-186-5p-low expression (Figure 6(d)). Then, we found that miR-186-5p mimics could reduce the luciferase activities of WT SNHG3 3’UTR rather than Mut SNHG3 3’UTR in HEK293T cells (Figure 6(e)). The qRT-PCR showed that knockdown of SNHG3 increased the expression levels of miR-186-5p in MKN-45, and BGC-823 cells (Figure 6(f) and (g)). MTT assay indicated that overexpression of SNHG3 promoted cell proliferation activity, but miR-186-5p mimics reversed SNHG3-induced proliferation-promoting effect in GC cells (Figure 6(h)). The miR-186-5p was sponged by SNHG3 and indicated favorable survival in GC. (a) The interaction of miR-186-5p with WT or Mut SNHG3 3’UTR. (b) Examination of the levels of miR-186-5p relative to the expression of U6 in GC. (c) The correlation between miR-186-5p and SNHG3 expression in GC. (d) Kaplan–Meier analysis of the association of miR-186-5p with overall survival in patients with GC. (e) The luciferase activities of WT or Mut SNHG3 3’UTR after treatment with miR-186-5p mimics in HEK293T cells. (f), (g) Examination of the levels of miR-186-5p after transfection with si-SNHG3 in GC cells. (h) Examination of the cell proliferation after treatment with miR-186-5p mimics and SNHG3 plasmids in GC cells.
The lncRNA SNHG3 abrogates METTL3 knockdown-induced miR-186-5p/cyclinD2 dysregulation in GC cells
To further confirm the regulation of METTL3 on lncRNA SNHG3 in GC cells, we first identified cyclinD2 as the potential target of miR-186-5p by starBaseV2.0 (https://starbase.sysu.edu.cn/starbase2/browseIntersectTargetSite.php) and RT-qPCR (Figure 7(a)), and Western blotting analysis (Figure 7(b)) indicated that miR-186-5p mimics could reduce the mRNA and protein expression levels of cyclinD2 in GC cells. Then, knockdown of METTL3 repressed the expression levels of SNHG3, but overexpression of SNHG3 reversed METTL3 knockdown-induced inhibitory effects in GC cells (Figure 7(c)). Reversely, knockdown of METTL3 increased the expression levels of miR-186-5p, but overexpression of SNHG3 reversed METTL3 knockdown-induced miR-186-5p upregulation in MKN-45 and BGC-823 cells (Figure 7(d)). Moreover, RT-qPCR (Figure 7(e)) and Western blotting analysis (Figure 7(f)) unveiled that knockdown of METTL3 could downregulate cyclinD2, but overexpression of SNHG3 reversed METTL3 knockdown-caused cyclinD2 downregulation in MKN-45 and BGC-823 cells. SNHG3 reversed METTL3 knockdown-induced miR-186-5p/cylcinD2 dysregulation in GC cells. (a) RT-qPCR and (b) Western blotting analysis of the mRNA and protein levels of cylcinD2 after the transfection with miR-186-5p mimics into MKN-45 and BGC-823 cells. (c) Examination of the levels of SNHG3 after co-transfection with si-METTL3 and SNHG3 overexpression plasmids in GC cells. (d) Examination of the levels of miR-186-5p relative to the expression of U6 after co-transfection with si-METTL3 and SNHG3 plasmids in GC cells. (e) RT-qPCR and (f) Western blotting analysis of the mRNA and protein levels of cyclinD2 after co-transfection with si-METTL3 and SNHG3 plasmids in GC cells.
Discussion
Increasing data reveals that m6A RNA methylation is responsible for regulating tumorigenesis and progression.5–11 METTL3 has been proved to be upregulated in multiple malignances including GC,5,12–17 and indicate poor prognosis,5,14,16,17 but it is downregulated and associated with favorable prognosis in RCC. 19 In the present study, in accordance with previous reports,5,20–23 we found that elevated expression of METTL3 was related to poor overall survival, acting as a prognostic marker in GC. METTL3 might be a potential prognostic factor in GC.
METTL3 can accelerate carcinogenesis,12–17 epithelial-mesenchymal transition (EMT), and liver metastasis of GC.5,20 However, METTL3 can act as a tumor suppressor in RCC and impair cell proliferation and invasion. 19 In consistence with previous reports,5,20 we unveiled that knockdown of METTL3 inhibited GC progression, whereas ectopic expression of METTL3 harbored the reverse effects, revealing that METTL3 might be a pro-oncogenic factor in GC.
METTL3 can act by mediating m6A modification of mRNAs 4 and noncoding RNAs 24 in cancer. METTL3 promotes tumorigenesis and progression by m6A modification of HDGF and ZMYM1 in GC,5,20 USP4 in GC, 12 YAP in NSCLC, 13 Notch1 in ESCC, 14 Myc in BCa, 15 BMI1 in OSCC, 16 and CRB3 in CRC. 17 METTL3 promotes the maturation of pri-miR221/222 in BCa 25 and pri-miR-1246 in CRC. 26 METTL3 mediates lncRNA stability, such as FOXD2-AS1 in cervical cancer, 27 SNHG7 in GC, 28 and MALAT1 in glioma. 29 LncRNAs also play crucial roles in GC progression. 30 It has been shown that lncRNA SNHG3 acts as a tumor-promoting factor in multiple malignancies including GC31–33 but acts as a tumor suppressor in papillary thyroid carcinoma. 34 We herein found that knockdown of METTL3 decreased the m6A modification levels of SNHG3, and overexpression of SNHG3 abolished METTL3 knockdown-led proliferation inhibition and unveiled poor survival. METTL3-mediated m6A modification of SNHG3 promoted GC growth.
It is widely acknowledged that lncRNA SNHG3 can sponge miRNAs to participate in cancer progression. For example, SNHG3 acts by regulating miR-539/RUNX2 axis in CRC, 35 sponging miR-154-3p in breast cancer, 36 regulating miR-1343-3p/NFIX pathway in NSCLC, 37 and targeting miR-139-5p/-326 in GC.38,39 In addition, miR-186-5p has been reported to suppress tumor growth and metastasis by targeting ZEB1 in CRC, 40 RAB27A/B in BCa, 41 and ITGA6 in OSCC. 42 We herein found that, on the one hand, miR-186-5p was downregulated in GC tissue samples and associated with favorable survival in patients with GC. On the other hand, miR-186-5p could be sponged by SNHG3 and reverse SNHG3 knockdown-induced anti-GC proliferation. Furthermore, we identified cyclinD2 as a target of miR-186-5p, and SNHG3 abrogated METTL3 knockdown-induced miR-186-5p upregulation and cyclinD2 downregulation in GC cells.
However, our present study had some shortcomings. On the one hand, we needed to increase the sample size to verify the expression and prognosis of METTL3 in GC. On the other hand, the in vivo study or the xenograft tumor model was required for validating the effect of METTL3 on the tumorigenesis of GC. In addition to lncRNA SNHG3, whether METTL3 can mediate the m6A medication of other noncoding RNAs such as circRNAs to influence GC progression need be further explored.
Conclusion
METTL3-mediated m6A modification of lncRNA SNHG3 promotes GC growth by regulating miR-186-5p/cyclinD2 axis and may provide a potential therapeutic target for GC.
Supplemental Material
Supplemental Material - METTL3-mediated m6A modification of lncRNA SNHG3 accelerates gastric cancer progression by modulating miR-186-5p/cyclinD2 axis
Supplemental Material for METTL3-mediated m6A modification of lncRNA SNHG3 accelerates gastric cancer progression by modulating miR-186-5p/cyclinD2 axis by Guo Ji, Xiu Wang and Hao Xi in International Journal of Immunopathology and Pharmacology.
Ethical statement
Ethical approval
Ethical approval for this study was obtained from the Ethics Committee of Shanghai Tenth People’s Hospital (2021-YS-009).
Footnotes
Author contributions
HX designed the manuscript. GJ and XW performed the experiments and contributed equally to the present work. GJ wrote the manuscript, analyzed the data, and HX revised the manuscript. All the authors checked and finalized the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by Shanghai Natural Science Foundation (21ZR1448700).
Informed consent
The Cancer Genome Atlas database was used for analysis of the expression and prognosis of METTL3, SNHG3, and miR-186-5p in GC. 10 pairs of GC samples were provided by Department of Pathology, Shanghai Tenth People’s Hospital for analysis of the expression of METTL3 in GC.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
